Abstract
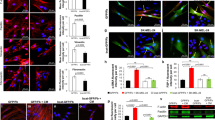
Cancer-associated fibroblasts (CAFs) are the crucial components of the dynamic tumor microenvironment, which not only supports the growth and metastasis of melanoma but also contributes to drug resistance in melanoma treatment. We recently discovered that loss of β-catenin signaling deactivated stromal fibroblasts and reduced the production of paracrine factors and extracellular matrix proteins. Based on this finding, we aimed to determine whether melanoma growth could be suppressed by targeted deactivation of CAFs via β-catenin ablation using a combination of in vitro and in vivo approaches. Using an in vitro three-dimensional (3D) tumor co-culture model, we showed that β-catenin-deficient fibroblasts lost the ability to respond to melanoma cell stimulation and to support the growth of B16F10 melanoma cells. To determine the in vivo effects of CAF deactivation on melanoma growth, we designed a novel genetic approach to ablate β-catenin expression in melanoma-associated fibroblasts only after melanoma tumor was formed. As expected, our observation showed that development of B16F10 melanoma was significantly delayed when β-catenin expression was ablated in CAFs. We determined that inhibition of tumor growth was due to decreased melanoma cell proliferation and increased cell death. Further analysis revealed that CAF deactivation caused the downregulation of the MAPK/ERK signaling cascade and S and G2/M phase cell cycle arrest in B16F10 melanoma cells. Overall, our data emphasize the significance of targeting CAFs as a potential novel therapeutic approach to improve melanoma treatment by creating a tumor-suppressive microenvironment through tumor-stroma interactions.






Similar content being viewed by others
References
Bertolotto C. Melanoma: from melanocyte to genetic alterations and clinical options. Scientifica. 2013;2013:635203. doi:10.1155/2013/635203.
Siegel RL, Miller KD, Jemal A. Cancer statistics, 2015. CA Cancer J Clin. 2015;65(1):5–29. doi:10.3322/caac.21254.
Schadendorf D, Hauschild A. Melanoma in 2013: melanoma—the run of success continues. Nat Rev Clin Oncol. 2014;11(2):75–6. doi:10.1038/nrclinonc.2013.246.
Livingstone E, Zimmer L, Vaubel J, Schadendorf D. BRAF, MEK and KIT inhibitors for melanoma: adverse events and their management. Chin Clin Oncol. 2014;3(3):29–47.
Lindsay JN, Barras M. Facing the challenges of new melanoma-targeted therapies: treatment of severe fevers associated with dabrafenib/trametinib combination therapy. J Oncol Pharm Pract : Off Publ Int Soc Oncol Pharm Pract. 2015;21(4):293–5. doi:10.1177/1078155214527859.
Slominski AT, Carlson JA. Melanoma resistance: a bright future for academicians and a challenge for patient advocates. Mayo Clin Proc. 2014;89(4):429–33. doi:10.1016/j.mayocp.2014.02.009.
Slominski A, Tobin DJ, Shibahara S, Wortsman J. Melanin pigmentation in mammalian skin and its hormonal regulation. Physiol Rev. 2004;84(4):1155–228. doi:10.1152/physrev.00044.2003.
Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011;144(5):646–74. doi:10.1016/j.cell.2011.02.013.
Hanahan D, Weinberg RA. The hallmarks of cancer. Cell. 2000;100(1):57–70. doi:10.1016/S0092-8674(00)81683-9.
Lu P, Weaver VM, Werb Z. The extracellular matrix: a dynamic niche in cancer progression. J Cell Biol. 2012;196(4):395–406. doi:10.1083/jcb.201102147.
Peng Q, Zhao L, Hou Y, Sun Y, Wang L, Luo H, et al. Biological characteristics and genetic heterogeneity between carcinoma-associated fibroblasts and their paired normal fibroblasts in human breast cancer. PLoS One. 2013;8(4):e60321. doi:10.1371/journal.pone.0060321.
Cirri P, Chiarugi P. Cancer associated fibroblasts: the dark side of the coin. Am J Cancer Res. 2011;1(4):482–97.
Rasanen K, Vaheri A. Activation of fibroblasts in cancer stroma. Exp Cell Res. 2010;316(17):2713–22. doi:10.1016/j.yexcr.2010.04.032.
Cornil I, Theodorescu D, Man S, Herlyn M, Jambrosic J, Kerbel RS. Fibroblast cell interactions with human melanoma cells affect tumor cell growth as a function of tumor progression. Proc Natl Acad Sci U S A. 1991;88(14):6028–32.
Zhou L, Yang K, Andl T, Wickett RR, Zhang Y. Perspective of targeting cancer-associated fibroblasts in melanoma. J Cancer. 2015;6(8):717–26. doi:10.7150/jca.10865.
Gaggioli C, Sahai E. Melanoma invasion—current knowledge and future directions. Pigment Cell Res / Sponsored Eur Soc Pigment Cell Res Int Pigment Cell Soc. 2007;20(3):161–72. doi:10.1111/j.1600-0749.2007.00378.x.
Li G, Satyamoorthy K, Meier F, Berking C, Bogenrieder T, Herlyn M. Function and regulation of melanoma-stromal fibroblast interactions: when seeds meet soil. Oncogene. 2003;22(20):3162–71. doi:10.1038/sj.onc.1206455.
Flach EH, Rebecca VW, Herlyn M, Smalley KS, Anderson AR. Fibroblasts contribute to melanoma tumor growth and drug resistance. Mol Pharm. 2011;8(6):2039–49. doi:10.1021/mp200421k.
Santos AM, Jung J, Aziz N, Kissil JL, Pure E. Targeting fibroblast activation protein inhibits tumor stromagenesis and growth in mice. J Clin Invest. 2009;119(12):3613–25. doi:10.1172/JCI38988.
Shao H, Cai L, Grichnik JM, Livingstone AS, Velazquez OC, Liu ZJ. Activation of Notch1 signaling in stromal fibroblasts inhibits melanoma growth by upregulating WISP-1. Oncogene. 2011;30(42):4316–26. doi:10.1038/onc.2011.142.
Guan JC, J. Tumor microenvironment: the promising target for tumor therapy. Cancer Cell Microenviron. 2014(1):17–9. doi:10.14800/ccm.81.
Zhou L, Yang K, Randall Wickett R, Zhang Y. Dermal fibroblasts induce cell cycle arrest and block epithelial-mesenchymal transition to inhibit the early stage melanoma development. Cancer Med. 2016;5(7):1566–79. doi:10.1002/cam4.707.
Zheng B, Zhang Z, Black CM, de Crombrugghe B, Denton CP. Ligand-dependent genetic recombination in fibroblasts : a potentially powerful technique for investigating gene function in fibrosis. Am J Pathol. 2002;160(5):1609–17. doi:10.1016/S0002-9440(10)61108-X.
Zhang Y, Tomann P, Andl T, Gallant NM, Huelsken J, Jerchow B, et al. Reciprocal requirements for EDA/EDAR/NF-kappaB and Wnt/beta-catenin signaling pathways in hair follicle induction. Dev Cell. 2009;17(1):49–61. doi:10.1016/j.devcel.2009.05.011.
Whittaker P, Kloner RA, Boughner DR, Pickering JG. Quantitative assessment of myocardial collagen with picrosirius red staining and circularly polarized light. Basic Res Cardiol. 1994;89(5):397–410.
Metzger D, Clifford J, Chiba H, Chambon P. Conditional site-specific recombination in mammalian-cells using a ligand-dependent chimeric Cre recombinase. P Natl Acad Sci USA. 1995;92(15):6991–5. doi:10.1073/pnas.92.15.6991.
Brault V, Moore R, Kutsch S, Ishibashi M, Rowitch DH, McMahon AP, et al. Inactivation of the beta-catenin gene by Wnt1-Cre-mediated deletion results in dramatic brain malformation and failure of craniofacial development. Development. 2001;128(8):1253–64.
Nyga A, Cheema U, Loizidou M. 3D tumour models: novel in vitro approaches to cancer studies. J Cell Commun Signal. 2011;5(3):239–48. doi:10.1007/s12079-011-0132-4.
Dufau I, Frongia C, Sicard F, Dedieu L, Cordelier P, Ausseil F, et al. Multicellular tumor spheroid model to evaluate spatio-temporal dynamics effect of chemotherapeutics: application to the gemcitabine/CHK1 inhibitor combination in pancreatic cancer. BMC Cancer. 2012;12:15. doi:10.1186/1471-2407-12-15.
Laurent J, Frongia C, Cazales M, Mondesert O, Ducommun B, Lobjois V. Multicellular tumor spheroid models to explore cell cycle checkpoints in 3D. BMC Cancer. 2013;13:73. doi:10.1186/1471-2407-13-73.
Pietras K, Pahler J, Bergers G, Hanahan D. Functions of paracrine PDGF signaling in the proangiogenic tumor stroma revealed by pharmacological targeting. PLoS Med. 2008;5(1):e19. doi:10.1371/journal.pmed.0050019.
Choi SY, Sung R, Lee SJ, Lee TG, Kim N, Yoon SM, et al. Podoplanin, alpha-smooth muscle actin or S100 A4 expressing cancer-associated fibroblasts are associated with different prognosis in colorectal cancers. J Korean Med Sci. 2013;28(9):1293–301. doi:10.3346/jkms.2013.28.9.1293.
Collins CA, Kretzschmar K, Watt FM. Reprogramming adult dermis to a neonatal state through epidermal activation of β-catenin. Development. 2011;138(23):5189–99.
Driskell RR, Lichtenberger BM, Hoste E, Kretzschmar K, Simons BD, Charalambous M, et al. Distinct fibroblast lineages determine dermal architecture in skin development and repair. Nature. 2013;504(7479):277–81. doi:10.1038/nature12783.
Sharon Y, Alon L, Glanz S, Servais C, Erez N. Isolation of normal and cancer-associated fibroblasts from fresh tissues by Fluorescence Activated Cell Sorting (FACS). J Vis Exp : JoVE. 2013;71:e4425. doi:10.3791/4425.
Rieger AM, Nelson KL, Konowalchuk JD, Barreda DR. Modified annexin V/propidium iodide apoptosis assay for accurate assessment of cell death. J Vis Exp : JoVE. 2011;50:2597. doi:10.3791/2597.
Klein EA, Assoian RK. Transcriptional regulation of the cyclin D1 gene at a glance. J Cell Sci. 2008;121(Pt 23):3853–7. doi:10.1242/jcs.039131.
Giacinti C, Giordano A. RB and cell cycle progression. Oncogene. 2006;25(38):5220–7. doi:10.1038/sj.onc.1209615.
Hocker TL, Singh MK, Tsao H. Melanoma genetics and therapeutic approaches in the 21st century: moving from the benchside to the bedside. J Invest Dermatol. 2008;128(11):2575–95. doi:10.1038/jid.2008.226.
Maio M, Grob JJ, Aamdal S, Bondarenko I, Robert C, Thomas L, et al. Five-year survival rates for treatment-naive patients with advanced melanoma who received ipilimumab plus dacarbazine in a phase III trial. J Clin Oncol Off J Am Soc Clin Oncol. 2015;33(10):1191–6. doi:10.1200/JCO.2014.56.6018.
Luo H, Tu G, Liu Z, Liu M. Cancer-associated fibroblasts: a multifaceted driver of breast cancer progression. Cancer Lett. 2015;361(2):155–63. doi:10.1016/j.canlet.2015.02.018.
Orimo A, Weinberg RA. Stromal fibroblasts in cancer: a novel tumor-promoting cell type. Cell Cycle. 2006;5(15):1597–601. doi:10.4161/cc.5.15.3112.
Gonda TA, Varro A, Wang TC, Tycko B. Molecular biology of cancer-associated fibroblasts: can these cells be targeted in anti-cancer therapy? Semin Cell Dev Biol. 2010;21(1):2–10. doi:10.1016/j.semcdb.2009.10.001.
Loeffler M, Kruger JA, Niethammer AG, Reisfeld RA. Targeting tumor-associated fibroblasts improves cancer chemotherapy by increasing intratumoral drug uptake. J Clin Invest. 2006;116(7):1955–62. doi:10.1172/JCI26532.
Ivascu A, Kubbies M. Rapid generation of single-tumor spheroids for high-throughput cell function and toxicity analysis. J Biomol Screen. 2006;11(8):922–32. doi:10.1177/1087057106292763.
Kaufmann WK, Nevis KR, Qu P, Ibrahim JG, Zhou T, Zhou Y, et al. Defective cell cycle checkpoint functions in melanoma are associated with altered patterns of gene expression. J Invest Dermatol. 2008;128(1):175–87. doi:10.1038/sj.jid.5700935.
Weinberg RA. The retinoblastoma protein and cell cycle control. Cell. 1995;81(3):323–30.
Acknowledgments
We thank Dr. Benoit de Crombrugghe for the Col1α2-CreER mice.
Funding
This work was supported by the Elsa U. Pardee Foundation, University of Cincinnati-MRA Young Investigator Award (grant number 300586), the Cincinnati Cancer Center—Mentor-Mentee Award, the Skin Cancer Foundation—The Dr. Marcia Robbins-Wilf Research Grant Award, and the Harry J. LIoyd Trust Research Award.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
None.
Rights and permissions
About this article
Cite this article
Zhou, L., Yang, K., Wickett, R.R. et al. Targeted deactivation of cancer-associated fibroblasts by β-catenin ablation suppresses melanoma growth. Tumor Biol. 37, 14235–14248 (2016). https://doi.org/10.1007/s13277-016-5293-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13277-016-5293-6