Abstract
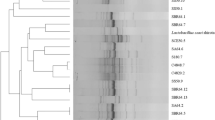
Italian ryegrass (IRG), barley, and rye are types of forage that are difficult to ensile with an assurance of good quality. Therefore, the addition of lactic acid bacteria (LAB) is the best way to enhance the preservation of this silage. However, applications of LAB have been impeded due to its poor growth characteristics, the sudden decline in pH, and other non-beneficial microbial growth associated with its presence. To overcome these limitations, a new Lactobacillus sp. KCC-10 and KCC-19 strain was isolated from well-fermented IRG silage samples. Biochemical and physiological studies revealed that the strains were Gram-positive, catalase-negative, and produced gas from glucose, and produced more lactic acid in fermentation. The 16S rRNA gene-based phylogenetic affiliation was determined by using bioinformatic tools that identified Lactobacillus sp. KCC-10 and KCC-19 with 100 % sequence similarity to Lactobacillus plantarum. Novel L. plantarum strains were deposited in the Korean Collection for Type Cultures under the accession numbers KACC 91785P and KACC 91758P, respectively. The shake-flask cultivation of these new strains under aerobic, microaerobic, and anaerobic conditions showed a higher specific growth rate than that achieved using the well-studied L. plantarum KACC 91016 and KACC 91096 on MRS broth and grass juice. Lactic acid was detected as the dominant organic acid in IRG (78.45 mM), barley (51.28 mM), corn (16.28 mM), and rice paddy (11.05 mM), followed by acetic acid and succinic acid. The KCC-10 in the silage was observed to increase from 2.4 × 105 CFU/g per sample at day 0 to 0.58, 0.60, and 0.59 × 109 CFU/g at day 5 for IRG, barley, and rye, respectively. The growth of KCC-10 and KCC-19 in all the silages decreased, as the storage period increased from 5 to 50 days. Whereas, KCC-19 was noted to increase from 2.7 × 105 CFU/g per sample at day 0 to 0.71, 0.72, and 0.711 × 109 CFU/g at day 5 for IRG, barley, and rye. Among the total organic acids, lactic acid was detected as the dominant acid present in IRG, barley, and rye silages. From these results, we concluded that strains KCC-10 and KCC-19 can be used as appropriate inoculants to prolong the stability of silage and fermentation quality.






Similar content being viewed by others
References
Arasu VM, Jung MW, Ilavenil S, Jane M, Kim DH, Lee KD, Park HS, Hur TY, Choi GJ, Lim YC, Al-Dhabi NA, Choi KC (2013) Isolation and characterization of antifungal compound from Lactobacillus plantarum KCC-10 from forage silage with potential beneficial properties. J Appl Microbiol 115(5):1172–1185
Ashbell G, Ashbell G, Lisker N (1988) Aerobic deterioration in maize silage stored in a bunker silos under farm conditions in a subtropical climate. J Sci Food Agric 45:307–315
Avila CLS, Valeriano AR, Pinto JC, Figueiredo AVR, Schwan RF (2010) Chemical and microbiological characteristics of sugar cane silages treated with microbial inoculants. Rev Bras Zootec 39:25–32
Bach SJ, McAllister TA, Baah J, Yanke LJ, Veira DM, Gannon VPJ, Holley RA (2002) Persistence of Escherichia coli O157: H7 in barley silage: effect of a bacterial inoculant. J Appl Microbiol 93:288–294
Barnett AJG (1954) Silage fermentation. Butterworth Scientific Publication, London, pp 111–123
Cai Y, Ohmomo S, Ogawa M, Kumai S (1999) Effect of applying lactic acid bacteria isolated from forage crops on fermentation characteristics and aerobic deterioration of silage. J Dairy Sci 82:520–526
De Angelis M, de Candia S, Calasso MP, Faccia M, Guinee TP, Simonetti MC, Gobbetti M (2008) Selection and use of autochthonous multiple strain cultures for the manufacture of high moisture traditional Mozzarella cheese. Int J Food Microbiol 125:123–132
Eitan BD, Shapiro OH, Siboni N, Kushmaro A (2006) Advantage of using inosine at the 3_ termini of 16S rRNA gene universal primers for the study of microbial diversity. Appl Environ Microbiol 72:6902–6906
Ennahar S, Cai Y, Yasuhito F (2003) Phylogenetic diversity of lactic acid bacteria associated with paddy rice silage as determined by 16S ribosomal DNA analysis. Appl Environ Microbiol 69:444–451
Georgieva R, Iliev I, Chipeva VA, Dimitonova SP, Samelis J, Danova S (2008) Identification and vitro characterization of Lactobacillus plantarum strains from artisanal Bulgarian white brined cheeses. J Basic Microb 48:234–244
Giraffa G, Chanishvili N, Widyastuti Y (2010) Importance of lactobacilli in food and feed biotechnology. Res Microbiol 161:480–487
Lin C, Bolsen KK, Brent BE, Hart RA, Dickerson JT, Feyerherm AM, Aimutis WR (1991) Epiphytic microflora on alfalfa and whole-plant corn. J Dairy Sci 75:2484–2493
Lin C, Bolsen KK, Brent BE, Fung DYC (1992) Epiphytic lactic acid bacteria succession during the pre-ensiling and ensiling periods of alfalfa and maize. J Appl Bacteriol 73:375–387
McAllister TA, Feniuk R, Mir Z, Mir P, Selinger LB, Cheng K-J (1998) Inoculants for alfalfa silage: effects on aerobic stability, digestibility and the growth performance of feedlot steers. Livest Prod Sci 53:171–181
McDonald P, Henderson AR, Heron SJE (1991) The biochemistry of silage, 2nd edn. Chalcombe Publications, Aberystwyth, pp 20–102
Nakui T, Masaki S, Aihara T, Yahara N, Takai S (1988) The making of rice whole crop silage and an evaluation of its value as forage for ruminants. Bull Tohoku Natl Agric Exp Stn 78:173–180
Rodrigues PHM, de Andrade SJT, de Almeida LFS, Meyer PM, de Lima FR, Lucci CD (2001) Microbial inoculation of alfalfa for ensiling on apparent digestibility in wethers. Revista Brasileira Zootecnia-Brazilian. J Anim Sci 30:1925–1930
Sebastian S, Phillip LE, Fellner V, Idziak ES (1996) Comparative assessment of bacterial inoculated corn and sorghum silages. J Anim Sci 71:505–514
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28(10):2731–2739
Valan Arasu M, Ilavenil S, Kim DH, Lee JC, Choi KC (2014). In vitro and in vivo enhancement of adipogenesis by Italian Ryegrass (Lolium Multiflorum) in 3T3-L1 cells and mice. PLOS ONE. doi:10.1371/journal.pone.0085297
Weinberg ZG, Muck RE (1996) New trends and opportunities in the development and use of inoculants for silage. FEMS Microbiol Rev 19:53–68
Weinberg ZG, Muck RE, Weimer PJ (2003) The survival of silage inoculant lactic acid bacteria in rumen fluid. J Appl Microbiol 94:1066–1071
Winters AL, Fychan R, Jones R (2001) Effect of formic acid and a bacterial inoculant on the amino acid composition of grass silage and on animal performance. Grass Forage Sci 56:181–192
Woolford MK (1975) Microbiological screening of the straight chain fatty acids (C1-C12) as potential silage additives. J Sci Food Agric 26:219–228
Acknowledgement
This work was carried out with the support of the “Cooperative Research Program for Agriculture Science & Technology Development (Project No. PJ008502)”, Rural Development Administration, Republic of Korea.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(DOCX 25 kb)
Rights and permissions
About this article
Cite this article
Valan Arasu, M., Jung, M.W., Kim, D.H. et al. Identification and phylogenetic characterization of novel Lactobacillus plantarum species and their metabolite profiles in grass silage. Ann Microbiol 65, 15–25 (2015). https://doi.org/10.1007/s13213-014-0830-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13213-014-0830-2