Abstract
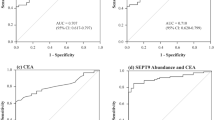
To determine the level of cell-free DNA (cfDNA), Septin 9 (SEPT9) and tumor markers (CEA, AFP, CA19–9, TPA, CA72–4). Plasma samples were collected four times a day (06:00, 12:00, 18:00, 24:00) from 9 patients with CRC (5 stage I-II, 4 stage III-IV), from one with colorectal adenoma and from one healthy control. CfDNA was isolated, quantified and bisulfite-converted. CfDNA and methylated SEPT9 were determined by RT-PCR. Plasma levels of conventional tumor markers were also measured. The lowest cfDNA concentrations were observed at 24:00 and 18:00 in stage I-III patients. In stage IV samples low cfDNA level (mean 48.2 ng/ml) were observed at several time points (6:00, 12:00, 18:00). The highest cfDNA levels were measured at 6:00 and 12:00 in CRC I-III stages and at 24:00 in stage IV samples (78.65 ng/ml). Higher in-day differences were found in stage II (43–48%) than in stage I samples (22%). Interestingly, the highest SEPT9 methylation level was found at 24:00 in most CRC cases, in contrast to the cfDNA levels. At 24:00, all cancer and adenoma cases were positive for SEPT9 methylation. At other time points (6:00, 12:00, 18:00) only 77.7% of CRC samples showed SEPT9 positivity. Stage I samples were SEPT9 positive only at 24:00. CEA and CA19–9 levels displayed correlation with the amount of cfDNA in case of late stage cases. Daytime activity can influence SEPT9 positivity in cases with low concentration of cfDNA. Thus, it may improve screening sensitivity by collecting samples earlier in the morning.


Similar content being viewed by others
References
Crowley E, Di Nicolantonio F, Loupakis F, Bardelli A (2013) Liquid biopsy: monitoring cancer-genetics in the blood. Nat Rev Clin Oncol 10:472–484. doi:10.1038/nrclinonc.2013.110
Mandel P, Métais P (1948) Nucleic acids in human blood plasma. C R Acad Sci Paris 15:241–243
Leon SA, Shapiro B, Sklaroff DM, Yaros MJ (1977) Free DNA in the serum of cancer patients and the effect of therapy. Cancer Res 15:646–650
Stroun M, Anker P, Maurice P, Lyautey J, Lederrey C, Beljanski M (1989) Neoplastic characteristics of the DNA found in the plasma of cancer patients. Oncology 15:318–322. doi:10.1159/000226740
Jahr S, Hentze H, Englisch S, Hardt D, Fackelmayer FO, Hesch RD, Knippers R (2001) DNA fragments in the blood plasma of cancer patients: quantitations and evidence for their origin from apoptotic and necrotic cells. Cancer Res 15:1659–1665
Brancaccio P, Lippi G, Maffulli N (2010) Biochemical markers of muscular damage. Clin Chem Lab Med 48:757–767. doi:10.1515/CCLM.2010.179
Chevion S, Moran DS, Heled Y, Shani Y, Regev G, Abbou B, Berenshtein E, Stadtman ER, Epstein Y (2003) Plasma antioxidant status and cell injury after severe physical exercise. Proc Natl Acad Sci U S A 100:5119–5123. doi:10.1073/pnas.0831097100
Fatouros IG, Destouni A, Margonis K, Jamurtas AZ, Vrettou C, Kouretas D, Mastorakos G, Mitrakou A, Taxildaris K, Kanavakis E, Papassotiriou I (2006) Cell-free plasma DNA as a novel marker of aseptic inflammation severity related to exercise overtraining. Clin Chem 52:1820–1824. doi:10.1373/clinchem.2006.070417
Brinkmann V, Zychlinsky A (2007) Beneficial suicide: why neutrophils die to make NETs. Nat Rev Microbiol 5:577–582. doi:10.1038/nrmicro1710
Tjoa ML, Cindrova-Davies T, Spasic-Boskovic O, Bianchi DW, Burton GJ (2006) Trophoblastic oxidative stress and the release of cell-free feto-placental DNA. Am J Pathol 169:400–404. doi:10.2353/ajpath.2006.060161
Maron JL, Bianchi DW (2007) Prenatal diagnosis using cell-free nucleic acids in maternal body fluids: a decade of progress. Am J Med Genet C Semin Med Genet 145C:5–17. doi:10.1002/ajmg.c.30115
Leung TN, Zhang J, Lau TK, Chan LYS, Lo D (2001) Increased maternal plasma fetal DNA concentrations in women who eventually develop preeclampsia. Clin Chem 47:137–139
Velders M, Treff G, Machus K, Bosnyák E, Steinacker J, Schumann U (2013) Exercise is a potent stimulus for enhancing circulating DNase activity. Clin Biochem 47:471–474. doi:10.1016/j.clinbiochem.2013.12.017
Stroun M, Lyautey J, Lederrey C, Olson-Sand A, Anker P (2001) About the possible origin and mechanism of circulating DNA apoptosis and active DNA release. Clin Chim Acta 15:139–142. doi:10.1016/S0009-8981(01)00665-9
Kettner NM, Katchy CA, Fu L (2014) Circadian gene variants in cancer. Ann Med 46:208–220. doi:10.3109/07853890.2014.914808
Lee JH, Sancar A (2011) Regulation of apoptosis by the circadian clock through NF-kappa B signaling. Proc Natl Acad Sci U S A 108:12036–12041. doi:10.1073/pnas.1108125108
Haus E, Smolensky MH (1999) Biologic rhythms in the immune sys-tem. Chronobiol Int 16:581–622
Born J, Lange T, Hansen K, Molle M, Fehm HL (1997) Effects of sleep and circadian rhythm on human circulating immune cells. J Immunol 158:4454–4464
Kawate T, Abo T, Hinuma S, Kumagai K (1981) Studies of the bioperio-dicity of the immune response. II. Co-variations of murine T and B cells and a role of corticosteroid. J Immunol 126:1364–1367
Braude S, Beck A (2013) Complete blood counts with differential: more accurate reference ranges based oncircadian leukocyte trafficking. J Clin Pathol 66:909–910. doi:10.1136/jclinpath-2013-201776
Vitaterna MH, King DP, Chang AM, Kornhauser JM, Lowrey PL, McDonald JD, Dove WF, Pinto LH, Turek FW, Takahashi JS (1994) Muta-genesis and mapping of a mouse gene, clock, essential for circadian behavior. Science 264:719–725
Lee C, Etchegaray JP, Cagampang FR, Loudon AS, Reppert SM (2001) Posttranslational mechanisms regulate the mammalian circadian clock. Cell 107:855–867
Panda S, Antoch MP, Miller BH, Su AI, Schook AB, Straume M, Schultz PG, Kay SA, Takahashi JS, Hogenesch JB (2002) Coordinated transcription of key pathways in the mouse by the circadian clock. Cell 109:307–320. doi:10.1016/S0092-8674(02)00722-5
Wood PA, Du-Quiton J, You S, Hrushesky WJ (2006) Circadian clock coordinates cancer cell cycle progression, thymidylate synthase, and 5-fluorouracil therapeutic index. Mol Cancer Ther 5:2023–2033. doi:10.1158/1535-7163.MCT-06-0177
Filipski E, King VM, Li X, Granda TG, Mormont MC, Liu X, Claustrat B, Hastings MH, Lévi F (2002) Host circadian clock as a control point in tumor progression. J Natl Cancer Inst 94:690–697. doi:10.1093/jnci/94.9.690
Schernhammer ES, Laden F, Speizer FE, Willett WC, Hunter DJ, Kawachi I, Colditz GA (2001) Rotating night shifts and risk of breast cancer in women participating in the nurses’ health study. J Natl Cancer Inst 93:1563–1568. doi:10.1093/jnci/93.20.1563
Cash E, Sephton SE, Chagpar AB, Spiegel D, Rebholz WN, Zimmaro LA, Tillie JM, Dhabhar FS (2015) Circadian disruption and biomarkers of tumor progression in breast cancer patients awaiting surgery. Brain Behav Immun 48:102–114. doi:10.1016/j.bbi.2015.02.017
Focan C, Focan-Henrard D, Collette J, Mechkouri M, Levi F, Hrushesky W, Touitou Y, Franchimont P (1986) Cancer-associated alteration of circadian rhythms in carcinoembryonic antigen (CEA) and alpha-fetoprotein (AFP) in humans. Anticancer Res 6:1137–1144
Frederiksen CB, Lomholt AF, Lottenburger T, Davis GJ, Dowell BL, Blankenstein MA, Christensen IJ, Brunner N, Nielsen HJ (2008) Assessment of the biological variation of plasma tissue inhibitor of metalloproteinases-1. Int J Biol Markers 23:42–47
Grützmann R, Molnar B, Pilarsky C, Habermann JK, Schlag PM, Saeger HD, Miehlke S, Stolz T, Model F, Roblick UJ, Bruch HP, Koch R, Liebenberg V, Devos T, Song X, Day RH, Sledziewski AZ, Lofton-Day C (2008) Sensitive detection of colorectal cancer in peripheral blood by septin 9 DNA methylation assay. PLoS One 3:e3759. doi:10.1371/journal.pone.0003759
deVos T, Tetzner R, Model F, Weiss G, Schuster M, Distler J, Steiger KV, Grützmann R, Pilarsky C, Habermann JK, Fleshner PR, Oubre BM, Day R, Sledziewski AZ, Lofton-Day C (2009) Circulating methylated SEPT9 DNA in plasma is a biomarker for colorectal cancer. Clin Chem 55:1337–1346. doi:10.1373/clinchem.2008.115808
Tänzer M, Balluff B, Distler J, Hale K, Leodolter A, Röcken C, Molnar B, Schmid R, Lofton-Day C, Schuster T, Ebert MP (2010) Performance of epigenetic markers SEPT9 and ALX4 in plasma for detection of colorectal precancerous lesions. PLoS One 5:e9061. doi:10.1371/journal.pone.0009061
Johnson DA, Barclay RL, Mergener K, Weiss G, König T, Beck J, Potter NT (2014) Plasma Septin9 versus fecal immunochemical testing for colorectal cancer screening: a prospective multicenter study. PLoS One 9:e98238. doi:10.1371/journal.pone.0098238
Potter NT, Hurban P, White MN, Whitlock KD, Lofton-Day CE, Tetzner R, Koenig T, Quigley NB, Weiss G (2014) Validation of a real-time PCR-based qualitative assay for the detection of methylated SEPT9 DNA in human plasma. Clin Chem 60:1183–1191. doi:10.1373/clinchem.2013.221044
Jin P, Kang Q, Wang X, Yang L, Yu Y, Li N, He YQ, Han X, Hang J, Zhang J, Song L, Han Y, Sheng JQ (2014) Performance of a second generation methylated SEPT9 test in detecting colorectal neoplasm. J Gastroenterol Hepatol 30:830–833. doi:10.1111/jgh.12855
Tóth K, Sipos F, Kalmár A, Patai AV, Wichmann B, Stoehr R, Golcher H, Schellerer V, Tulassay Z, Molnár B (2012) Detection of methylated SEPT9 in plasma is a reliable screening method for both left- and right-sided colon cancers. PLoS One 7:e46000. doi:10.1371/journal.pone.0046000
Church TR, Wandell M, Lofton-Day C, Mongin SJ, Burger M, Payne SR, Castaños-Vélez E, Blumenstein BA, Rösch T, Osborn N, Snover D, Day RW, Ransohoff DF, PRESEPT Clinical Study Steering Committee, Investigators and Study Team (2014) Prospective evaluation of methylated SEPT9 in plasma for detection of asymptomatic colorectal cancer. Gut 63:317–325. doi:10.1136/gutjnl-2012-304149
Tóth K, Wasserkort R, Sipos F, Kalmár A, Wichmann B, Leiszter K, Valcz G, Juhász M, Miheller P, Patai ÁV, Tulassay Z, Molnár B (2014) Detection of methylated septin 9 in tissue and plasma of colorectal patients with neoplasia and the relationship to the amount of circulating cell-free DNA. PLoS One 9:e115415. doi:10.1371/journal.pone.0115415
Velders M, Treff G, Machus K, Bosnyák E, Steinacker J, Schumann U (2014) Exercise is a potent stimulus for enhancing circulating DNase activity. Clin Biochem 47:471–474. doi:10.1016/j.clinbiochem.2013.12.017
Karantanos T, Theodoropoulos G, Pektasides D, Gazouli M (2014) Clock genes: their role in colorectal cancer. World J Gastroenterol 20:1986–1992. doi:10.3748/wjg.v20.i8.1986
Mazzoccoli G, Panza A, Valvano MR, Palumbo O, Carella M, Pazienza V, Biscaglia G, Tavano F, Di Sebastiano P, Andriulli A, Piepoli A (2011) Clock gene expression levels and relationship with clinical and pathological features in colorectal cancer patients. Chronobiol Int 28:841–851. doi:10.3109/07420528.2011.615182
Warren JD, Xiong W, Bunker AM, Vaughn CP, Furtado LV, Roberts WL, Fang JC, Samowitz WS, Heichman KA (2011) Septin 9 methylated DNA is a sensitive and specific blood test for colorectal cancer. BMC Med 9:133. doi:10.1186/1741-7015-9-133
Ali T, Choe J, Awab A, Wagener TL, Orr WC (2013) Sleep, immunity and inflammation in gastrointestinal disorders. World J Gastroenterol 19:9231–9239. doi:10.3748/wjg.v19.i48.9231
Zubelewicz-Szkodzinska B, Muc-Wierzgon M, Wierzgon J, Brodziak A (2001) Dynamics of circadian fluctuations in serum concentration of cortisol and TNF-alpha soluble receptors in gastrointestinal cancer patients. Oncol Rep 8:207–212. doi:10.3892/or.8.1.207
Rich T, Innominato PF, Boerner J, Mormont MC, Iacobelli S, Baron B, Jasmin C, Lévi F (2005) Elevated serum cytokines correlated with altered behavior, serum cortisol rhythm, and dampened 24-hour rest-activity patterns in patients with metastatic colorectal cancer. Clin Cancer Res 11:1757–1764. doi:10.1158/1078-0432.CCR-04-2000
Halberg F, Haus E, Lakatua DJ, Antinozzi R, Cornélissen G (1995) Cancer marker assessment: case report on salivary and urinary CEA. In Vivo 9:311–314
Acknowledgements
We thank both the Endoscopy Unit of the 2nd Department of Internal Medicine, Semmelweis University, and the nurses at the 2nd Department of Internal Medicine for their technical assistance. We also thank Bernadett Tóth for blood sample and data collection and plasma preparation. We thank Prof. Barna Vásárhelyi, Tünde Holczer and all the colleagues at the Department of Laboratory Medicine for the technical assistance. Finally, we thank all of the included patients for the blood collection.
The authors thank Theo deVos the help, scientific comments and support our review paper.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Tóth, K., Patai, Á.V., Kalmár, A. et al. Circadian Rhythm of Methylated Septin 9, Cell-Free DNA Amount and Tumor Markers in Colorectal Cancer Patients. Pathol. Oncol. Res. 23, 699–706 (2017). https://doi.org/10.1007/s12253-016-0174-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12253-016-0174-2