Abstract
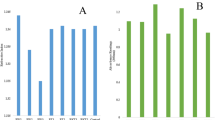
The purpose of this research was to isolate microorganisms from coffee fermentation processes and screen them for their potential to improve the flavor of Arabica coffee using a new approach that included pectin degradation ability and growth in mucilage broth. All of the studied microorganisms were isolated from 38 different samples of fresh coffee cherries, coffee mucilage and coffee pulp. A total of 262 microbial isolates were obtained and subjected to screening using pectinase screening agar medium for pectinolytic organisms. The results of the pectinase production test showed that 18 yeast isolates were found to produce pectinase that could degrade the pectin present in solid media. The sugar assimilation profiles and growth of selected strains in mucilage broth were studied. Therefore, 18 isolates from the selected yeasts were subjected to molecular identification by the use of 18S rRNA gene sequencing. The diversity of the yeast isolates was studied, and they were identified as Wickerhamomyces anomalus, Naganishia liquefaciens, Pichia kudriavzevii, Kazachstania naganishii and Kazachstania sp. Moreover, isolates SWU3YWP1-3, SWU3YSK9 and INFCY1-4 were used as a seed culture for Arabica coffee fermentation. The cupping sensory scores of the control (without yeast inoculation) and those inoculated with three isolated yeast strains that were determined by Q-Arabica Graders were 73.75, 84.75, 80.25 and 75.00, respectively. Unique flavors and aromas were detected. This is the first report of screening microorganisms from the Arabica coffee fermentation process by the combination of various properties with success in improving the quality of coffee beverage.




Similar content being viewed by others
References
Agate AD, Bhat JV (1966) Role of pectinolytic yeasts in the degradation of mucilage layer of Coffea robusta cherries. Appl Microbiol 14(2):256–260. https://doi.org/10.1128/am.14.2.256-260.1966
Avallone S, Guyot B, Brillouet JM, Olguin E, Guiraud JP (2001) Microbiological and biochemical study of coffee fermentation. Curr Microbiol 42(4):252–256. https://doi.org/10.1007/s002840110213
Avallone S, Guiraud JP, Guyot B, Olguin E, Brillouet JM (2006) Polysaccharide constituents of coffee-bean mucilage. J Food Sci 65(8):1308–1311. https://doi.org/10.1111/j.1365-2621.2000.tb10602.x
Batista L, Chalfoun DSS, Silva e BatistaSchwan CR (2016) Coffee: types and production. Encycl Food Health. https://doi.org/10.1016/B978-0-12-384947-2.00184-7
Bhumiratana N, Adhikari K, Chambers E (2011) Evolution of sensory aroma attributes from coffee beans to brewed coffee. LWT Food Sci Technol 44(10):2185–2192. https://doi.org/10.1016/j.lwt.2011.07.001
Bressani APP, Martinez SJ, Batista NN, Simão JBP, Dias DR, Schwan RF (2021) Co-inoculation of yeasts starters: A strategy to improve quality of low altitude Arabica coffee. Food Chem 361:130133. https://doi.org/10.1016/j.foodchem.2021.130133
Elhalis H, Cox J, Frank D, Zhao J (2021) Microbiological and biochemical performances of six yeast species as potential starter cultures for wet fermentation of coffee beans. Food Sci Technol 137:110430. https://doi.org/10.1016/j.lwt.2020.110430
Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39:783–791
Gonzalez-Rios O, Suarez-Quiroz ML, Boulanger R, Barel M, Guyot B, Guiraud JP (2007) Impact of ‘“ecological”’ post-harvest processing on coffee aroma: II. Roasted Coffee. J Food Compost Anal 20(3–4):297–307. https://doi.org/10.1016/j.jfca.2006.12.004
Haile M, Kang WH (2019) Isolation, identification, and characterization of pectinolytic yeasts for starter culture in coffee fermentation. Microorganisms 7(10):401. https://doi.org/10.1155/2019/4836709
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for windows 95/98/NT. Nucl Acids Symp Ser 41:95–98
Jackels SC, Jackels CF (2005) Characterization of the coffee mucilage fermentation process using chemical indicators: a field study in Nicaragua. J Food Sci 70(5):321–325. https://doi.org/10.1111/j.1365-2621.2005.tb09960
Jackels SC, Jackels CF, Vallejos C, Kleven S, Rivas R, Fraser-Dauphinee S (2006) Control of the coffee fermentation process and quality of resulting roasted coffee: Studies in the field laboratory and on small farms in Nicaragua during the 2005–06 harvest. In: 21st International Scientific Colloquium on Coffee—Post-harvest processing and green coffee quality. Montpellier, France
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874. https://doi.org/10.1093/molbev/msw054
Lin CC (2010) Approach of improving coffee industry in Taiwan—promote quality of coffee bean by fermentation. J Int Manag Stud 5(1):154–159
Martinez SJ, Bressani APP, Dias DR, Simão JBP, Schwan RF (2019) Effect of bacterial and yeast starters on the formation of volatile and organic acid compounds in coffee beans and selection of flavors markers precursors during wet fermentation. Front Microbiol 10:1287. https://doi.org/10.3389/fmicb.2019.01287
Masoud W, Cesar LB, Jespersen L, Jakobsen M (2004) Yeast involved in fermentation of Coffea arabica in East Africa determined by genotyping and by direct denaturating gradient gel electrophoresis. Yeast 21(7):549–556. https://doi.org/10.1016/j.ijfoodmicro.2006.04.030
Nasanit R, Satayawut K (2015) Microbiological study during coffee fermentation of Coffea arabica var. chiangmai 80 in Thailand. Kasetsart J (nat Sci) 49:32–41
Nei M, Kumar S (2000) Molecular evolution and phylogenetics. Oxford University Press, New York
Pereira DMGV, Soccol VT, Pandey A, Medeiros ABP, Lara JMRA, Gollo AL, Soccol CR (2014) Isolation, selection and evaluation of yeasts for use in fermentation of coffee beans by the wet process. Int J Food Microbiol 188:60–66. https://doi.org/10.1016/j.ijfoodmicro.2014.07.008
Pereira DMGV, Neto ESVT, Medeiros ABP, Woiciechowski AL, Soccol CR (2015) Conducting starter culture-controlled fermentations of coffee beans during on-farm wet processing: Growth, metabolic analyses and sensorial effects. Food Res Int 75:348–356. https://doi.org/10.1016/j.foodres.2015.06.027
Pereira DMGV, da Silva VA, de Carvalho Neto DP, Muynarsk ESM, Soccol VT, Soccol CR (2020) Lactic acid bacteria: what coffee industry should know? Curr Opin Food Sci 31:1–8. https://doi.org/10.1016/j.cofs.2019.07.004
Ribeiro LS, Miguel MGCP, Martinez SJ, Bressani APP, Evangelista SR, e BatistaSchwan RF, CFSRF (2020) The use of mesophilic and lactic acid bacteria strains as starter cultures for improvement of coffee beans wet fermentation. World J Microbiol Biotechnol 36(12):1–15. https://doi.org/10.1007/s11274-020-02963-7
Siridevi GB, Havare D, Basavaraj K, Murthy PS (2019) Coffee starter microbiome and in-silico approach to improve Arabica coffee. LWT - Food Sci Technol 114:108382. https://doi.org/10.1016/j.lwt.2019.108382
Sunarharum WB, Williams DJ, Smyth HE (2014) Complexity of coffee flavor: a compositional and sensory perspective. Food Res Int 62:315–325. https://doi.org/10.1016/j.foodres.2014.02.030
White TJ, Bruns T, Lee S, Taylor J (1999) Amplification and direct sequencing of fungal ribosomal genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ (eds) PCR protocols: a guide to methods and applications. Academic press, New York, pp 315–322
Acknowledgements
This research was conducted in collaboration with UCC-K2 Co., Ltd., who provided assistance and contacted a coffee plantation to collect samples and conduct research by Mr. Meechai Amornpattanakul, President of the Thai Barista Association. This collaboration also included the Bank for Agriculture and Cooperatives, which supported this research. In addition, the researcher would like to thank the Royal Project Foundation for arranging contact with the coffee plantations in Chiang Mai under supervision for the researcher to travel to collect samples. In addition, this work was supported by the Kasetsart University Research and Development Institute (KURDI) grant no. FF(KU)18.64 under the coffee innovation research unit at Srinakharinwirot University grant no. 680/2563. Moreover, this work was funded by the Faculty of Science Srinakharinwirot University research fund Grant no. 491/2562.
Author information
Authors and Affiliations
Contributions
SK: methodology, data curation, formal analysis, investigation, writing—original draft, supervision. PS: methodology, experimenting. PT: review, manuscript revision, English editing. RM: methodology, experimenting. YM: methodology, experimenting. TP: experimenting. SS: review. KD: review. RJ: review. WL: writing—original draft, review and editing. NS: funding acquisition, project administration.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflicts of interest.
Compliance with ethics requirements
This article contains any studies with human subjects. This research has been certified ethics in human number SWUEC 041/62.
Informed consent
The authors used informed consent form for co-authors according to the regulations of the Ethics Committee for reviewing human research projects at Srinakharinwirot university.
Rights and permissions
About this article
Cite this article
Krajangsang, S., Seephin, P., Tantayotai, P. et al. New approach for screening of microorganisms from Arabica coffee processing for their ability to improve Arabica coffee flavor. 3 Biotech 12, 143 (2022). https://doi.org/10.1007/s13205-022-03203-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13205-022-03203-5