Abstract
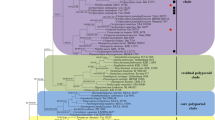
Fragiliporiaceae fam. nov., a new poroid wood-inhabiting family, is introduced based on the combination of molecular and morphological data, and is typified by Fragiliporia gen. nov. The phylogenetic analysis shows that Fragiliporia fragilis sp. nov. forms a monophyletic group within Polyporales and warrants the introduction of a new fragiliporia clade based on molecular data of ITS + nLSU rRNA gene regions. Combined ITS, nLSU, mtSSU, TEF1 and RPB2 sequence data also demonstrated that the new family Fragiliporiaceae also formed a monophyletic lineage (70 % BS, 57 % BP, 0.99 BPP), and grouped with the phlebioid clade, residual polyporoid clade and core polyporoid clade. Fragiliporiaceae has unique macromorphological characters in having resupinate basidiocarps with very soft tubes when fresh, which become brittle when dry (becoming almost powdery when bruised); a monomitic hyphal system with thick-walled generative hyphae, clamp connections, and frequently H-, W- or Y-shaped hyphae branching from the clamp connections.




Similar content being viewed by others
References
Binder M, Hibbett DS, Larsson KH, Larsson E, Langer E, Langer G (2005) The phylogenetic distribution of resupinate forms across the major clades of mushroom-forming fungi (Homobasidiomycetes). Syst Biodivers 3:113–157. doi:10.1017/S1477200005001623
Binder M, Justo A, Riley R, Salamov A, López-Giráldez F, Sjökvist E, Copeland A, Foster B, Sun H, Larsson E, Larsson KH, Townsend J, Grigoriev IV, Hibbett DS (2013) Phylogenetic and phylogenomic overview of the Polyporales. Mycologia 105:1350–1373. doi:10.3852/13-003
Cao Y, Wu SH, Dai YC (2012) Species clarification of the prize medicinal Ganoderma mushroom “Lingzhi”. Fungal Divers 56:49–62. doi:10.1007/s13225-012-0178-5
Cui BK, Zhao CL, Dai YC (2011) Melanoderma microcarpum gen. et sp. nov. (Basidiomycota) from China. Mycotaxon 116:295–302. doi:10.5248/116.295
Dai YC (2010) Hymenochaetaceae (Basidiomycota) in China. Fungal Divers 45:131–343. doi:10.1007/s13225-010-0066-9
Dai YC (2012) Polypore diversity in China with an annotated checklist of Chinese polypores. Mycoscience 53:49–80. doi:10.1007/s10267-011-0134-3
Dai YC, Cui BK, Yuan HS, Li BD (2007) Pathogenic wood-decaying fungi in China. Forest Pathol 37:105–120. doi:10.1111/j.1439-0329.2007.00485.x
Dai YC, Yang ZL, Cui BK, Yu CJ, Zhou LW (2009) Species diversity and utilization of medicinal mushrooms and fungi in China (Review). Int J Med Mushrooms 11:287–302. doi:10.1615/IntJMedMushr.v11.i3.80
Dai YC, Xue HJ, Vlasák J, Rajchenberg M, Wang B, Zhou LW (2014) Phylogeny and global diversity of Polyporus group Melanopus (Polyporales, Basidiomycota). Fungal Divers 64:133–144. doi:10.1007/s13225-013-0248-3
Donk MA (1933) Revision de niederlandischen homobasidiomycetes. Aphyllophoraceae 2. Medd Bot Mus Herb Rijhs Universit Utrecht 9:1–278
Felsenstein J (1985) Confidence intervals on phylogenetics: an approach using bootstrap. Evolution 39:783–791
Floudas D, Binder M, Riley R, Barry K, Blanchette RA, Henrissat B, Martínez AT, Otillar R, Spatafora JW, Yadav JS, Aerts A, Benoit I, Boyd A, Carlson A, Copeland A, Coutinho PM, de Vries RP, Ferreira P, Findley K, Foster B, Gaske J, Glotzer D, Górecki P, Heitman J, Hesse C, Hori C, Igarashi K, Jurgens JA, Kallen N, Kersten P, Kohler A, Kües U, Kumar TK, Kuo A, LaButti K, Larrondo LF, Lindquist E, Ling A, Lombard V, Lucas S, Lundell T, Martin R, McLaughlin DJ, Morgenstern I, Morin E, Murat C, Nagy LG, Nolan M, Ohm RA, Patyshakuliyeva A, Rokas A, Ruiz-Dueñas FJ, Sabat G, Salamov A, Samejima M, Schmutz J, Slot JC, St John F, Stenlid J, Sun H, Sun S, Syed K, Tsang A, Wiebenga A, Young D, Pisabarro A, Eastwood DC, Martin F, Cullen D, Grigoriev IV, Hibbett DS (2012) The Paleozoic origin of enzymatic lignin decomposition reconstructed from 31 fungal genomes. Science 336:1715–1719. doi:10.1126/science.1221748
Garcia-Sandoval R, Wang Z, Binder M, Hibbett DS (2011) Molecular phylogenetics of the Gloeophyllales and relative ages of clades of Agaricomycotina producing a brown rot. Mycologia 103:510–524. doi:10.3852/10-209
Gilbertson RL, Ryvarden L (1986) North American polypores 1. Fungiflora, Oslo
Ginns J (1984) Griseoporia a new genus for Hexagonia carbonaria (Polyporaceae). Mycotaxon 20:559–565
Hall TA (1999) Bioedit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98
He SH, Dai YC (2012) Taxonomy and phylogeny of Hymenochaete and allied genera of Hymenochaetaceae (Basidiomycota) in China. Fungal Divers 56:77–93. doi:10.1007/s13225-012-0174-9
Hibbett DS, Binder M (2002) Evolution of complex fruiting body morphologies in homobasidiomycetes. Proc Biol Sci 269:1963–1969. doi:10.1098/rspb.2002.2123
Hibbett DS, Binder M, Bischoff JF, Blackwell M, Cannon PF, Eriksson OE, Huhndorf S, James T, Kirk PM, Lücking R, Thorsten Lumbsch H, Lutzoni F, Matheny PB, McLaughlin DJ, Powell MJ, Redhead S, Schoch CL, Spatafora JW, Stalpers JA, Vilgalys R, Aime MC, Aptroot A, Bauer R, Begerow D, Benny GL, Castlebury LA, Crous PW, Dai YC, Gams W, Geiser DM, Griffith GW, Gueidan C, Hawksworth DL, Hestmark G, Hosaka K, Humber RA, Hyde KD, Ironside JE, Kõljalg U, Kurtzman CP, Larsson KH, Lichtwardt R, Longcore J, Miadlikowska J, Miller A, Moncalvo JM, Mozley-Standridge S, Oberwinkler F, Parmasto E, Reeb V, Rogers JD, Roux C, Ryvarden L, Sampaio JP, Schüssler A, Sugiyama J, Thorn RG, Tibell L, Untereiner WA, Walker C, Wang Z, Weir A, Weiss M, White MM, Winka K, Yao YJ, Zhang N (2007) A higher-level phylogenetic classification of the Fungi. Mycol Res 111:509–547. doi:10.1016/j.mycres.2007.03.004
Hillis DM, Bull JJ (1993) An empirical test of bootstrapping as a method for assessing confidence in phylogenetic analysis. Syst Biol 42:182–192. doi:10.1093/sysbio/42.2.182
Hjortstam K, Tellería MT (1990) Columnocystis, a synonym of Veluticeps. Mycotaxon 37:53–56
Hyde KD, Nilsson RH, Alias SA, Ariyawansa HA, Blair JE, Cai L, de Cock AWAM, Dissanayake AJ, Glockling SL, Goonasekara ID, Gorczak M, Hahn M, Jayawardena RS, van Kan IAL, Laurence MH, Lévesque CA, Li XH, Liu JK, Maharachchikumbura SSN, Manamgoda DS, Martin FN, McKenzie EHC, McTaggart AR, Mortimer PE, Nair PVR, Pawłowska J, Rintoul TL, Shivas TG, Spies ARCFJ, Summerell BA, Taylor PWJ, Terhem RB, Udayanga D, Vaghefi N, Walther G, Wilk M, Wrzosek M, Xu JC, Yan JY, Zhou N (2014) One stop shop: backbones trees for important phytopathogenic genera. Fungal Diversity 68 (in press)
Hyde KD, Jones EBG, Liu JK, Ariyawansha H, Boehm E, Boonmee S, Braun U, Chomnunti P, Crous P, Dai DQ, Diederich P, Dissanayake A, Doilom M, Doveri F, Hongsanan S, Jayawardena R, Lawrey JD, Li YM, Liu YX, Lücking R, Monkai J, Nelsen MP, Phookamsak R, Muggia L, Pang KL, Senanayake I, Shearer CA, Wijayawardene N, Wu HX, Thambugala M, Suetrong S, Tanaka K, Wikee S, Zhang Y, Hudson BA, Alias SA, Aptroot A, Bahkali AH, Bezerra LJ, Bhat JD, Camporesi E, Chukeatirote E, Hoog SD, Gueidan C, Hawksworth DL, Hirayama K, Kang JC, Knudsen K, Li WJ, Liu ZY, McKenzie EHC, Miller AN, Nadeeshan D, Phillip AJL, Mapook A, Raja HA, Tian Q, Zhang M, Scheuer C, Schumm F, Taylor J, Yacharoen S, Tibpromma S, Wang Y, Yan J, Li X (2013) Families of Dothideomycetes. Fungal Divers 63:1–313. doi:10.1007/s13225-013-0263-4
James TY, Kauff F, Schoch CL, Matheny PB, Hofstetter V, Cox CJ, Celio G, Gueidan C, Fraker E, Miadlikowska J, Lumbsch HT, Rauhut A, Reeb V, Arnold AE, Amtoft A, Stajich JE, Hosaka K, Sung GH, Johnson D, O'Rourke B, Crockett M, Binder M, Curtis JM, Slot JC, Wang Z, Wilson AW, Schüssler A, Longcore JE, O'Donnell K, Mozley-Standridge S, Porter D, Letcher PM, Powell MJ, Taylor JW, White MM, Griffith GW, Davies DR, Humber RA, Morton JB, Sugiyama J, Rossman AY, Rogers JD, Pfister DH, Hewitt D, Hansen K, Hambleton S, Shoemaker RA, Kohlmeyer J, Volkmann-Kohlmeyer B, Spotts RA, Serdani M, Crous PW, Hughes KW, Matsuura K, Langer E, Langer G, Untereiner WA, Lücking R, Büdel B, Geiser DM, Aptroot A, Diederich P, Schmitt I, Schultz M, Yahr R, Hibbett DS, Lutzoni F, McLaughlin DJ, Spatafora JW, Vilgalys R (2006) Reconstructing the early evolution of fungi using a six-gene phylogeny. Nature 443:818–822. doi:10.1038/nature05110
Jia BS, Zhou LW, Cui BK, Rivoire B, Dai YC (2014) Taxonomy and phylogeny of Ceriporia (Polyporales, Basidiomycota) with an emphasis of Chinese collections. Mycol Prog 13:81–93. doi:10.1007/s11557-013-0895-5
Jülich W (1981) Higher taxa of Basidiomycetes. Biblthca Mycol 85:1–485
Justo A, Hibbett DS (2011) Phylogenetic classification of Trametes (Basidiomycota, Polyporales) based on a five-marker dataset. Taxon 60:1567–1583
Kim SY, Jung HS (2000) Phylogenetic relationships of the Aphyllophorales inferred from sequence analysis of nuclear small subunit ribosomal DNA. J Microbiol 38:122–131
Kim KM, Lee JS, Jung HS (2007) Fomitopsis incarnatus sp. nov. based on generic evaluation of Fomitopsis and Rhodofomes. Mycologia 99:833–841. doi:10.3852/mycologia.99.6.833
Kirk PM, Cannon PF, David JC, Minter DW, Stalpers JA (2008) Ainsworth and bisby's dictionary of the fungi, 10th edn. CAB International Press, Wallingford
Li HJ, Cui BK (2013) Taxonomy and phylogeny of the genus Megasporoporia and its related genera. Mycologia 105:368–383. doi:10.3852/12-114
Liu YL, Whelen S, Hall BD (1999) Phylogenetic relationships among ascomycetes: evidence from an RNA polymerase II subunit. Mol Biol Evol 16:1799–1808
Matheny PB (2005) Improving phylogenetic inference of mushrooms with RPB1 and RPB2 nucleotide sequences (Inocybe; Agaricales). Mol Phylogenet Evol 35:1–20. doi:10.1016/j.ympev.2004.11.014
Miettinen O, Rajchenberg M (2012) Obba and Sebipora, new polypore genera related to Cinereomyces and Gelatoporia (Polyporales, Basidiomycota). Mycol Prog 11:131–147. doi:10.1007/s11557-010-0736-8
Miettinen O, Larsson E, Sjökvist E, Larsson KL (2012) Comprehensive taxon sampling reveals unaccounted diversity and morphological plasticity in a group of dimitic polypores (Polyporales, Basidiomycota). Cladistics 28:251–270. doi:10.1111/j.1096-0031.2011.00380.x
Nakasone KK (1990) Taxonomic study of Veluticeps (Aphyllophorales). Mycologia 82:622–641
Nilsson RH, Hyde KD, Pawłowska J, Ryberg M, Tedersoo L, Aas AB, Alias SA, Alves A, Anderson CL, Antonelli A, Arnold AE, Bahnmann B, Bahram M, Bengtsson-Palme J, Berlin A, Branco S, Chomnunti P, Dissanayake A, Drenkhan R, Friberg H, Frøslev TG, Halwachs B, Hartmann M, Henricot B, Jayawardena R, Jumpponen A, Kauserud H, Koskela S, Kulik T, Liimatainen K, Lindahl BD, Lindner D, Liu J-K, Maharachchikumbura S, Manamgoda D, Martinsson S, Neves MA, Niskanen T, Nylinder S, Pereira OL, Pinho DB, Porter TM, Queloz V, Riit T, Sánchez-García M, Sousa F, Stefańczyk E, Tadych M, Takamatsu S, Tian Q, Udayanga D, Unterseher M, Wang Z, Wikee S, Yan J, Larsson E, Larsson K-H, Kõljalg U, Abarenkov K (2014) Improving ITS sequence data for identification of plant pathogenic fungi. Fungal Divers. doi:10.1007/s13225-014-0291-8, in press
Núñez M, Ryvarden L (2001) East Asian polypores 2. Synopsis Fungorum 14:165–522
Nylander JAA (2004) MrModeltest v2. Program distributed by the author. Evolutionary Biology Centre, Uppsala University
Petersen JH (1996) Farvekort. The Danish Mycological Society’s colour-chart. Foreningen til Svampekundskabens Fremme, Greve
Posada D, Crandall KA (1998) Modeltest: testing the model of DNA substitution. Bioinformatics 14:817–818
Rehner SA, Buckley E (2005) A Beauveria phylogeny inferred from nuclear ITS and EF1-alpha sequences: evidence for cryptic diversification and links to Cordyceps teleomorphs. Mycologia 97:84–98. doi:10.3852/mycologia.97.1.84
Robledo GL, Amalfi M, Castillo G, Rajchenberg M, Decock C (2009) Perenniporiella chaquenia sp. nov. and further notes on Perenniporiella and its relationships with Perenniporia (Poriales, Basidiomycota). Mycologia 101:657–673. doi:10.3852/08-040
Ronquist F, Huelsenbeck JP (2003) MrBayes 3: bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572–1574. doi:10.1093/bioinformatics/btg180
Rune F (1994) Neolentinus: a well founded genus in Pleurotaceae that includes Heliocybe. Mycol Res 98:542–544. doi:10.1016/S0953-7562(09)80476-0
Ryvarden L, Gilbertson RL (1993) European polypores 1. Synopsis Fungorum 6:1–387
Si J, Peng F, Cui BK (2013) Purification, biochemical characterization and dye decolorization capacity of an alkali-resistant and metal-tolerant laccase from Trametes pubescens. Bioresour Technol 128:49–57. doi:10.1016/j.biortech.2012.10.085
Sjökvist E, Larsson E, Eberhardt U, Ryvarden L, Larsson KH (2012) Stipitate stereoid basidiocarps have evolved multiple. Mycologia 104:1046–1055. doi:10.3852/11-174
Sotome K, Hattori T, Ota Y, To-anun C, Salleh B, Kakishima M (2008) Phylogenetic relationships of Polyporus and morphologically allied genera. Mycologia 100:603–615. doi:10.3852/07-191R
Stamatakis A (2006) RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 22:2688–2690. doi:10.1093/bioinformatics/btl446
Suhara H, Maekawa N, Kaneko S, Hattori T, Sakai K, Kondo R (2003) A new species, Ceriporia lacerata, isolated from white-rotted wood. Mycotaxon 86:335–347
Swofford DL (2002) PAUP*: phylogenetic analysis using parsimony (*and other methods). Version 4.0b10. Sinauer Associates, Massachusetts
Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The CLUSTAL X windows interface: flexible strategies formultiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 25:4876–4882. doi:10.1093/nar/25.24.4876
Tian XM, Yu HY, Zhou LW, Decock C, Vlasák J, Dai YC (2013) Phylogeny and taxonomy of the Inonotus linteus complex. Fungal Divers 58:159–169. doi:10.1007/s13225-012-0202-9
Tomšovský M, Menkis A, Vasaitis R (2010) Phylogenetic relationships in European Ceriporiopsis species inferred from nuclear and mitochondrial ribosomal DNA sequences. Fungal Biol 114:350–358. doi:10.1016/j.funbio.2010.02.004
Wang W, Yuan TQ, Wang K, Cui BK, Dai YC (2012) Combination of biological pretreatment with liquid hot water pretreatment to enhance enzymatic hydrolysis of Populus tomentosa. Bioresour Technol 107:282–286. doi:10.1016/j.biortech.2011.12.116
White TJ, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ (eds) PCR protocols: a guide to methods and applications. Academic, San Diego, pp 315–322
Wu SH, Nilsson HR, Chen CT, Yu SY, Hallenberg N (2010) The white-rotting genus Phanerochaete is polyphyletic and distributed throughout the phleboid clade of the Polyporales (Basidiomycota). Fungal Divers 42:107–188. doi:10.1007/s13225-010-0031-7
Wu G, Feng B, Xu JP, Zhu XT, Li YC, Zeng NK, Hosen Md I, Yang ZL (2014) Molecular phylogenetic analyses redefine seven major clades and reveal 22 new generic clades in the fungal family Boletaceae. Fungal Divers. doi:10.1007/s13225-014-0283-8, in press
Younes SB, Mechichi T, Sayadi S (2007) Purification and characterization of the laccase secreted by the white rot fungus Perenniporia tephropora and its role in the decolourization of synthetic dyes. J Appl Microbiol 102:1033–1042
Zhang Y, Crous PW, Schoch CL, Bahkali AH, Guo LD, Hyde KD (2011) A molecular, morphological and ecological re-appraisal of Venturiales―a new order of Dothideomycetes. Fungal Divers 51:249–277. doi:10.1007/s13225-011-0141-x
Zhao CL, Cui BK (2014) Phylogeny and taxonomy of Ceriporiopsis (Polyporales) with descriptions of two new species from southern China. Phytotaxa 164:17–28. doi:10.11646/phytotaxa.164.1.2
Zhao CL, Cui BK, Dai YC (2013) New species and phylogeny of Perenniporia based on morphological and molecular characters. Fungal Divers 58:47–60. doi:10.1007/s13225-012-0177-6
Zhao CL, He XS, WangHe KY, Cui BK, Dai YC (2014) Flammeopellis bambusicola gen. et. sp. nov. (Polyporales, Basidiomycota) evidenced by morphological characters and phylogenetic analysis. Mycol Prog 13:771–780. doi:10.1007/s11557-014-0960-8
Zhou LW, Hao ZQ, Wang Z, Wang B, Dai YC (2011) Comparison of ecological patterns of polypores in three forest zones in China. Mycology 2:260–275
Acknowledgments
The research is supported by the Fundamental Research Funds for the Central Universities (Project No. JC2013-1) and Beijing Higher Education Young Elite Teacher Project (YETP0774).
Author information
Authors and Affiliations
Corresponding author
Additional information
Chang-Lin Zhao and Bao-Kai Cui contributed equally to this work and shared the first author
Rights and permissions
About this article
Cite this article
Zhao, CL., Cui, BK., Song, J. et al. Fragiliporiaceae, a new family of Polyporales (Basidiomycota). Fungal Diversity 70, 115–126 (2015). https://doi.org/10.1007/s13225-014-0299-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13225-014-0299-0