Abstract
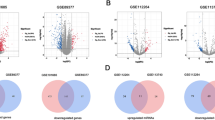
The purpose of this study was to explore potential biomarkers in the diagnosis of hepatocellular carcinoma (HCC) based on RNA-seq. The microarray data GSE98269 were downloaded from the GEO database, including the miRNA, mRNA and lncRNA expression profiles of 3 HCC tissues and 3 normal liver tissues from 3 HCC patients. The limma package was used to identify the differentially expressed miRNAs (DEMs) and the differentially expressed lncRNAs in HCC tissues compared with normal liver tissues. Database of DAVID, KEGG PATHWAY and Reactome were used to perform the functional and pathway enrichment. Putative targets for DEMs, and the miRNA-gene pairs were predicted via the miRWalk V2.0 database. The protein-protein pairs of DEGs were screened via String software. The expression features of the differentially expressed lncRNAs were analyzed. The regulated network of DEGs and DEMs were constructed, and related genes and miRNAs were detected in the HCC tissues and normal liver samples with Q-PCR. A total of 678 DEGs, 32 DEMs and 411 differential expressed lncRNAs were identified. The DEGs were enriched in 196 GO terms and 79 pathways. 38 negative regulation miRNA-gene pairs and 1205 protein-protein interactions were screened out, and the regulated network was constructed based on them. KNG1, CDK1, EHHADH, CYP3A4, hsa-miR-199a-5p and hsa-miR-455-3p might be biomarkers in the occurrence of HCC.

Similar content being viewed by others
Data Availability
The datasets used during the present study are available from the corresponding author upon reasonable request.
References
McGuire S (2016) World Cancer Report 2014. Geneva, Switzerland: World Health Organization, International Agency for Research on Cancer, WHO Press, 2015. Advances in Nutrition (Bethesda, Md) 7:418–419
Ferlay J, Soerjomataram I, Dikshit R et al (2015) Cancer incidence and mortality worldwide: sources, methods and major patterns in GLOBOCAN 2012. International Journal of Cancer 136:E359–E386
Siegel RL, Miller KD, Jemal A (2017) Cancer statistics, 2017. CA Cancer J Clin 67(9):7–30
Diaz-Gonzalez A, Forner A, Rodriguez de Lope C, Varela M (2016) New challenges in clinical research on hepatocellular carcinoma. Revista Espanola De Enfermedades Digestivas : Organo Oficial De La Sociedad Espanola De Patologia Digestiva 108:485–493
Li D, Satomura S (2015) Biomarkers for hepatocellular carcinoma (HCC): an update. Adv Exp Med Biol 867:179–193
Kiyokawa H, Yasuda H (2017) Serum Monomeric Laminin-Gamma2 as a Novel Biomarker for Hepatocellular Carcinoma. Cancer Science 108:1432–1439
Shaker O, Alhelf M, Morcos G, Elsharkawy A (2017) miRNA-101-1 and miRNA-221 expressions and their polymorphisms as biomarkers for early diagnosis of hepatocellular carcinoma. Infection, Genetics and Evolution : Journal of Molecular Epidemiology and Evolutionary Genetics in Infectious Diseases 51:173–181
Shen Q, Fan J, Yang XR et al (2012) Serum DKK1 as a protein biomarker for the diagnosis of hepatocellular carcinoma: a large-scale, multicentre study. Lancet Oncol 13:817–826
Huang S, Jiang F, Wang Y et al (2017) Diagnostic performance of tumor markers AFP and PIVKA-II in Chinese hepatocellular carcinoma patients. Tumour Biology : the Journal of the International Society for Oncodevelopmental Biology and Medicine 39:1010428317705763
Saitta C, Raffa G, Alibrandi A et al (2017) PIVKA-II is a useful tool for diagnostic characterization of ultrasound-detected liver nodules in cirrhotic patients. Medicine 96:e7266
Iarovaia GA (2001) Kallikrein-kinin system: novel facts and concepts (literature review). Vopr Med Khim 47:20–42
Colman RW (2006) Regulation of angiogenesis by the kallikrein-kinin system. Curr Pharm Des 12:2599–2607
Quesada-Calvo F, Massot C, Bertrand V et al (2017) OLFM4, KNG1 and Sec24C identified by proteomics and immunohistochemistry as potential markers of early colorectal cancer stages. Clin Proteomics 14:9
Ren Y, Ji N, Kang X et al (2016) Aberrant ceRNA-mediated regulation of KNG1 contributes to glioblastoma-induced angiogenesis. Oncotarget
Cho SJ, Lee SS, Kim YJ et al (2010) Xylocydine, a novel Cdk inhibitor, is an effective inducer of apoptosis in hepatocellular carcinoma cells in vitro and in vivo. Cancer Lett 287:196–206
Luo HB, Chen GW, Shao YX, Li Z, Liu M, Liu PQ (2012) 3D-QSAR studies on pyrimidine analogues as cyclin-dependent kinase 1 inhibitors. Shenzhen Daxue Xuebao 29:438–443
Zhao J, Han SX, Ma JL et al (2013) The role of CDK1 in apoptin-induced apoptosis in hepatocellular carcinoma cells. Oncol Rep 30:253–259
Yi Z, Wei H, Yan R et al (2015) miR-582-5p inhibits proliferation of hepatocellular carcinoma by targeting CDK1 and AKT3. Tumor Biol 36:8309–8316
Baes M, Van Veldhoven PP (2016) Hepatic dysfunction in peroxisomal disorders. BBA - Molecular Cell Research 1863:956–970
Houten SM, Denis S, Argmann CA (2012) Peroxisomal L-bifunctional enzyme (Ehhadh) is essential for the production of medium-chain dicarboxylic acids. J Lipid Res 53:1296–1303
Suto K, Kajihara-Kano H, Yokoyama Y et al (1999) Decreased expression of the peroxisomal bifunctional enzyme and carbonyl reductase in human hepatocellular carcinomas. J Cancer Res Clin Oncol 125:83–88
Zanger UM, Schwab M (2013) Cytochrome P450 enzymes in drug metabolism: regulation of gene expression, enzyme activities, and impact of genetic variation. Pharmacol Ther 138:103–141
Ali GT, Al-Azhary NM, Mokhtar DA (2014) Frequency and prognostic significant of CYP3A4-a-290G polymorphism in acute myeloid leukemia. J Adv Res 5:657–661
Rodrigues RM, De Kock J, Doktorova TY, Rogiers V, Vanhaecke T (2015) Measurement of cytochrome P450 enzyme induction and inhibition in human hepatoma cells. Methods Mol Biol 1250:279–285
Williams AE (2008) Functional aspects of animal microRNAs. Cell Mol Life Sci 65:545–562
Eulalio A, Huntzinger E, Nishihara T, Rehwinkel J, Fauser M, Izaurralde E (2009) Deadenylation is a widespread effect of miRNA regulation. RNA (New York, NY) 15:21–32
Hawkins PG, Morris KV (2008) RNA and transcriptional modulation of gene expression. Cell cycle (Georgetown, Tex) 7:602–607
Schultz S, Neubauer H, Bartsch H et al (2014) Hsa-miR-199a-5p is functionally relevant in progression from ductal carcinoma in situ to invasive breast tumours. Oncology Research & Treatment 37:68–68
Shen Q, Cicinnati VR, Zhang X et al (2010) Role of microRNA-199a-5p and discoidin domain receptor 1 in human hepatocellular carcinoma invasion. Mol Cancer 9:227
Sand M, Skrygan M, Sand D et al (2012) Expression of microRNAs in basal cell carcinoma. Br J Dermatol 167:847–855
Chickooree D, Zhu K, Ram V, Wu HJ, He ZJ, Zhang S (2016) A preliminary microarray assay of the miRNA expression signatures in buccal mucosa of oral submucous fibrosis patients. J Oral Pathol Med 45:691–697
Acknowledgements
We would like to thank all the members of our research group for their enthusiastic participation in this study.
Funding
Tianjin Clinical Research Center for Organ Transplantation (Grant No.15ZXLCSY00070); International Science and Technology Cooperation Program of China (Grant No. 2015DFG31850); National Key Clinical Specialty Construction Project of Organ Transplantation Department (Grant No.2013544); Science and Technology Fund of Tianjin Municipal Health Bureau (Grant No. 2013 KZ034); The study of precisely medical technology in the perioperative period of liver transplantation for liver cancer (Grant No. 17ZXMFSY00040); Spring Buds Project (Grant No. CL201801); General Program of The National Natural Science Foundation of China (Grant No. 81870444); Trial Registration Clinical Trial Registration URL: http://www.chictr.org.cn/showproj.aspx?proj=19662 webcite Trial Registration number: ChiCTR-ROR-17012157.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Jiang, W., Zhang, L., Guo, Q. et al. Identification of the Pathogenic Biomarkers for Hepatocellular Carcinoma Based on RNA-seq Analyses. Pathol. Oncol. Res. 25, 1207–1213 (2019). https://doi.org/10.1007/s12253-019-00596-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12253-019-00596-2