Abstract
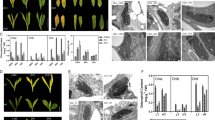
The spikelet is a unique structure of grass plants, and its development involved with complicated molecular regulation network. nsg (nonstop glumes) mutant affecting spikelet development was identified from EMS-treated Jinhui10 (Oryza sativa L. ssp. indica.). Mutant plants had normal glumes (inner rudimentary glume, empty glumes and lemma/palea) and pedicel at the early flowering stage, but had longer ones at later stage. An extra glume-like organ was found in 84 % of mutant individuals. The number of stamens decreased in most mutant individuals whereas three stigmas or two carpels were found in some mutant individuals. The mutant phenotype suggests that NSG is involved in the whole rice spikelet development. NSG was mapped to a 15 kb region on the chromosome 4. According to sequence analysis, a gene encoding a protein with C2H2 domain exhibited a 13 bp insertion, causing a frame shift in genomics DNA and cDNA in nsg. This gene was identified as the candidated gene of NSG. The mutation of NSG influenced the transcription level of some floral hometic genes. The expression of OsMADS4, OsMADS16, DL and OsMADS3 decreased distinctly, and OsMADS1 increased in nsg panicle, suggests that NSG affected spikelet development through influencing the expression of floral hometic genes.




Similar content being viewed by others

Abbreviations
- ORG:
-
Outer rudimentary glume
- IRG:
-
Inner rudimentary glume
- OEG:
-
Outer empty glume
- IEG:
-
Inner empty glume
References
Abe M, Kobayashi Y, Yamamoto S, Daimon Y, Yamaguchi A, Ikeda Y, Ichinoki H, Notaguchi M, Goto K, Araki T (2005) FD, a bZIP protein mediating signals from the floral pathway integrator FT at the shoot apex. Science 309:1052–1056
Agrawal KG, Abe K, Yamazaki M, Miyao A, Hirochika A (2005) Conservation of the E-function for floral organ identity in rice revealed by the analysis of tissue culture-induced loss of function mutants of the OsMADS1 gene. Plant Mol Biol 59:125–135
Bradley D, Ratcliffe O, Vincent C, Carpenter R, Coen E (1997) Inflorescence commitment and architecture in Arabidopsis. Science 275:80–83
Chung YY, Kim SR, Kang HG, Noh YS, Park MC, Finkel D, An G (1995) Characterization of two rice MADS-box genes homologous to GLOBOSA. Plant Sci 109:45–56
Coen ES, Meyerowitz EM (1991) The war of the whorls: genetic interactions controlling flower development. Nature 353:31–37
Fornara F, Parenicova L, Falasca G, Pelucchi N, Masiero S, Ciannamea S, Lopez-Dee Z, Altamura MM, Colombo L, Kater MM (2004) Functional characterization of OsMADS18, a member of the AP1/SQUA subfamily of MADS-box genes. Plant Physiol 135:2207–2219
Gao X, Liang W, Yin C, Ji S, Wang H, Su X, Guo C, Kong H, Xue H, Zhang D (2010) The SEPALLATA-like gene OsMADS34 is required for rice inflorescence and spikelet development. Plant Physiol 153:728–740
Greco R, Stagi L, Colombo L, Agenent GC, Sari-Gorla M, Pè ME (1997) MADS-box genes expressed in developing inflorescences of rice and sorghum. Mol Gen Genet 253:615–623
Huang T, Bohlenius H, Eriksson S, Parcy F, Nilsson O (2005) The mRNA of the Arabidopsis gene FT moves from leaf to shoot apex and induces flowering. Science 309:1694–1696
Jeon JS, Lee S, Jung KH, Yang WS, Yi GH, Oh BG, An GH (2000) Production of transgenic rice plants showing reduced heading date and plant height by ectopic expression of rice MADS-box genes. Mol Breed 6:581–592
Kang HG, An G (1997) Isolation and characterization of a rice MADS-box gene belonging to the AGL2 gene family. Mol Cell 7:45–51
Kater MM, Dreni L, Colombo L (2006) Functional conservation of MADS-box factors controlling floral organ identity in rice and Arabidopsis. J Exp Bot 57:3433–3444
Kobayashi Y, Kaya H, Goto K, Iwabuchi M, Araki T (1999) A pair of related genes with antagonistic roles in mediating flowering signals. Science 286:1960–1962
Kobayashi K, Maekawa M, Miyao A, Hirochika H, Kyozuka J (2010) PANICLE PHYTOMER2 (PAP2), encoding a SEPALLATA subfamily MADS-box protein, positively controls spikelet meristem identity in rice. Plant Cell Physiol 51:47–57
Komatsu M, Chujo A, Nagato Y, Shimamoto K, Kyozuka J (2003) FRIZZY PANICLE is required to prevent the formation of axillary meristems and to establish floral meristem identity in rice spikelets. Development 130:3841–3850
Kyozuka J, Kobayashi T, Morita M, Shimamoto K (2000) Spatially and temporally regulated expression of rice MADS-box genes with similarity to Arabidopsis class A, B and C genes. Plant Cell Physiol 41:710–718
Lee DY, Lee J, Moon S, Park SY, An G (2007) The rice heterochronic gene SUPERNUMERARY BRACT regulates the transition from spikelet meristem to floral meristem. Plant J 49:64–78
Li H, Liang W, Yin C, Zhu L, Zhang D (2011) Genetic interaction of OsMADS3, DROOPING LEAF, and OsMADS13 in specifying rice floral organ identities and meristem determinacy. Plant Physiol 156:263–274
Liljegren SJ, Gustafson-Brown C, Pinyopich A, Ditta GS, Yanofsky MF (1999) Interactions among APETALA1, LEAFY, and TERMINAL FLOWER1 specify meristem fate. Plant Cell 11:1007–1018
Mandel MA, Yanofsky MF (1995) A gene triggering flower formation in Arabidopsis. Nature 377:522–524
McSteen P, Laudencia-Chingcuanco D, Colasanti J (2000) A floret by any other name: control of meristem identity in maize. Trends Plant Sci 5:61–66
Nagasawa N, Miyoshi M, Sano Y, Satoh H, Hirano HY, Sakai H, Nagato Y (2003) SUPERWOMAN1 and DROOPING LEAF genes control floral organ identity in rice. Development 130:705–718
Pelucchi N, Fornara F, Favalli C, Masiero S, Lago C, Pe ME, Colombo L, Kater MM (2002) Comparative analysis of rice MADS-box genes expressed during flower development. Sex Plant Reprod 15:113–122
Poethig RS (2003) Phase change and the regulation of developmental timing in plants. Science 301:334–336
Prasad K, Vijayraghavan U (2003) Double-stranded RNA interference of a rice PI/GLO paralog OsMADS2, uncovers its second- whorl-specific function in floral organ patterning. Genetics 165:2301–2305
Prasad K, Parameswaran S, Vijayraghavan U (2005) OsMADS1, a rice MADS-box factor, controls differentiation of specific cell types in the lemma and palea and is an early acting regulator of inner floral organs. Plant J 43:915–928
Ruiz-Garcia L, Madueno F, Wilkinson M, Haughn G, Salinas J, Martinez-Zapater JM (1997) Different roles of flowering-time genes in the activation of floral initiation genes in Arabidopsis. Plant Cell 9:1921–1934
Sablowski R (2007) Flowering and determinacy in Arabidopsis. J Exp Bot 58:899–907
Schmidt RJ, Ambrose BA (1998) The blooming of grass flower development. Curr Opin Plant Biol 1:60–67
Schultz EA, Haughn GW (1991) LEAFY, a homeotic gene that regulates inflorescence development in Arabidopsis. Plant Cell 3:771–781
TheiBen G, Saedler H (2001) Plant biology. Floral quartets. Nature 409:469–471
Wagner D, Sablowski RW, Meyerowitz EM (1999) Transcriptional activation of APETALA1 by LEAFY. Science 285:582–584
Wang K, Tang D, Hong L, Xu W, Huang J, Li M, Gu M, Xue Y, Cheng Z (2010) DEP and AFO regulate reproductive habit in rice. PLoS Genet 6:e1000818
Weigel D, Nilsson O (1995) A developmental switch sufficient for flower initiation in diverse plants. Nature 377:495–500
Weigel D, Alvarez J, Smyth DR, Yanofsky MF, Meyerowitz EM (1992) LEAFY controls floral meristem identity in Arabidopsis. Cell 69:843–859
Yamaguchi T, Hirano HY (2006) Function and diversification of MADS-box genes in rice. Sci World J 6:1923–1932
Yamaguchi T, Nagasawa N, Kawasaki S, Matsuoka M, Nagato Y, Hirano HY (2004) The YABBY gene DROOPING LEAF regulates carpel specification and midrib development in rice. Plant Cell 16:500–509
Yamaguchi T, Lee DY, Miyao A, Hirochika H, An G, Hirano HY (2006) Functional diversification of the two C-class genes OsMADS3 and OsMADS58 in Oryza Sativa. Plant Cell 18:15–18
Acknowledgments
This research was supported by funds from the National Natural Science Foundation of China (31071390) and the Excellent Youth Foundation Project of Chongqing (2008BA1033).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wang, N., Li, Y., Sang, X. et al. nonstop glumes (nsg), a novel mutant affects spikelet development in rice. Genes Genom 35, 149–157 (2013). https://doi.org/10.1007/s13258-013-0067-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13258-013-0067-7