Abstract
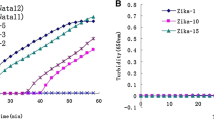
The establishment of highly sensitive diagnostic methods is critical in the early diagnosis and control of Zika virus (ZIKV) and in preventing serious neurological complications of ZIKV infection. In this study, we established micro-droplet digital polymerase chain reaction (ddPCR) and real-time quantitative PCR (RT-qPCR) protocols for the detection of ZIKV based on the amplification of the NS5 gene. For the ZIKV standard plasmid, the RT-qPCR results showed that the cycle threshold (Ct) value was linear from 101 to 108 copy/μL, with a standard curve R2 of 0.999 and amplification efficiency of 92.203%; however, a concentration as low as 1 copy/μL could not be detected. In comparison with RT-qPCR, the ddPCR method resulted in a linear range of 101–104 copy/μL and was able to detect concentrations as low as 1 copy/μL. Thus, for detecting ZIKV from clinical samples, RT-qPCR is a better choice for high-concentration samples (above 101 copy/μL), while ddPCR has excellent accuracy and sensitivity for low-concentration samples. These results indicate that the ddPCR method should be of considerable use in the early diagnosis, laboratory study, and monitoring of ZIKV.



Similar content being viewed by others
References
Balm MN, Lee CK, Lee HK, Chiu L, Koay ES, Tang JW (2012) A diagnostic polymerase chain reaction assay for Zika virus. J Med Virol 84:1501–1505
Bizouarn F (2014) Introduction to digital PCR. Meth Mol Biol 1160:27–41
Calvert AE, Biggerstaff BJ, Tanner NA, Lauterbach M, Lanciotti RS (2017) Rapid colorimetric detection of Zika virus from serum and urine specimens by reverse transcription loop-mediated isothermal amplification (RT-LAMP). PLoS ONE 12:e0185340
Cao Y, Raith MR, Griffith JF (2015) Droplet digital PCR for simultaneous quantification of general and human-associated fecal indicators for water quality assessment. Water Res 70:337–349
Chouin-Carneiro T, Vega-Rua A, Vazeille M, Yebakima A, Girod R, Goindin D, Dupont-Rouzeyrol M, Lourenço-de-Oliveira R, Failloux AB (2016) Differential susceptibilities of Aedes aegypti and Aedes albopictus from the Americas to Zika virus. PLoS Negl Trop Dis 10:e0004543
Dick GW, Kitchen SF, Haddow AJ (1952) Zika virus (I). Isolations and serological specificity. Trans R Soc Trop Med Hyg 46:509–520
Duffy MR, Chen TH, Hancock WT, Powers AM, Kool JL, Lanciotti RS, Pretrick M, Marfel M, Holzbauer S, Dubray C, Guillaumot L, Griggs A, Bel M, Lambert AJ, Laven J, Kosoy O, Panella A, Biggerstaff BJ, Fischer M, Hayes EB (2009) Zika virus outbreak on Yap Island, federated states of Micronesia. N Engl J Med 360:2536–2543
Faye O, Freire CCM, Iamarino A, Faye O, de Oliveira JVC, Diallo M, Zanotto PMA, Sall AA (2014) Molecular evolution of Zika virus during its emergence in the 20(th) century. PLoS Negl Trop Dis 8:e2636
Floren C, Wiedemann I, Brenig B, Schutz E, Beck J (2014) Species identification and quantification in meat and meat products using droplet digital PCR (ddPCR). Food Chem 173:1054–1058
Fontes-Garfias CR, Shan C, Luo H, Muruato AE, Medeiros DBA, Mays E, Xie X, Zou J, Roundy CM, Wakamiya M, Rossi SL, Wang T, Weaver SC, Shi PY (2017) Functional analysis of glycosylation of Zika virus envelope protein. Cell Rep 21:1180–1190
Frankel MB, Pandya K, Gersch J, Siddiqui S, Schneider GJ (2017) Development of the Abbott Real Time ZIKA assay for the qualitative detection of Zika virus RNA from serum, plasma, urine, and whole blood specimens using the m2000 system. J Virol Methods 246:117–124
Gourinat AC, O’Connor O, Calvez E, Goarant C, Dupont-Rouzeyrol M (2015) Detection of Zika virus in urine. Emerg Infect Dis 21:84–86
Hancock WT, Marfel M, Bel M (2014) Zika virus, French Polynesia, South Pacific, 2013. Emerg Infect Dis 20:1085–1086
Hayden RT, Gu Z, Ingersoll J, Abdul-Ali D, Shi L, Pounds S, Caliendo AM (2013) Comparison of droplet digital PCR to real-time PCR for quantitative detection of cytomegalovirus. J Clin Microbiol 51:540–546
Hayes EB (2009) Zika virus outside Africa. Emerg Infect Dis 15:1347–1350
Hindson BJ, Ness KD, Masquelier DA, Masquelier DA, Belgrader P, Heredia NJ, Makarewicz AJ, Bright IJ, Lucero MY, Hiddessen AL, Legler TC, Kitano TK, Hodel MR, Petersen JF, Wyatt PW, Steenblock ER, Shah PH, Bousse LJ, Troup CB, Mellen JC, Wittmann DK, Erndt NG, Cauley TH, Koehler RT, So AP, Dube S, Rose KA, Montesclaros L, Wang S, Stumbo DP, Hodges SP, Romine S, Milanovich FP, White HE, Regan JF, Karlin-Neumann GA, Hindson CM, Saxonov S, Colston BW (2011) High-throughput droplet digital PCR system for absolute quantitation of DNA copy number. Anal Chem 83:8604–8610
Nakano M, Nakai N, Kurita H, Komatsu J, Takashima K, Katsura S, Mizuno A (2005) Single-molecule reverse transcription polymerase chain reaction using water-in-oil emulsion. J Biosci Bioeng 99:293–295
Nathan LM, Simmons M, Wegleitner BJ, Jerde CL, Mahon AR (2014) Quantifying environmental DNA signals for aquatic invasive species across multiple detection platforms. Environ Sci Technol 48:12800–12806
Pinheiro LB, Coleman VA, Hindson CM, Herrmann J, Hindson BJ, Bhat S, Emslie KR (2012) Evaluation of a droplet digital polymerase chain reaction format for DNA copy number quantification. Anal Chem 84:1003–1011
Priyamvada L, Quicke KM, Hudson WH, Onlamoon N, Sewatanon J, Edupuganti S, Pattanapanyasat K, Chokephaibulkit K, Mulligan MJ, Wilson PC, Ahmed R, Suther MS, Wrammert J (2016) Human antibody responses after dengue virus infection are highly cross-reactive to Zika virus. Proc Natl Acad Sci 113:7852–7857
Rossini G, Gaibani P, Vocale C, Cagarelli R, Landini MP (2017) Comparison of Zika virus (ZIKV) RNA detection in plasma, whole blood and urine–Case series of travel-associated ZIKV infection imported to Italy. J Infect 75:242–245
Rutsaert S, Bosman K, Trypsteen W, Nijhuis M, Vandekerckhove L (2018) Digital PCR as a tool to measure HIV persistence. Retrovirology 15:16
Sedlak RH, Jerome KR (2013) Viral diagnostics in the era of digital polymerase chain reaction. Diagn Microbiol Infect Dis 75:1–4
Sidransky D, Tokino T, Hamilton SR, Kinzler KW, Levin B, Frost P, Vogelstein B (1992) Identification of ras oncogene mutations in the stool of patients with curable colorectal tumors. Science 256:102–105
Strain MC, Lada SM, Luong T, Rought SE, Gianella S, Terry VH, Spina CA, Woelk CH, Richman DD (2013) Highly precise measurement of HIV DNA by droplet digital PCR. PLoS ONE 8:e55943
Sykes PJ, Neoh SH, Brisco MJ, Hughes E, Condon J, Morley AA (1992) Quantitation of targets for PCR by use of limiting dilution. Biotechniques 13:444–449
Taylor SC, Carbonneau J, Shelton DN, Boivin G (2015) Optimization of Droplet digital PCR from RNA and DNA extracts with direct comparison to RT-qPCR: clinical implications for quantification of Oseltamivir-resistant subpopulations. J Virol Methods 224:58–66
Weaver SC, Costa F, Garcia-Blanco MA, Ko AI, Ribeiro GS, Saade G, Shi PY, Vasilakis N (2016) Zika virus: history, emergence, biology, and prospects for control. Antiviral Res 130:69–80
Xin H, Xuan Z, Xiao H, Pei W, Shuai Y, Li W, Yu Z, Qian X, Bao Z, Wei Z (2016) Detection of infectious dengue virus by selective real-time quantitative polymerase chain reaction. Virol Sin 31:342–345
Xu MY, Liu SQ, Deng CL, Zhang QY, Zhang B (2016) Detection of Zika virus by SYBR green one-step real-time RT-PCR. J Virol Methods 236:93–97
Zhao H, Wilkins K, Damon IK, Li Y (2013) Specific qPCR assays for the detection of virus, Pseudocowpox virus and bovine papular stomatitis virus. J Virol Methods 194:229–234
Acknowledgements
This study was supported by the National Natural Science Foundation of China (Nos. 31470271 and 81730110) and Guangzhou Science and Technology Program key projects (No. 201803040006). We thank Changwen Ke, De Wu, and Jiufeng Sun from the Guangdong Center for Disease Control and Prevention for providing the Asian ZIKV Z16006 strain. The authors would like to thank anonymous referees for their valuable suggestions, which have improved the paper immensely.
Author information
Authors and Affiliations
Contributions
ZQ and YH designed the experiments. ZW, LZ, JL, XL, HF, and SF carried out the experiments. JY, XH and WL analyzed the data. WX, and QW wrote the paper. BZ and WZ checked and finalized the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Animal and Human Rights Statement
The study was approved by the Ethics Committees of Southern Medical University. All participants provided written informed consent.
Additional information
Yuan Hui, Zhiming Wu, Zhiran Qin and Li Zhu have contribued equally to this work.
Rights and permissions
About this article
Cite this article
Hui, Y., Wu, Z., Qin, Z. et al. Micro-droplet Digital Polymerase Chain Reaction and Real-Time Quantitative Polymerase Chain Reaction Technologies Provide Highly Sensitive and Accurate Detection of Zika Virus. Virol. Sin. 33, 270–277 (2018). https://doi.org/10.1007/s12250-018-0037-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12250-018-0037-y