Abstract
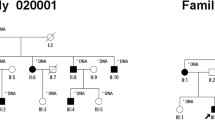
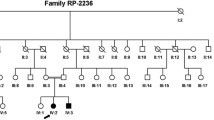
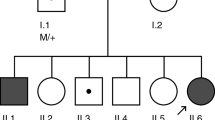
Retinitis pigmentosa (RP) is a rare and heterogeneous group of inherited ocular diseases. However, the relationship between CACNA2D4 mutations and RP is not well understood. In this study, a Chinese autosomal recessive retinitis pigmentosa (arRP) pedigree was enrolled and targeted next-generation sequencing was employed for identifying the causative gene in the proband. These steps were followed by confirmatory Sanger sequencing and segregation analysis. RNA-sequencing (RNA-seq) data and semi-quantitative reverse transcription polymerase chain reaction analysis were then applied to examine the expressions in the human and mouse tissues. Novel compound heterozygous, deleterious missense variants of the CACNA2D4 gene, NM_172364.4: c.G955A (p.D319N) and c.A1822C (p.I608L), were identified in the arRP pedigree, co-segregating with the clinical phenotype in the patient. The CACNA2D4 protein is highly conserved among species. The CACNA2D4 mRNA expression showed the highest expression in the retina of humans and in the later four developmental stages/times of retinal tissues in mice, indicating its role in retina/eye functions and developments. This study is the first to identify novel compound heterozygous mutations c.G955A (p.D319N) and c.A1822C (p.I608L) in the CACNA2D4 gene. These might be disease-causing mutations, thereby extending the mutational spectra. The identification of pathogenic CACNA2D4 variants is expected to enhance our understanding of the genotype–phenotype correlations of arRP for disease diagnosis and genetic counseling. The relationship between the CACNA2D4 variants and diseases/phenotypes other than RP has also been reviewed and discussed in this paper.




Similar content being viewed by others
Abbreviations
- RP:
-
Retinitis pigmentosa
- arRP:
-
Autosomal recessive retinitis pigmentosa
- CACNA2D4:
-
Homo sapiens calcium voltage-gated channel auxiliary subunit alpha2delta 4 gene
- NGS:
-
Next-generation sequencing
- TGS:
-
Targeted next-generation sequencing
- RNA-seq:
-
RNA-sequencing
- RT-PCR:
-
Revere transcriptional-polymerase chain reaction
- ExAC:
-
The Exome Aggregation Consortium database
- HGMD:
-
The Human Gene Mutation Database
- RPKM:
-
Reads Per Kilobase of transcript per Million
- OMIM:
-
Online Mendelian Inheritance in Man
- NX:
-
Consensus normalized expression
- VGCC:
-
Voltage-gated calcium channels
- BWA:
-
Burrow–Wheeler Aligner
- ERG:
-
Electroretinogram
References
Adams DR, Eng CM (2018) Next-generation sequencing to diagnose suspected genetic disorders. N Engl J Med 379(14):1353–1362. https://doi.org/10.1056/NEJMra1711801
Ali MU, Rahman MSU, Cao J, Yuan PX (2017) Genetic characterization and disease mechanism of retinitis pigmentosa; current scenario. 3 Biotech 7(4):251. https://doi.org/10.1007/s13205-017-0878-3
Ba-Abbad R, Arno G, Carss K, Stirrups K, Penkett CJ, Moore AT, Michaelides M, Raymond FL, Webster AR, Holder GE (2016) Mutations in CACNA2D4 cause distinctive retinal dysfunction in humans. Ophthalmology 123(3):668–671. https://doi.org/10.1016/j.ophtha.2015.09.045
Challis D, Yu J, Evani US, Jackson AR, Paithankar S, Coarfa C, Milosavljevic A, Gibbs RA, Yu F (2012) An integrative variant analysis suite for whole exome next-generation sequencing data. BMC Bioinf 13:8. https://doi.org/10.1186/1471-2105-13-8
Cheng J, Fu J, Zhou Q, Xiang X, Wei C, Yang L, Fu S, Khan MA, Lv H, Fu J (2019) A novel splicing mutation in the PRPH2 gene causes autosomal dominant retinitis pigmentosa in a Chinese pedigree. J Cell Mol Med 23(5):3776–3780. https://doi.org/10.1111/jcmm.14278
Cheng J, Peng J, Fu J, Khan MA, Tan P, Wei C, Deng X, Chen H, Fu J (2020) Identification of a novel germline BRCA2 variant in a Chinese breast cancer family. J Cell Mol Med 24(2):1676–1683. https://doi.org/10.1111/jcmm.14861
Fu J, Li L, Lu G (2002) Relationship between microdeletion on Y chromosome and patients with idiopathic azoospermia and severe oligozoospermia in the Chinese. Chin Med J 115(1):72–75
Fu JW, Ma L, Cheng JL, Yang LS, Wei CL, Fu SY, Lv HB, Chen R, Fu JJ (2018) A novel, homozygous nonsense variant of the CDHR1 gene in a Chinese family causes autosomal recessive retinal dystrophy by NGS-based genetic diagnosis. J Cell Mol Med 22(11):5662–5669. https://doi.org/10.1111/jcmm.13841
Fu J, Cheng J, Zhou Q, Wei C, Chen H, Lv H, Fu J (2019) A novel missense variant c.G644A (p.G215E) of the RPGR gene in a Chinese family causes X-linked retinitis pigmentosa. Biosci Rep 39:10. https://doi.org/10.1042/BSR20192235
Fu J, Shen S, Cheng J, Lv H, Fu J (2020a) A case of Usher syndrome type IIA caused by a rare USH2A homozygous frameshift variant with maternal uniparental disomy (UPD) in a Chinese family. J Cell Mol Med 24(14):7743–7750. https://doi.org/10.1111/jcmm.15405
Fu J, Zhou B, Zhang L, Balaji KS, Wei C, Liu X, Chen H, Peng J, Fu J (2020b) Expressions and significances of the angiotensin-converting enzyme 2 gene, the receptor of SARS-CoV-2 for COVID-19. Mol Biol Rep 47(6):4383–4392. https://doi.org/10.1007/s11033-020-05478-4
Genomes Project C, Abecasis GR, Altshuler D, Auton A, Brooks LD, Durbin RM, Gibbs RA, Hurles ME, McVean GA (2010) A map of human genome variation from population-scale sequencing. Nature 467(7319):1061–1073. https://doi.org/10.1038/nature09534
Goodloe AH, Evans JM, Middha S, Prasad A, Olson TM (2014) Characterizing genetic variation of adrenergic signalling pathways in Takotsubo (stress) cardiomyopathy exomes. Eur J Heart Fail 16(9):942–949. https://doi.org/10.1002/ejhf.145
Gustafson K, Duncan JL, Biswas P, Soto-Hermida A, Matsui H, Jakubosky D, Suk J, Telenti A, Frazer KA, Ayyagari R (2017) Whole genome sequencing revealed mutations in two independent genes as the underlying cause of retinal degeneration in an Ashkenazi Jewish Pedigree. Genes 8:9. https://doi.org/10.3390/genes8090210
Hamel CP (2014) Gene discovery and prevalence in inherited retinal dystrophies. CR Biol 337(3):160–166. https://doi.org/10.1016/j.crvi.2013.12.001
Hartong DT, Berson EL, Dryja TP (2006) Retinitis pigmentosa. Lancet 368(9549):1795–1809. https://doi.org/10.1016/S0140-6736(06)69740-7
Hoppa MB, Lana B, Margas W, Dolphin AC, Ryan TA (2012) alpha2delta expression sets presynaptic calcium channel abundance and release probability. Nature 486(7401):122–125. https://doi.org/10.1038/nature11033
Huang XF, Huang F, Wu KC, Wu J, Chen J, Pang CP, Lu F, Qu J, Jin ZB (2015) Genotype-phenotype correlation and mutation spectrum in a large cohort of patients with inherited retinal dystrophy revealed by next-generation sequencing. Genetics Med 17(4):271–278. https://doi.org/10.1038/gim.2014.138
Imani S, Cheng J, Mobasher-Jannat A, Wei C, Fu S, Yang L, Jadidi K, Khosravi MH, Mohazzab-Torabi S, Shasaltaneh MD, Li Y, Chen R, Fu J (2018a) Identification of a novel RPGRIP1 mutation in an Iranian family with leber congenital amaurosis by exome sequencing. J Cell Mol Med 22(3):1733–1742. https://doi.org/10.1111/jcmm.13454
Imani S, Ijaz I, Shasaltaneh MD, Fu S, Cheng J, Fu J (2018b) Molecular genetics characterization and homology modeling of the CHM gene mutation: a study on its association with choroideremia. Mutat Res 775:39–50. https://doi.org/10.1016/j.mrrev.2018.02.001
Kerov V, Laird JG, Joiner ML, Knecht S, Soh D, Hagen J, Gardner SH, Gutierrez W, Yoshimatsu T, Bhattarai S, Puthussery T, Artemyev NO, Drack AV, Wong RO, Baker SA, Lee A (2018) alpha2delta-4 Is required for the molecular and structural organization of rod and cone photoreceptor synapses. J Neurosci 38(27):6145–6160. https://doi.org/10.1523/JNEUROSCI.3818-16.2018
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25(14):1754–1760. https://doi.org/10.1093/bioinformatics/btp324
Liu X, Cheng J, Mei Z, Wei C, Khan MA, Peng J, Fu J (2020) SCAR marker for identification and discrimination of specific medicinal Lycium chinense Miller from Lycium species from ramp-PCR RAPD fragments. 3 Biotech 10(8):334. https://doi.org/10.1007/s13205-020-02325-y
Marchler-Bauer A, Bo Y, Han L, He J, Lanczycki CJ, Lu S, Chitsaz F, Derbyshire MK, Geer RC, Gonzales NR, Gwadz M, Hurwitz DI, Lu F, Marchler GH, Song JS, Thanki N, Wang Z, Yamashita RA, Zhang D, Zheng C, Geer LY, Bryant SH (2017) CDD/SPARCLE: functional classification of proteins via subfamily domain architectures. Nucleic Acids Res 45(D1):D200–D203. https://doi.org/10.1093/nar/gkw1129
Psaty BM, O’Donnell CJ, Gudnason V, Lunetta KL, Folsom AR, Rotter JI, Uitterlinden AG, Harris TB, Witteman JC, Boerwinkle E, Consortium C (2009) Cohorts for Heart and Aging Research in Genomic Epidemiology (CHARGE) Consortium: design of prospective meta-analyses of genome-wide association studies from 5 cohorts. Circ Cardiovasc Genet 2(1):73–80. https://doi.org/10.1161/CIRCGENETICS.108.829747
Purcell SM, Moran JL, Fromer M, Ruderfer D, Solovieff N, Roussos P, O’Dushlaine C, Chambert K, Bergen SE, Kahler A, Duncan L, Stahl E, Genovese G, Fernandez E, Collins MO, Komiyama NH, Choudhary JS, Magnusson PK, Banks E, Shakir K, Garimella K, Fennell T, DePristo M, Grant SG, Haggarty SJ, Gabriel S, Scolnick EM, Lander ES, Hultman CM, Sullivan PF, McCarroll SA, Sklar P (2014) A polygenic burden of rare disruptive mutations in schizophrenia. Nature 506(7487):185–190. https://doi.org/10.1038/nature12975
Qin N, Yagel S, Momplaisir ML, Codd EE, D’Andrea MR (2002) Molecular cloning and characterization of the human voltage-gated calcium channel alpha(2)delta-4 subunit. Mol Pharmacol 62(3):485–496
Rahbaran M, Hassani Doabsari M, Salavitabar S, Mokhberian N, Morovvati Z, Morovvati S (2019) A novel frameshift mutation in the EDA gene in an Iranian patient affected by X-linked hypohidrotic ectodermal dysplasia. Cell Mol Biol Lett 24:54. https://doi.org/10.1186/s11658-019-0174-9
Salvo J, Lyubasyuk V, Xu M, Wang H, Wang F, Nguyen D, Wang K, Luo H, Wen C, Shi C, Lin D, Zhang K, Chen R (2015) Next-generation sequencing and novel variant determination in a cohort of 92 familial exudative vitreoretinopathy patients. Invest Ophthalmol Vis Sci 56(3):1937–1946. https://doi.org/10.1167/iovs.14-16065
Tennessen JA, Bigham AW, O’Connor TD, Fu W, Kenny EE, Gravel S, McGee S, Do R, Liu X, Jun G, Kang HM, Jordan D, Leal SM, Gabriel S, Rieder MJ, Abecasis G, Altshuler D, Nickerson DA, Boerwinkle E, Sunyaev S, Bustamante CD, Bamshad MJ, Akey JM, Broad GO, Seattle GO, Project NES (2012) Evolution and functional impact of rare coding variation from deep sequencing of human exomes. Science 337(6090):64–69. https://doi.org/10.1126/science.1219240
Uhlen M, Oksvold P, Fagerberg L, Lundberg E, Jonasson K, Forsberg M, Zwahlen M, Kampf C, Wester K, Hober S, Wernerus H, Bjorling L, Ponten F (2010) Towards a knowledge-based Human Protein Atlas. Nat Biotechnol 28(12):1248–1250. https://doi.org/10.1038/nbt1210-1248
Valencia CA, Husami A, Holle J, Johnson JA, Qian Y, Mathur A, Wei C, Indugula SR, Zou F, Meng H, Wang L, Li X, Fisher R, Tan T, Hogart Begtrup A, Collins K, Wusik KA, Neilson D, Burrow T, Schorry E, Hopkin R, Keddache M, Harley JB, Kaufman KM, Zhang K (2015) Clinical Impact and Cost-Effectiveness of Whole Exome Sequencing as a Diagnostic Tool: A Pediatric Center’s Experience. Front Pediatr 3:67. https://doi.org/10.3389/fped.2015.00067
Van Den Bossche MJ, Strazisar M, De Bruyne S, Bervoets C, Lenaerts AS, De Zutter S, Nordin A, Norrback KF, Goossens D, De Rijk P, Green EK, Grozeva D, Mendlewicz J, Craddock N, Sabbe BG, Adolfsson R, Souery D, Del-Favero J (2012) Identification of a CACNA2D4 deletion in late onset bipolar disorder patients and implications for the involvement of voltage-dependent calcium channels in psychiatric disorders. Am J Med Genet B Neuropsychiatr Genet 159B(4):465–475. https://doi.org/10.1002/ajmg.b.32053
Wang K, Li M, Hakonarson H (2010) ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res 38(16):e164. https://doi.org/10.1093/nar/gkq603
Wang F, Wang H, Tuan HF, Nguyen DH, Sun V, Keser V, Bowne SJ, Sullivan LS, Luo H, Zhao L, Wang X, Zaneveld JE, Salvo JS, Siddiqui S, Mao L, Wheaton DK, Birch DG, Branham KE, Heckenlively JR, Wen C, Flagg K, Ferreyra H, Pei J, Khan A, Ren H, Wang K, Lopez I, Qamar R, Zenteno JC, Ayala-Ramirez R, Buentello-Volante B, Fu Q, Simpson DA, Li Y, Sui R, Silvestri G, Daiger SP, Koenekoop RK, Zhang K, Chen R (2014) Next generation sequencing-based molecular diagnosis of retinitis pigmentosa: identification of a novel genotype-phenotype correlation and clinical refinements. Hum Genet 133(3):331–345. https://doi.org/10.1007/s00439-013-1381-5
Weeke P, Muhammad R, Delaney JT, Shaffer C, Mosley JD, Blair M, Short L, Stubblefield T, Roden DM, Darbar D, National Heart L, Blood Institute GOESP (2014) Whole-exome sequencing in familial atrial fibrillation. Eur Heart J 35(36):2477–2483. https://doi.org/10.1093/eurheartj/ehu156
Wei C, Xiao T, Cheng J, Fu J, Zhou Q, Yang L, Lv H, Fu J (2020) Novel compound heterozygous EYS variants may be associated with arRP in a large Chinese pedigree. Biosci Rep 40:6. https://doi.org/10.1042/BSR20193443
Wycisk KA, Budde B, Feil S, Skosyrski S, Buzzi F, Neidhardt J, Glaus E, Nurnberg P, Ruether K, Berger W (2006a) Structural and functional abnormalities of retinal ribbon synapses due to Cacna2d4 mutation. Invest Ophthalmol Vis Sci 47(8):3523–3530. https://doi.org/10.1167/iovs.06-0271
Wycisk KA, Zeitz C, Feil S, Wittmer M, Forster U, Neidhardt J, Wissinger B, Zrenner E, Wilke R, Kohl S, Berger W (2006b) Mutation in the auxiliary calcium-channel subunit CACNA2D4 causes autosomal recessive cone dystrophy. Am J Hum Genet 79(5):973–977. https://doi.org/10.1086/508944
Zhang Q, Xu M, Verriotto JD, Li Y, Wang H, Gan L, Lam BL, Chen R (2016) Next-generation sequencing-based molecular diagnosis of 35 Hispanic retinitis pigmentosa probands. Sci Rep 6:32792. https://doi.org/10.1038/srep32792
Zhou B, Wei C, Khan MA, Chen H, Fu J (2019) Characterization and molecular cloning of novel isoforms of human spermatogenesis associated gene SPATA3. Mol Biol Rep 46(4):3827–3834. https://doi.org/10.1007/s11033-019-04825-4
Acknowledgements
This work was supported by the National Natural Science Foundation of China (31701087), the Special Training Program for Young Science and Technology Talents from Southwest Medical University (00031726), the Joint Research Foundation of Luzhou City and Southwest Medical University (2018LZXNYD-YL01).
Author information
Authors and Affiliations
Contributions
JF was in charge of the idea. J.F. and S.A. supervised the project. JC, JF, LZ, and CW performed DNA extraction, PCR, sequencing and data analysis. HL and QZ recruited the clinical patients and were in charge of the clinical assessment. JF and SK wrote the manuscript, and JF, SA and JC revised the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
The study has the Ethical Committees approval granted by the Southwest Medical University. All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards.
Consent for publication
Written informed consent was obtained from all participants.
Competing interests
The authors declare no conflict of interests.
Rights and permissions
About this article
Cite this article
Cheng, J., Zhou, Q., Fu, J. et al. Novel compound heterozygous missense variants (c.G955A and c.A1822C) of CACNA2D4 likely causing autosomal recessive retinitis pigmentosa in a Chinese patient. 3 Biotech 11, 208 (2021). https://doi.org/10.1007/s13205-021-02761-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13205-021-02761-4