Abstract
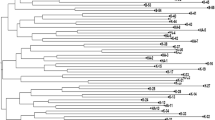
The genetic variation, marker attributes and population structure was assessed in 104 genotypes of cucumber using 23 SSR primer pairs. The total number of alleles produced was 67 with an average of 2.91 per locus. Allele frequency was in the range of 0.215 to 0.561 with mean value of 0.403, polymorphic information content ranged from 0.158 to 0.495 with the mean of 0.333, marker index ranged from 0.316 to 1.54 with an average value of 0.954 and resolving power ranged from 0.346 to 2.692 with mean of 1.392. The maximum allele frequency was reported with primer SSR65, whereas the maximum value of polymorphic information content and resolving power was found with SSR61 and the maximum value of marker index was reported with SSR60. Jaccard’s similarity coefficient ranged from 0.07 to 0.897 with maximum similarity between genotype G40 and G41 and minimum between G16 and G20, and G16 and G100. Clustering and PCA grouped the genotypes in two clusters, and majority of them were found in cluster B. The population structure analysis also showed two major populations, in which 47 genotypes were found in population 1, 39 genotypes in population 2, whereas remaining 18 genotypes were admixtures. The study provides researchers a valuable information for genotype identification, gene mapping, molecular breeding, and future exploration of cucumber germplasm in India and other major cucumber growing countries.



Similar content being viewed by others
References
Ammar MH, Alghamdi SS, Migdadi HM, Khan MA, El-Harty EH, Al-Faif SA (2015) Assessment of genetic diversity among faba bean genotypes using agro-morphological and molecular markers. Saudi J Biol Sci 22:340–350
Beckmann JS, Soller M (1990) Toward a unified approach to genetic mapping of eukaryotes based on sequence tagged microsatellite sites. Biotechnology 8:30–32
Bell CJ, Ecker JR (1994) Assignment of 30 microsatellite loci to the linkage map of Arabidopsis. Genomics 19:137–144
Botstein D, White RL, Skolnick M, Davis RW (1980) Construction of a genetic linkage map in man using restriction fragment length polymorphisms. Am J Hum Genet 32:314–331
Bradeen JM, Staub JE, Wye C, Antonise R, Peleman J (2001) Towards an expanded and integrated linkage map of cucumber (Cucumis sativus L.). Genome 44:111–119
Dang X, Thi TGT, Dong G, Wang H, Edzesi WM, Hong D (2014) Genetic diversity and association mapping of seed vigor in rice (Oryza sativa L.). Planta 239:1309–1319
Dar AA, Mudigunda S, Mittal PK, Arumugam N (2017) Comparative assessment of genetic diversity in Sesamum indicum L. using RAPD and SSR markers. 3. Biotech 7:10. doi:10.1007/s13205-016-0578-4
Dijkhuizen A, Kennard WC, Havey MJ, Staub JE (1996) RFLP variation and genetic relationships in cultivated cucumber. Euphytica 90:79–87
Dossa K, Wei X, Zhang Y, Fonceka D, Yang W et al (2016) Analysis of genetic diversity and population structure of sesame accessions from Africa and Asia as major centers of its cultivation. Genes 7:14. doi:10.3390/genes7040014
Doyle JJ, Doyle JL (1990) Isolation of plant DNA from fresh tissue. Focus 12:13–15
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software structure: a simulation study. Mol Ecol 14:2611–2620
FAO Statistical Database (2014) http://www.fao.org/faostat/en/#data/QC. Accessed 12 Jan 2017
Fukino N, Yoshioka Y, Kubo N, Hirai M, Sugiyama M, Sakata Y, Matsumoto S (2008) Development of 101 novel SSR markers and construction of an SSR-based genetic linkage map in cucumber (Cucumis sativus L.). Breed Sci 58:475–483
Garzon-Martinez GA, Osorio-Guarin JA, Delgadillo-Duran P, Mayorga F, Enciso-Rodriguez FE et al (2015) Genetic diversity and population structure in Physalis peruviana and related taxa based on InDels and SNPs derived from COSII and IRG markers. Plant Gene 4:29–37
Govindaraj M, Vetriventhan M, Srinivasan M (2015) Importance of genetic diversity assessment in crop plants and its recent advances: an overview of its analytical perspectives. Genet Res Int 2015:431487. doi:10.1155/2015/431487
Horejsi T, Staub JE (1999) Genetic variation in cucumber (Cucumis sativus L.) as assessed by random amplified polymorphic DNA. Genet Res Crop Evol 46:337–350
Horejsi T, Box JM, Staub JE (1999) Efficiency of randomly amplified polymorphic DNA to sequence characterized amplified region marker conversion and their comparative polymerase chain reaction sensitivity in cucumber. J Amer Soc Hort Sci 124:128–135
Hu J, Wang L, Li J (2011) Comparison of genomic SSR and EST-SSR markers for estimating genetic diversity in cucumber. Biol Plant 55:577–580
Hua J, Zhoub X, Li J (2010) Development of novel EST-SSR markers for cucumber (Cucumis sativus) and their transferability to related species. Sci Hortic 125:534–538
Kaçar YA, Simsek O, Solmaz I, Sari N, Mendi YY (2012) Genetic diversity among melon accessions (Cucumis melo) from Turkey based on SSR markers. Genet Mol Res 11:4622–4631
Liu K, Muse SV (2005) Power Marker. Integrated analysis environment for genetic marker data. Bioinformatics 21:2128–2129
Liu K, Goodman M, Muse S, Smith JS, Buckler E et al (2003) Genetic structure and diversity among maize inbred lines as inferred from DNA microsatellites. Genetics 165:2117–2128
Lv J, Qi J, Shi Q, Shen D, Zhang S et al (2012) Genetic diversity and population structure of cucumber (Cucumis sativus L.). PLoS One 7(10):e46919. doi:10.1371/journal.pone.0046919
Mahajan R, Zargar SM, Singh R, Salgotra RK, Farhat S, Sonah H (2016) Population structure analysis and selection of core set among common bean genotypes from Jammu and Kashmir., India. Appl. Biochem. doi:10.1007/s12010-016-2307-1
Messmer MM, Melchinger AE, Herrmann RG, Boppenmaier J (1993) Relationships among early European maize inbreds: II. Comparison of pedigree and RFLP data. Crop Sci 33:944–950
Miao H, Zhang S, Wang X, Zhang Z, Li M et al (2011) A linkage map of cultivated cucumber (Cucumis sativus L.) with 248 microsatellite marker loci and seven genes for horticulturally important traits. Euphytica 182:167–176
Pandey S, Ansari WA, Mishra VK, Singh AK, Singh M (2013) Genetic diversity in Indian cucumber based on microsatellite and morphological markers. Biochem Syst Ecol 51:19–27
Phan HTT, Ellwood SR, Hane JK, Ford R, Materne M, Oliver RP (2007) Extensive macrosynteny between Medicago truncatula and Lens culinaris ssp. culinaris. Theor Appl Genet 114:549–558
Powell W, Margenta M, Andre C, Hanfrey M, Vogel J, Tingey S, Rafalsky A (1996) The utility of RFLP, RAPD, AFLP and SSR (microsatellite) markers for germplasm analysis. Mol Breed 2:225–238
Prevost A, Wilkinson MJ (1999) A new system of comparing PCR primers applied to ISSR fingerprinting of potato cultivars. Theor Appl Genet 98:107–112
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Ranc N, Munos S, Santoni S, Causse M (2008) A clarified position for Solanum lycopersicum var. cerasiforme in the evolutionary history of tomatoes (solanaceae). BMC Plant Biol 8:130
Ren Y, Zhang ZH, Liu JH, Staub JE, Han YH et al (2009) An integrated genetic and cytogenentic map of the cucumber genome. PLoS One 4:e5795
Salem KFM, Roder MS, Borner A (2015) Assessing genetic diversity of Egyptian hexaploid wheat (Triticum aestivum L.) using microsatellite markers. Genet Res Crop Evol 62:377–385
Semagn K, Magorokosho C, Ogugo V, Makumbi D, Warburton ML (2014) Genetic relationships and structure among open-pollinated maize varieties adapted to eastern and southern Africa using microsatellite markers. Mol Breed 34:1423–1435
Sensoy S, Buyukalaca S, Abak K (2007) Evaluation of genetic diversity in Turkish melons (Cucumis melo L.) based on phenotypic characters and RAPD markers. Genet Resour Crop Evol 54:1351–1365
Shehata AI, Al-Ghethar HA, Al-Homaidan AA (2009) Application of simple sequence repeat (SSR) markers for molecular diversity and heterozygosity analysis in maize inbred lines. Saudi J Biol Sci 16:57–62
Staub JE, Chung Fazio G (2005) Conformity and genetic relatedness estimation in crop species having a narrow genetic base: the case of cucumber (Cucumis sativus L.). Plant Breed 124:44–53
Staub JE, Serquen FC, McCreight JD (1997) Genetic diversity in cucumber (Cucumis sativus L.): III. An evaluation of Indian germplasm. Genet Resour Crop Evol 44:315–326
Varshney RK, Chabane K, Hendre PS, Aggarwal RK, Graner A (2007) Comparative assessment of EST-SSR, EST-SNP and AFLP markers for evaluation of genetic diversity and conservation of genetic resources using wild, cultivated and elite barleys. Plant Sci 173:638–649
Watcharawongpaiboon N, Chunwongse J (2008) Development and characterization of microsatellite markers from an enriched genomic library of cucumber (Cucumis sativus). Plant Breed 127:74–81
Weng Y (2010) Genetic diversity among Cucumis metuliferus populations revealed by cucumber microsatellites. Hort Sci 45:214–219
Wright S (1978) Evolution and the genetics of populations: Variability within and among natural populations, vol 4. The University of Chicago Press, Chicago
Wu YQ, Huang Y (2006) An SSR genetic map of Sorghum bicolour (L.) Moench and its comparison to a published genetic map. Genome 50:84–89
Yang YT, Liu Y, Qi F, Xu LL, Li XZ et al (2015) Assessment of genetic diversity of cucumber cultivars in China based on simple sequence repeats and fruit traits. Genet Mol Res 14:19028–19039
Yuan XJ, Pan JS, Cai R, Guan Y, Liu LZ, Zhang WW, Li Z, He HL, Zhang C, Si LT, Zhu LH (2008) Genetic mapping and QTL analysis of fruit and flower related traits in cucumber (Cucumis sativus L.) using recombinant inbred lines. Euphytica 164:473–491
Zhou H, Xie Z, Ge S (2003) Microsatellite analysis of genetic diversity and population genetic structure of a wild rice (Oryza rufipogon Griff.) in China. Theor Appl Genet 107:332–339
Zhu H, Guo L, Song P, Luan F, Hu J, Sun X, Yang L (2016) Development of genome-wide SSR markers in melon with their cross-species transferability analysis and utilization in genetic diversity study. Mol Breed 36:153
Acknowledgements
We thank field assistants, Mr. Neetu Ram and Mr. Trivandrum for maintaining and growing the germplasm in our experimental field. We also thank DBT (Project sanction No. BT/PR11500/PBD/16/1101/2014) for providing us financial support for conducting the research.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Dar, A.A., Mahajan, R., Lay, P. et al. Genetic diversity and population structure of Cucumis sativus L. by using SSR markers. 3 Biotech 7, 307 (2017). https://doi.org/10.1007/s13205-017-0944-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13205-017-0944-x