Abstract
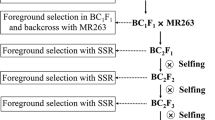
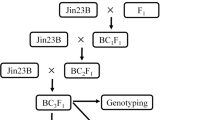
The study was conducted to identify the suitable SSR markers in relation to blast disease of rice. Backcross breeding was applied and BC2F1 generation was produced using rice variety Pongsu Seribu 2 as blast resistant parent and MR219 as susceptible parent. Eleven SSR markers related to blast resistance genes (Pi gene) were found highly polymorphic between the parental lines. According to single-gene model only two markers, RM208 and RM206, fitted to the expected test cross ratio (1:1) in BC2F1 generation with the relation to resistant and susceptible plants. Further BC2F1 population carrying 320 plants were inoculated with the highly virulent pathotype P7.2 of Magnaporthe oryze. Chi-square (χ2) test for single-gene model showed goodness of fit (p = 0.45) to the expected segregation ratio (1:1) for phenotypic segregation in BC2F1 population. In marker segregation analysis two markers RM208 (χ2 = 1.513, p = 0.218) and RM206 (χ2 = 0.613, P = 0.433) clearly produced goodness of fit to the expected test cross ratio (1:1) for the single-gene model. The present finding could help the rice breeders to monitor the blast resistance genes with appropriate and highly segregating SSR markers in rice varietal development program against blast disease.




Similar content being viewed by others
References
Akagi H, Yokozeki Y, Inagaki A, Fujimura T (1996) Microsatellite DNA markers for rice chromosomes. Theor Appl Genet 93:1071–1077
Ashkani S, Rafii M, Sariah M, Siti NAA, Rusli I, Harun A, Latif M (2011) Analysis of simple sequence repeat markers linked with blast disease resistance genes in a segregating population of rice (Oryza sativa). Genet Mol Res 10:1345–1355
Ballini E, Morel JB, Droc G, Price A, Courtois B, Notteghem JL, Tharreau D (2008) A genome-wide meta-analysis of rice blast resistance genes and quantitative trait loci provides new insights into partial and complete resistance. Mol Plant Microbe Interact 21:859–868
Barman S, Gowda M, Venu R, Chattoo B (2004) Identification of a major blast resistance gene in the rice cultivar ‘Tetep’. Plant Breed 123:300–302
Bres-Patry C, Lorieux M, Clement G, Bangratz M, Ghesquière A (2001) Heredity and genetic mapping of domestication-related traits in a temperate japonica weedy rice. Theor Appl Genet 102:118–126
Chen H (2001) Population structure of Pyricularia grisea from central and southern China and comparative mapping of QTL for blast-and bacterial blight-resistance in rice and barley. PhD thesis, Huazhong Agriculture University, Wuhan, China
Chen X, Shang J, Chen D, Lei C, Zou Y, Zhai W, Liu G, Xu J, Ling Z, Cao G, Ma B, Wang Y, Zhao X, Li S, Zhu L (2006) A B-lectin receptor kinase gene conferring rice blast resistance. Plant J 46:794–804
Cho YG et al (2000) Diversity of microsatellites derived from genomic libraries and GenBank sequences in rice (Oryza sativa L.). Theor Appl Genet 100:713–722
Correa-Victoria FJ, Zeigler RS (1995) Stability of partial and complete resistance in rice to Pyricularia grisea under rainfed upland conditions in eastern Colombia. Phytopathology 85(9):977–982
Deng Y, Zhu X, Shen Y, He Z (2006) Genetic characterization and fine mapping of the blast resistance locus Pigm(t) tightly linked to Pi2 and Pi9 in a broad-spectrum resistant Chinese variety. Theor Appl Genet 113:705–713
Doyle JJ, Doyle JL (1990) Isolation of plant DNA from fresh tissue. Focus 12:13–15
Ellingboe AH, Chao CCT (1994) Genetic interactions in Magnaporthe grisea that affect cultivar 463 specific avirulence/virulence in rice. In: Zeigler RS, Leong SA, Teng PS (eds) Rice blast disease. CAB International, Wallingford, pp 51–64
Filippi MC, Prabhu AS (2001) Phenotypic virulence analysis of Pyricularia grisea isolates from Brazilian upland rice cultivars. Pesq Agrop Brasileira 36(1):27–35
Huang H, Huang L, Feng G, Wang S, Wang Y, Liu J, Jiang N, Yan W, Xu L, Sun P, Liu Z, Pan S, Liu X, Xiao Y, Liu E, Dai L, Wang G (2010) Molecular mapping of the new blast resistance genes Pi47 and Pi48 in the durably resistant local rice cultivar Xiangzi 3150. Phytopathology 101(5):620–626
IRRI (1996) Standard evaluation system for rice. 4th edn. The International Network for Genetic Evaluation of Rice, Genetic Resources Center. International Rice Research Institute, Manila, 52
Kumbhar S, Kulwal P, Patil J, Gaikwad A, Jadhav A (2013) Inheritance of blast resistance and identification of SSR marker associated with it in rice cultivar RDN 98-2. J Genet 92(2):317–321
Lin F, Chen S, Que ZQ, Wang L, Liu XQ, Pan QH (2007) The blast resistance gene Pi37 encodes a nucleotide binding site-leucine rich repeat protein and is a member of a resistance gene cluster on rice chromosome 1. Genetics 177:1871–1880
Liu XQ, Wang L, Chen S, Lin F, Pan QH (2005) Genetic and physical mapping of Pi36(t), a novel rice blast resistance gene located on rice chromosome 8. Mol Gen Genomics 274:394–401
Liu JL, Wang XJ, Thomas M, Hu YJ, Liu XL, Dai LY, Wang GL (2010) Recent progress and understanding of the molecular mechanisms of the rice—Magnaporthe oryzae interaction. Mol Plant Pathol 11(3):419–427
Mackill D, Bonman J, Suh H, Srilingam R (1985) Genes for resistance to Philippine isolates of the rice blast pathogen. Rice Genet Newsl 2:80–81
McCouch SR, Kochert G, Yu Z, Wang Z, Khush G, Coffman W, Tanksley S (1988) Molecular mapping of rice chromosomes. Theor Appl Genet 76:815–829
McCouch SR, Teytelman L, Xu Y, Lobos KB, Clare K, Walton M, Fu B, Maghirang R, Li Z, Xing Y (2002) Development and mapping of 2240 new SSR markers for rice (Oryza sativa L.). DNA Res 9(6):199–207
Miah G, Rafii MY, Ismail MR, Puteh AB, Rahim HA, Islam KN, Latif MA (2013) A review of microsatellite markers and their applications in rice breeding programs to improve blast disease resistance. Int J Mol Sci 14(11):22499–22528
Moncada P et al (2001) Quantitative trait loci for yield and yield components in an Oryza sativa× Oryza rufipogon BC2F2 population evaluated in an upland environment. Theor Appl Genet 102:41–52
Pan Q, Wang L, Ikehashi H, Tanisaka T (1996) Identification of a new blast resistance gene in the indica rice cultivar Kasalath using Japanese differential cultivars and isozyme markers. Phytopathology 86:1071–1075
Persaud M, Kumar A, Sengar RBS, Abhinav S, Dantre RK, Shrivastava MN (2007) Genetic analysis of blast (Pyricularia grisea Sacc.) resistance in rice (Oryza sativa L.). J Biol Sci 7(1):215–217
Prabhu M, Veeraragavathatham D, Srinivasa K (2003) Effect of nitrogen and phosphorous on growth and yield of brinjal. South Indian Hortic 51:152–156
Rahim HA, Bhuiyan MAR, Saad A, Azhar M, Wickneswari R (2013) Identification of virulent pathotypes causing rice blast disease (Magnaporthe oryzae) and study on single nuclear gene inheritance of blast resistance in F2 population derived from Pongsu Seribu 2 × Mahshuri. Aust J Crop Sci 7:1597–1605
RoyChowdhury M, Jia Y, Jackson A, Jia MH, Fjellstrom R, Cartwright RD (2012) Analysis of rice blast resistance gene Pi-z in rice germplasm using pathogenicity assays and DNA markers. Euphytica 184:35–46
Sharma T et al (2005) High-resolution mapping, cloning and molecular characterization of the Pi-k h gene of rice, which confers resistance to Magnaporthe grisea. Mol Gen Genomics 274:569–578
Sharma R, Shrestha S, Pandey M (2007) Inheritance of blast resistance and associated microsatellite markers in rice cultivar ‘Laxmi’. J Phytopathol 155(11–12):749–753
Shu QY (2009) Induced plant mutations in the genomics era. Proceedings of an International Joint FAO/IAEA Symposium, Vienna, Austria, 2008. Paper presented at the Induced plant mutations in the genomics era. Proceedings of an International Joint FAO/IAEA Symposium, Vienna, Austria, 2008
Singh A et al (2012) Molecular breeding for the development of multiple disease resistance in Basmati rice AoB plants 2012:pls029
Sirithunya P, Tragoonrung S, Vanavichit A, Pa-In N, Vongsaprom C, Toojinda T (2002) Quantitative trait loci associated with leaf and neck blast resistance in recombinant inbred line population of rice (Oryza sativa). DNA Res 9:79–88
Tanweer FA, Rafii MY, Sijam K, Rahim HA, Ahmed F, Latif MA (2015) Cloning and characterization of two major blast resistance genes Pi-b and Pi-kh from Malaysian rice variety Pongsu Seribu 2. Plant Omic 8(3):257–263
Temnykh S et al (2000) Mapping and genome organization of microsatellite sequences in rice (Oryza sativa L.). Theor Appl Genet 100:697–712
Wu K-S, Tanksley SD (1993) Abundance, polymorphism and genetic mapping of microsatellites in rice. Mol Gen Genet MGG 241:225–235
Xiao WM, Yang QY, Wang H, Guo T, Liu YZ, Zhu XY, Chen ZQ (2010) Identification and fine mapping of a resistance gene to Magnaporthe oryzae in a space-induced rice mutant. Mol Breed 28:303–312
Xu X, Chen H, Fujimura T, Kawasaki S (2008) Fine mapping of a strong QTL of field resistance against rice blast, Pikahei-1(t), from upland rice Kahei, utilizing a novel resistance evaluation system in the green house. Theor Appl Genet 117:997–1008
Yu Z, Mackill D, Bonman J (1987) Inheritance of resistance to blast in some traditional and improved rice cultivars. Phytopathology (USA)
Zhu M, Wang L, Pan Q (2004) Identification and characterization of a new blast resistance gene located on rice chromosome 1 through linkage and differential analyses. Phytopathology 94:515–519
Zou J, Pan X, Chen Z, Xu J, Lu J, Zhai W, Zhu L (2000) Mapping quantitative trait loci controlling sheath blight resistance in two rice cultivars (Oryza sativa L.). Theor Appl Genet 101:569–573
Acknowledgments
The authors would like to acknowledge Long term Research Grant Scheme (LRGS), Food Security Project, Ministry of Higher Education, Malaysia, for the financial support to conduct research on rice breeding. The author would also like to acknowledge Sindh Agriculture University Tandojam Sindh Pakistan for providing financial support.
Conflict of interest
The authors declare that there is no conflict of interests regarding the publication of this paper.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Tanweer, F.A., Rafii, M.Y., Sijam, K. et al. Identification of suitable segregating SSR markers for blast resistance in rice using inheritance and disease reaction analysis in backcross families. Australasian Plant Pathol. 44, 619–627 (2015). https://doi.org/10.1007/s13313-015-0380-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13313-015-0380-5