Abstract
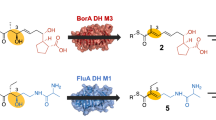
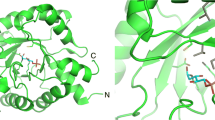
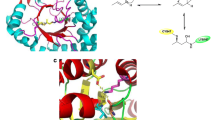
2-deoxyribose-5-phosphate aldolase (DERA) is a class I aldolase that catalyzes aldol condensation of two aldehydes in the active site, which is particularly germane in drug manufacture. Structural and biochemical studies have shown that the active site of DERA is typically loosely packed and displays broader substrate specificity despite sharing conserved folding architecture with other aldolases. The most distinctive structural feature of DERA compared to other aldolases is short and flexible C-terminal region. This region is also responsible for substrate recognition. Therefore, substrate tolerance may be related to the C-terminal structural features of DERA. Here, we determined the crystal structures of full length and C-terminal truncated DERA from Streptococcus suis (SsDERA). In common, both contained the typical (α/β)8 TIM-barrel fold of class I aldolases. Surprisingly, C-terminal truncation resulting in missing the last α9 and β8 secondary elements, allowed DERA to maintain activity comparable to the fulllength enzyme. Specifically, Arg186 and Ser205 residues at the C-terminus appeared mutually supplemental or less indispensible for substrate phosphate moiety recognition. Our results suggest that DERA might adopt a shorter C-terminal region than conventional aldolases during evolution pathway, resulting in a broader range of substrate tolerance through active site flexibility.
Similar content being viewed by others
References
Adams, P.D., Grosse-Kunstleve, R.W., Hung, L.W., Ioerger, T.R., McCoy, A.J., Moriarty, N.W., Read, R.J., Sacchettini, J.C., Sauter, N.K., and Terwilliger, T.C. 2002. Phenix: Building new software for automated crystallographic structure determination. Acta Crystallogr. D. Biol. Crystallogr. 58, 1948–1954.
Barth, P.T., Beacham, I.R., Ahmad, S.I., and Pritchard, R.H. 1968. The inducer of the deoxynucleoside phosphorylases and deoxyriboaldolase in Escherichia coli. Biochim. Biophys. Acta 161, 554–557.
Baugh, L., Phan, I., Begley, D.W., Clifton, M.C., Armour, B., Dranow, D.M., Taylor, B.M., Muruthi, M.M., Abendroth, J., Fairman, J.W., et al. 2015. Increasing the structural coverage of tuberculosis drug targets. Tuberculosis 95, 142–148.
Berkowitz, S.A. 2006. Role of analytical ultracentrifugation in assessing the aggregation of protein biopharmaceuticals. AAPS J. 8, e590–605.
Berthiaume, L., Loisel, T.P., and Sygusch, J. 1991. Carboxyl terminus region modulates catalytic activity of recombinant maize aldolase. J. Biol. Chem. 266, 17099–17105.
Blom, N. and Sygusch, J. 1997. Product binding and role of the cterminal region in class i d-fructose 1,6-bisphosphate aldolase. Nat. Struct. Biol. 4, 36–39.
Chen, V.B., Arendall, W.B., Headd, J.J., Keedy, D.A., Immormino, R.M., Kapral, G.J., Murray, L.W., Richardson, J.S., and Richardson, D.C. 2010. Molprobity: All-atom structure validation for macromolecular crystallography. Acta Crystallogr. D. Biol. Crystallogr. 66, 12–21.
Corsini, A., Maggi, F.M., and Catapano, A.L. 1995. Pharmacology of competitive inhibitors of HMg-CoA reductase. Pharmacol. Res. 31, 9–27.
DeSantis, G., Liu, J., Clark, D.P., Heine, A., Wilson, I.A., and Wong, C.H. 2003. Structure-based mutagenesis approaches toward expanding the substrate specificity of d-2-deoxyribose-5-phosphate aldolase. Bioorg. Med. Chem. 11, 43–52.
Edgar, R.C. 2004. Muscle: Multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797.
Emsley, P. and Cowtan, K. 2004. Coot: Model-building tools for molecular graphics. Acta Crystallogr. D. Biol. Crystallogr. 60, 2126–2132.
Endo, A. 1992. The discovery and development of hmgcoa reductase inhibitors. J. Lipid R. 33, 1569–1582.
Greenberg, W.A., Varvak, A., Hanson, S.R., Wong, K., Huang, H., Chen, P., and Burk, M.J. 2004. Development of an efficient, scalable, aldolase-catalyzed process for enantioselective synthesis of statin intermediates. Proc. Natl. Acad. Sci. USA 101, 5788–5793.
Guo, B.B., Devenish, S.R., Dobson, R.C., Muscroft-Taylor, A.C., and Gerrard, J.A. 2009. The c-terminal domain of Escherichia coli dihydrodipicolinate synthase (dhdps) is essential for maintenance of quaternary structure and efficient catalysis. Biochem. Biophys. Res. Commun. 380, 802–806.
Hannappel, E., MacGregor, J.S., Davoust, S., and Horecker, B.L. 1982. Limited proteolysis of liver and muscle aldolases: Effects of subtilisin, cathepsin b, and Staphylococcus aureus protease. Arch. Biochem. Biophys. 214, 293–298.
Heine, A., DeSantis, G., Luz, J.G., Mitchell, M., Wong, C.H., and Wilson, I.A. 2001. Observation of covalent intermediates in an enzyme mechanism at atomic resolution. Science 294, 369–374.
Heine, A., Luz, J.G., Wong, C.H., and Wilson, I.A. 2004. Analysis of the class i aldolase binding site architecture based on the crystal structure of 2-deoxyribose-5-phosphate aldolase at 0.99 Å resolution. J. Mol. Biol. 343, 1019–1034.
Humphreys, L., Reid, S., and Masters, C. 1986. Evidence for the spatial separation of the binding sites for substrate and for cytoskeletal proteins on the enzyme aldolase. Int. J. Biochem. 18, 7–13.
Istvan, E.S. and Deisenhofer, J. 2001. Structural mechanism for statin inhibition of HMg-CoA reductase. Science 292, 1160–1164.
Krissinel, E. and Henrick, K. 2004. Secondary-structure matching (ssm), a new tool for fast protein structure alignment in three dimensions. Acta Crystallogr. D. Biol. Crystallogr. 60, 2256–2268.
Krissinel, E. and Henrick, K. 2007. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797.
Liu, J., Andya, J.D., and Shire, S.J. 2006. A critical review of analytical ultracentrifugation and field flow fractionation methods for measuring protein aggregation. AAPS J. 8, e580–589.
Lokanath, N.K., Shiromizu, I., Ohshima, N., Nodake, Y., Sugahara, M., Yokoyama, S., Kuramitsu, S., Miyano, M., and Kunishima, N. 2004. Structure of aldolase from thermus thermophilus hb8 showing the contribution of oligomeric state to thermostability. Acta Crystallogr. D. Biol. Crystallogr. 60, 1816–1823.
Lun, Z.R., Wang, Q.P., Chen, X.G., Li, A.X., and Zhu, X.Q. 2007. Streptococcus suis: An emerging zoonotic pathogen. Lancet. Infect. Dis. 7, 201–209.
Machajewski, T.D. and Wong, C.H. 2000. The catalytic asymmetric aldol reaction. Angew. Chem. Int. Edi. Engl. 39, 1352–1374.
Otwinowski, Z. 1997. Processing of x-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 1–9.
Rashid, N., Imanaka, H., Fukui, T., Atomi, H., and Imanaka, T. 2004. Presence of a novel phosphopentomutase and a 2-deoxyribose 5-phosphate aldolase reveals a metabolic link between pentoses and central carbon metabolism in the hyperthermophilic archaeon Thermococcus kodakaraensis. J. Bacteriol. 186, 4185–4191.
Robert, X. and Gouet, P. 2014. Deciphering key features in protein structures with the new endscript server. Nucleic Acids Res. 42, W320–324.
Sakuraba, H., Tsuge, H., Shimoya, I., Kawakami, R., Goda, S., Kawarabayasi, Y., Katunuma, N., Ago, H., Miyano, M., and Ohshima, T. 2003. The first crystal structure of archaeal aldolase. Unique tetrameric structure of 2-deoxy-d-ribose-5-phosphate aldolase from the hyperthermophilic archaea Aeropyrum pernix. J. Biol. Chem. 278, 10799–10806.
Schuck, P. 2003. On the analysis of protein self-association by sedimentation velocity analytical ultracentrifugation. Anal. Biochem. 320, 104–124.
Sgarrella, F., Poddie, F.P.A., Meloni, M.A., Sciola, L., Pippia, P., and Tozzi, M.G. 1997. Channelling of deoxyribose moiety of exogenous DNA into carbohydrate metabolism: Role of deoxyriboaldolase. Comp. Biochem. Physiol. B. Biochem. Mol. Biol. 117, 253–257.
St-Jean, M. and Sygusch, J. 2007. Stereospecific proton transfer by a mobile catalyst in mammalian fructose-1,6-bisphosphate aldolase. J. Biol. Chem. 282, 31028–31037.
Staats, J.J., Feder, I., Okwumabua, O., and Chengappa, M.M. 1997. Streptococcus suis: Past and present. Vet. Res. Commun. 21, 381–407.
Sygusch, J., Beaudry, D., and Allaire, M. 1987. Molecular architecture of rabbit skeletal muscle aldolase at 2.7-a resolution. Proc. Natl. Acad. Sci. USA 84, 7846–7850.
Tozzi, M.G., Camici, M., Mascia, L., Sgarrella, F., and Ipata, P.L. 2006. Pentose phosphates in nucleoside interconversion and catabolism. FEBS J. 273, 1089–1101.
Vagin, A. and Teplyakov, A. 2010. Molecular replacement with molrep. Acta Crystallogr. D. Biol. Crystallogr. 66, 22–25.
Vedadi, M., Lew, J., Artz, J., Amani, M., Zhao, Y., Dong, A., Wasney, G.A., Gao, M., Hills, T., Brokx, S., et al. 2007. Genome-scale protein expression and structural biology of plasmodium falciparum and related apicomplexan organisms. Mol. Biochem. Parasitol. 151, 100–110.
Wertheim, H.F., Nghia, H.D., Taylor, W., and Schultsz, C. 2009. Streptococcus suis: An emerging human pathogen. Clin. Infect. Dis. 48, 617–625.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Cao, TP., Kim, JS., Woo, MH. et al. Structural insight for substrate tolerance to 2-deoxyribose-5-phosphate aldolase from the pathogen Streptococcus suis . J Microbiol. 54, 311–321 (2016). https://doi.org/10.1007/s12275-016-6029-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12275-016-6029-4