Abstract
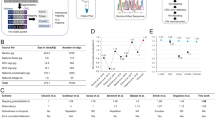
DNA has been recognized as a promising natural medium for information storage. The expensive DNA synthesis process makes it an important challenge to utilize DNA nucleotides optimally and increase the storage density. Thus, a novel scheme is proposed for the storage of digital information in synthetic DNA with high storage density and perfect error correction capability. The proposed strategy introduces quaternary Huffman coding to compress the binary stream of an original file before it is converted into a DNA sequence. The proposed quaternary Huffman coding is based on the statistical properties of the source and can gain a very high compression ratio for files with a non-uniform probability distribution of the source. Consequently, the amount of information that each base can store increases, and the storage density is also improved. In addition, quaternary Hamming code with low redundancy is proposed to correct errors occurring in the synthesis and sequencing. We have successfully converted a total of 5.2 KB of files into 3934 bits in DNA bases. The results of biological experiment indicate that the storage density of the proposed scheme is higher than that of state-of-the-art schemes.









Similar content being viewed by others
References
Ailenberg M, Rotstein O (2009) An improved Huffman coding method for archiving text, images, and music characters in DNA. Biotechniques 47:747–754
Akram F, Haq IU, Ali H, Laghari AT (2018) Trends to store digital data in DNA: an overview. Mol Biol Rep 45:1479–1490
Babu HMH, Mia MS, Biswas AK (2017) Efficient techniques for fault detection and correction of reversible circuits. J Electron Test 33:591
Bancroft C, Bowler T, Bloom B, Clelland CT (2001) Long-term storage of information in DNA. Science 5536:1763–1765
Blawat M, Gaedkea K, Hütter I, Chen XM, Turczyk B, Inverso S, Pruitt BW, Church GM (2016) Forward error correction for DNA data storage. Procedia Comput Sci 80:1011–1022
Borchert C, Schirmeier H, Spinczyk O (2013) Generative software-based memory error detection and correction for operating system data structures. In: Proceedings of the 2013 43rd annual IEEE/IFIP international conference on dependable systems and networks (DSN), pp 1–12
Bornholt J, Lopez R, Carmean DM (2016) A DNA-based archival storage system. IEEE Micro 99:637–649
Church GM, Kosuri S (2012) Next-generation digital information storage in DNA. Science 6102:1628
Davis J (1996) Microvenus. Art J 55:70–74
Dimopoulou M, Antonini M, Barbry P, Appuswamy R (2019) A biologically constrained encoding solution for long-term storage of images onto synthetic DNA. arXiv:1904:03024
Erlich Y, Zielinski D (2017) DNA fountain enables a robust and efficient storage architecture. Science 6328:950–954
Goda K, Kitsuregawa M (2012) The history of storage systems. Proc IEEE 2012:1433–1440
Goldman N, Bertone P, Chen SY, Dessimoz C, LeProust ME, Sipos B, Birney E (2013) Towards practical, high-capacity, low-maintenance information storage in synthesized DNA. Nature 7435:77–80
Grass RN, Heckel R, Puddu M, Paunescu D, Stark WJ (2015) Robust chemical preservation of digital information on DNA in silica with error-correcting codes. Angew Chem Int Ed 8:2552–2555
Hughes A, Ellington D (2017) Synthetic DNA synthesis and assembly: putting the synthetic in synthetic biology. Cold Spring Harbor Perspect Biol 1:a023812
Mardis R (2017) DNA sequencing technologies: 2006–2016. Nat Protoc 2:213–218
Panda D, Molla KA, Baig MJ, Swain A, Behera D, Dash M (2018) DNA as a digital information storage device: hope or hype? 3 Biotech 8:239
Rajaei N, Rajaei R, Tabandeh M (2017) A soft error tolerant register file for highly reliable microprocessor design. Int J High Perform Syst Archit 7:113–119
Shipman SL, Nivala J, Macklis JD, Church GM (2017) CRISPR-Cas encoding of a digital movie into the genomes of a population of living bacteria. Nature 7663:345–349
Yazdi SMHT, Kiah HM, Ruiz EG, Ma J, Zhao H, Milenkovic O (2015) DNA-based storage: trends and methods. IEEE Trans Mol Biol Multi-Scale Commun 1:230–248
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
On behalf of all authors, the corresponding author states that there is no conflict of interest.
Rights and permissions
About this article
Cite this article
Zhang, S., Huang, B., Song, X. et al. A high storage density strategy for digital information based on synthetic DNA. 3 Biotech 9, 342 (2019). https://doi.org/10.1007/s13205-019-1868-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13205-019-1868-4