Abstract
Background
The increase in genetic alterations targeted by specific chemotherapy in lung cancer has led to the need for universal use of more comprehensive genetic testing, which has highlighted the development of a lung cancer diagnostic panel using next-generation sequencing.
Objective
We developed a hybridization capture-based massively parallel sequencing assay named Friendly, Integrated, Research-based, Smart and Trustworthy (FIRST)-lung cancer panel (LCP), and evaluated its performance.
Methods
FIRST-LCP comprises 64 lung cancer-related genes to test for various kinds of genetic alterations including single nucleotide variations (SNVs), insertions and deletions (indels), copy number variations (CNVs), and structural variations. To assess the performance of FIRST-LCP, we compiled test sets using HapMap samples or tumor cell lines with disclosed genetic information, and also tested our clinical lung cancer samples whose genetic alterations were known by conventional methods.
Results
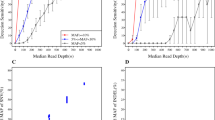
FIRST-LCP accomplished high sensitivity (99.4%) and specificity (100%) of the detection of SNVs. High precision was also achieved, with intra- or inter-run concordance rate of 0.99, respectively. FIRST-LCP detected indels and CNVs close to the expected allele frequency and magnitude, respectively. Tests with samples from lung cancer patients also identified all SNVs, indels and fusions.
Conclusion
Based on the current state of the art, continuous application of the panel design and analysis pipeline following up-to-date knowledge could ensure precision medicine for lung cancer patients.



Similar content being viewed by others
References
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30(15):2114–2120
Cheng DT, Mitchell TN, Zehir A et al (2015) Memorial Sloan Kettering-integrated mutation profiling of actionable cancer targets (MSK-IMPACT): a hybridization capture-based next-generation sequencing clinical assay for solid tumor molecular oncology. J Mol Diagn 17(3):251–264
Cibulskis K, Lawrence MS, Carter SL et al (2013) Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat Biotechnol 31(3):213–219
Costello M, Pugh TJ, Fennell TJ et al (2013) Discovery and characterization of artifactual mutations in deep coverage targeted capture sequencing data due to oxidative DNA damage during sample preparation. Nucleic Acids Res 41(6):e67
DePristo MA, Banks E, Poplin R et al (2011) A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat Genet 43(5):491–498
Frampton GM, Fichtenholtz A, Otto GA et al (2013) Development and validation of a clinical cancer genomic profiling test based on massively parallel DNA sequencing. Nat Biotechnol 31(11):1023–1031
Frampton GM, Ali SM, Rosenzweig M et al (2015) Activation of MET via diverse exon 14 splicing alterations occurs in multiple tumor types and confers clinical sensitivity to MET inhibitors. Cancer Discov 5(8):850–859
Jordan EJ, Kim HR, Arcila ME et al (2017) Prospective comprehensive molecular characterization of lung adenocarcinomas for efficient patient matching to approved and emerging therapies. Cancer Discov 7(6):596–609
Krumm N, Sudmant PH, Ko A et al (2012) Copy number variation detection and genotyping from exome sequence data. Genome Res 22(8):1525–1532
Levin PA, Brekken RA, Byers LA et al (2016) Axl receptor axis: a new therapeutic target in lung cancer. J Thorac Oncol 11(8):1357–1362
Li H, Durbin R (2010) Fast and accurate long-read alignment with Burrows–Wheeler transform. Bioinformatics 26(5):589–595
Pritchard CC, Salipante SJ, Koehler K et al (2014) Validation and implementation of targeted capture and sequencing for the detection of actionable mutation, copy number variation, and gene rearrangement in clinical cancer specimens. J Mol Diagn 16(1):56–67
Rausch T, Zichner T, Schlattl A et al (2012) DELLY: structural variant discovery by integrated paired-end and split-read analysis. Bioinformatics 28(18):i333–i339
Robinson JT, Thorvaldsdottir H, Winckler W et al (2011) Integr Genom View. Nat Biotechnol 29(1):24–26
Seo JS, Ju YS, Lee WC et al (2012) The transcriptional landscape and mutational profile of lung adenocarcinoma. Genome Res 22(11):2109–2119
Wang K, Li M, Hakonarson H (2010) ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res 38(16):e164
Funding
This research was supported by grants from the Korea Health Technology R&D Project through the Korea Health Industry Development Institute (KHIDI), funded by the Ministry of Health and Welfare, Republic of Korea (Grant numbers HI14C1277 and HI13C2148).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
Nak-Jung Kwon is an employee of Macrogen Inc. Sun-Wha Im, Jeesoo Chae, Se Song Jang, Jaeyong Choi, Jihui Yun, Soojin Cha, Yoon Kyung Jeon, Yoohwa Hwang, Miso Kim, Tae Min Kim, Dong-Wan Kim, Jong-Il Kim and Young Tae Kim declare that they have no conflict of interest.
Ethical approval
This study had been approved by the Seoul National University Hospital (No. 1012-130-346 and 1105-075-362). Informed consent was obtained from all individual participants included in the study.
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Im, SW., Chae, J., Jang, S.S. et al. A newly developed capture-based sequencing panel for genomic assay of lung cancer. Genes Genom 42, 751–759 (2020). https://doi.org/10.1007/s13258-020-00949-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13258-020-00949-1