Abstract
Background
There were significant differences in the change of moisture content and grain composition at the late stage of grain development among different maize varieties, but the regulation mechanism is not clear.
Objective
To explore the key genes causing the variation in physiological traits of two typical maize inbred lines in late grain development.
Methods
The grains at different development stages were selected as materials to determine the content of water, sucrose, starch and ABA. Transcriptomic and proteomic analysis of the materials were performed to screen relevant genes.
Results
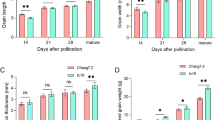
The grain dehydration rate and the content of sucrose, starch and ABA were showed significant differences between two varieties in the late stage of grain development. The enrichment analysis of common differentially expressed genes (proteins) showed that most of the genes (proteins) were enriched in the extracellular region. The downregulated genes were mainly concentrated in carbohydrate metabolism and lipid metabolism, while the upregulated genes were mainly in response to stress. Furthermore, this study also identified many key candidate genes (dehydrin genes, pathogenesis-related genes, sucrose synthase and secondary metabolites related genes) related to late grain development of maize.
Conclusions
The suggested genes related to late grain development of maize can be candidates for further functional study.






Similar content being viewed by others
References
Allagulova CR, Gimalov FR, Shakirova FM, Vakhitov VA (2003) The plant dehydrins: structure and putative functions. Biochem Mosc 68:945–951
Bufe A, Spangfort MD, Kahlert H, Schlaak M, Becker WM (1996) The major birch pollen allergen, Bet v 1, shows ribonuclease activity. Planta 199:413–415
Cavalieri AJ, Smith OS (1985) Grain filling and field drying of a set of maize hybrids released from 1930 to 1982. Crop Sci 25:856–860
Chen WP, Chen PD, Liu DJ, Kynast R, Friebe B, Velazhahan R, Muthukrishnan S, Gill BS (1999) Development of wheat scab symptoms is delayed in transgenic wheat plants that constitutively express a rice thaumatin-like protein gene. Theor Appl Genet 99:755–760
Chowdhury MH, Buchele WF (1978) The nature of corn kernel damage inflicted in the shelling crescent of grain combines. IJSTE 21:610–614
Close TJ (1996) Dehydrins: emergence of a biochemical role of a family of plant dehydration proteins. Physiol Plantarum 97:795–803
Close TJ, Kortt AA, Chandler PM (1989) A cDNA-based comparison of dehydration-induced proteins (dehydrins) in barley and corn. Plant Mol Biol 13:95–108
Cross HZ, Chyle JR, Hammond J (1987) Divergent selecting for ear moisture in early maize. Crop Sci 27:914–918
Dai L, Wu L, Dong Q, Zhang Z, Wu N, Song Y, Lu S, Wang P (2017) Genome-wide association study of field grain drying rate after physiological maturity based on a resequencing approach in elite maize germplasm. Euphytica 213:182
Danquah A, Zelicourt A, Colcombet J, Hirt H (2014) The role of ABA and MAPK signaling pathways in plant abiotic stress responses. Biotechnol Adv 32:40–52
Dixon RA (2001) Natural products and plant disease resistance. Nature 411:843–847
Dong NQ, Lin HX (2021) Contribution of phenylpropanoid metabolism to plant development and plant-environment interactions. J Integr Plant Biol 63:180–209
Drira M, Saibi W, Amara I, Masmoudi K, Hanin M, Brini F (2015) Wheat dehydrin K-segments ensure bacterial stress tolerance, antiaggregation and antimicrobial effects. Appl Biochem Biotechnol 175:3310–3321
Dudley JW, Lambert RJ, Alexander DE (1971) Variability and relationships among characters in Zea mays L. synthetics with improved protein quality 1. Crop Sci 11:512–514
Dure L, Crouch M, Harada J, Ho THD, Mundy J, Quatrano R, Thomas T, Sung ZR (1989) Common amino acid sequence domains among the LEA proteins of higher plants. Plant Mol Biol 12:475–486
Halder T, Agarwal T, Ray S (2016) Isolation, cloning, and characterization of a novel Sorghum dehydrin (SbDhn2)protein. Protoplasma 253:1475–1488
Hicks D, Geadelmann G, Peterson R (1976) Drying rates of frosted maturing maize. Agron J 68:452–455
Holton TA, Cornish EC (1995) Genetics and biochemistry of anthocyanin biosynthesis. Plant Cell 7:1071–1083
Jin X, Fu Z, Ding D, Li W, Liu Z, Tang J (2013) Proteomic identification of genes associated with maize grain-filling rate. PLoS ONE 8:e59353
Kasprzewska A (2003) Plant chitinases–regulation and function. Cell Mol Biol Lett 8:809–824
Kebebe AZ, Reid LM, Zhu X, Wu J, Woldemariam T, Voloaca C, Xiang K (2015) Relationship between kernel drydown rate and resistance to gibberella ear rot in maize. Euphytica 201:79–88
Keller L, Surette MG (2006) Communication in bacteria: an ecological and evolutionary perspective. Nat Rev Microbiol 4:249–258
Kim D, Langmead B, Salzberg SL (2015) HISAT: a fast spliced aligner with low memory requirements. Nat Methods 12:357–360
L CT, C MN, C TJ (1994) Purification of a maize dehydrin. Protein Expres Purif 5:266–269
Lee JH, Lee J (2010) Indole as an intercellular signal in microbial communities. FEMS Microbiol Rev 34:426–444
Li B, Dewey CN (2011) RSEM: accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinform 12:323
Li W, Yu Y, Wang L, Luo Y, Peng Y, Xu Y, Liu X, Wu S, Jian L, Xu J et al (2021) The genetic architecture of the dynamic changes in grain moisture in maize. Plant Biotechnol J 19:1195–1205
Loon LCV, Strien EAV (1999) The families of pathogenesis-related proteins, their activities, and comparative analysis of PR-1 type proteins. Physiol Mol Plant 55:85–97
Loon LCV, Rep M, Pieterse CMJ (2006) Significance of inducible defense-related proteins in infected plants. Annu Rev Phytopathol 44:135–162
Luo SH, Hua J, Niu XM, Liu Y, Li CH, Zhou YY, Jing SX, Zhao X, Li SH (2013) Defense sesterterpenoid lactones from Leucosceptrum canum. Phytochemistry 86:29–35
Mahmood T, Kakishima M, Komatsu S (2009) Proteome analysis of probenazole-effect in rice-bacterial blight interactions. Protein Pept Lett 16:1041–1052
Matsuoka Y, Vigouroux Y, Goodman MM, Sanchez GJ, Buckler E, Doebley J (2002) A single domestication for maize shown by multilocus microsatellite genotyping. Proc Natl Acad Sci USA 99:6080–6084
Nakashima K, Yamaguchi-Shinozaki K (2013) ABA signaling in stress-response and seed development. Plant Cell Rep 32:959–970
Ou-Lee TM, Setter TL (1985) Enzyme activities of starch and sucrose pathways and growth of apical and Basal maize kernels. Plant Physiol 79:848–851
Plett S (1994) Corn kernel breakage as a function of grain moisture at harvest in a prairie environment. Can J Plant Sci 74:543–544
Powers MJ, Trent MS (2019) Intermembrane transport: Glycerophospholipid homeostasis of the Gram-negative cell envelope. Proc Natl Acad Sci USA 116:17147–17155
Roitsch T, González MC (2004) Function and regulation of plant invertases: sweet sensations. Trends Plant Sci 9:606–613
Sala RG, Andrade FH, Camadro EL, Cerono JC (2006) Quantitative trait loci for grain moisture at harvest and field grain drying rate in maize (Zea mays, L.). Theor Appl Genet 112:462–471
Sentz JC (1971) Genetic variances in a synthetic variety of maize estimated by two mating designs. Crop Sci 11:234–238
Simmonds MS, Stevenson PC (2001) Effects of isoflavonoids from Cicer on larvae of Heliocoverpa armigera. J Chem Ecol 27:965–977
Svensson J, Ismail AM, Palva ET, Close TJ (2002) Chapter 11—Dehydrins. Cell Mol Response Stress 3:155–171
Troyer AF, Ambrose WB (1971) Plant characteristics affecting field drying rate of ear corn. Crop Sci 11:529–531
Velazhahan R, Muthukrishnan S (2003) Transgenic tobacco plants constitutively overexpressing a rice thaumatin-like protein (PR-5) show enhanced resistance to alternaria alternata. Biol Plantarum 47:347–354
Waelti H, Buchele WF (1969) Factors affecting corn kernel damage in combine cylinders. Lowa State Univ 12:55–59
Wang N, Xiao B, Xiong L (2011) Identification of a cluster of PR4-like genes involved in stress responses in rice. J Plant Physiol 168:2212–2224
Wang Z, Wang X, Zhang L, Liu X, Di H, Li T, Jin X (2012) QTL underlying field grain drying rate after physiological maturity in maize (Zea MaysL.). Euphytica 185:521–528
Wen B, Zhou R, Feng Q, Wang Q, Wang J, Liu S (2015) IQuant: an automated pipeline for quantitative proteomics based upon isobaric tags. Proteomics 14:2280–2285
Wisniewski M, Webb R, Balsamo R, Close TJ, Yu X-M, Griffit M (1999) Purification, immunolocalization, cryoprotective, and antifreeze activity of PCA60: a dehydrin from peach (Prunus persica). Physiol Plantarum 105:600–608
Yamaguchi Y, Sano H (2001) The sulfate assimilation pathway in higher plants: recent progress regarding multiple isoforms and regulatory mechanisms. Plant Tissue Cult Lett 18:17–25
Yamaguchi-Shinozaki K, Shinozaki K (2006) Transcriptional regulatory networks in cellular responses and tolerance to dehydration and cold stresses. Annu Rev Plant Biol 57:781–803
Zhan J, Thakare D, Ma C, Lloyd A, Nixon NM, Arakaki AM, Burnett WJ, Logan KO, Wang D, Wang X et al (2015) RNA sequencing of laser-capture microdissected compartments of the maize kernel identifies regulatory modules associated with endosperm cell differentiation. Plant Cell 27:513–531
Zhang L, Liang XG, Shen S, Yin H, Zhou LL, Gao Z, Lv XY, Zhou SL (2018) Increasing the abscisic acid level in maize grains induces precocious maturation by accelerating grain filling and dehydration. Plant Growth Regul 86:65–79
Zhang J, Zhang F, Tang B, Ding Y, Xia L, Qi J, Mu X, Gu L, Lu D, Chen Y (2020) Molecular mapping of quantitative trait loci for grain moisture at harvest and field grain drying rate in maize (Zea mays L.). Physiol Plantarum 169:64–72
Zhao J, Davis LC, Verpoorte R (2005) Elicitor signal transduction leading to production of plant secondary metabolites. Biotechnol Adv 23:283–333
Zhou G, Hao D, Xue L, Chen G, Lu H, Zhang Z, Shi M, Huang XL, Mao Y (2018) Genome-wide association study of kernel moisture content at harvest stage in maize. Breed Sci 68:622–628
Acknowledgements
The authors are grateful for the transcriptome and iTRAQ analyses performed by the BGI Gene Co., Ltd. (Shenzhen, China).
Funding
This study was funded by Key Science and Technology Projects in Henan Province (Grant no. 212102110305) and Important Science and Technology Specific Projects in Henan Province (Grant no. 201300111100).
Author information
Authors and Affiliations
Contributions
SLC, XNJ, and ZWH conceived and designed research. XNJ, PXW and XXZ conducted the experiments. XNJ and XYW contributed new reagents or analytical tools. HJZ and XNJ analyzed the data. XNJ wrote the manuscript. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
All the authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Jin, X., Zhai, H., Wang, P. et al. Physiological and omics analysis of maize inbred lines during late grain development. Genes Genom 44, 993–1006 (2022). https://doi.org/10.1007/s13258-022-01279-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13258-022-01279-0