Abstract
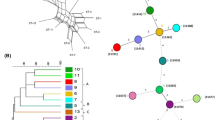
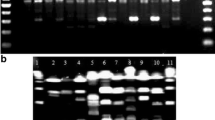
A high-throughput, medium-discrimination method for preliminary typing and selecting non-identical isolates of lactic acid bacteria in cheeses was developed. RAMP, a PCR with one microsatellite-targeted and one random primer, was used for preliminary typing of 1119 isolates of lactic acid bacteria from Slovak Bryndza cheese. A total of 59 genotypes were identified based on RAMP profiles consisting of 12–23 DNA fragments of 150–3000 bp. For example, 18, 17, 13 and 7 different RAMP-types were identified in Lactobacillus brevis, L. plantarum, L. paracasei and L. fermentum, respectively. The method facilitated well reproducible, medium-discrimination typing of Lactobacillus spp. and Pediococcus spp. at a subspecies level and proved to be suitable for preliminary typing of lactic acid bacteria isolated from cheese.
Similar content being viewed by others
Abbreviations
- LAB:
-
lactic acid bacteria
- MRS:
-
De Man-Rogosa-Sharpe (agar)
- PCR:
-
polymerase chain reaction
- RAMP:
-
randomly-amplified microsatellite polymorphism
- RAPD:
-
randomly-amplified polymorphic DNA
- SPE:
-
solid phase extraction
- UPGMA:
-
unweighted pair-group method with arithmetic averages
References
van Belkum A., Scherer S., VAN Alphen L., Verbrugh H.: Short-sequence DNA repeats in prokaryotic genomes. Microbiol.Mol. Biol.Rev.62, 275–293 (1998).
Boyd M.A., Antonio M.A.D., Hillier S.L.: Comparison of API 50 CH strips to whole-chromosomal DNA probes for identification of Lactobacillus species. J.Clin.Microbiol.43, 5309–5311 (2005).
de Angelis M., Corsetti A., Tosti N., Rossi J., Corbo M.R., Gobbetti M.: Characterization of non-starter lactic acid bacteria from Italian ewe cheeses based on phenotypic, genotypic, and cell wall protein analyses. Appl.Environ.Microbiol.67, 2011–2020 (2001).
de Candia S., DE Angelis M., Dunlea E., Minervini F., McSweeney P.L.H., Faccia M., Gobbetti M.: Molecular identification and typing of natural whey starter cultures and microbiological and compositional properties of related traditional Mozzarella cheeses. Internat.J.Food Microbiol.119, 182–191 (2007).
de León J.H., Jones W.A.: Detection of DNA polymorphisms in Homalodisca coagulata (Homoptera:Cicadellidae) by polymerase chain reaction-based DNA fingerprinting methods. Ann.Entomol.Soc.Am.97, 574–585 (2004).
Delfederico L., Hollmann A., Martínez M., Iglesias N.G., DE Antoni G., Semorile L.: Molecular identification and typing of lactobacilli isolated from kefir grains. J.Dairy Res.73, 20–27 (2006).
Fitzsimons N.A., Cogan T.M., Condon S., Bersford T.: Phenotypic and genotypic characterization of non-starter lactic acid bacteria in mature Cheddar cheese. Appl.Environ.Microbiol.65, 3418–3426 (1999).
Gevers D., Huys G., Swings J.: Applicability of rep-PCR fingerprinting for identification of Lactobacillus species. FEMS Microbiol. Lett.205, 31–36 (2001).
Giraffa G., Rossetti L., Neviani E.: An evaluation of chelex-based DNA purification protocols for the typing of lactic acid bacteria. J.Microbiol.Meth.42, 175–184 (2000).
Giraffa G., Andrighetto C., Antonello C., Gatti M., Lazzi C., Marcazzan G., Lombardi A., Neviani E.: Genotypic and phenotypic diversity of Lactobacillus delbrueckii subsp. lactis strains of dairy origin. Internat.J.Food Microbiol.91, 129–139 (2004).
Herman L., Heyndrickx M., Waes G.: Typing of Bacillus sporothermodurans and other Bacillus species isolated from milk by repetitive element sequence based PCR. Lett.Appl.Microbiol.26, 183–188 (1998).
Lane D.J.: 16S/23S rDNA sequencing, pp. 115–175 in E. Stackebrandt, M. Goodfellow (Eds): Nucleic Acid Techniques in Bacterial Systematics. John Wiley & Sons, Chichester 1991.
le Flèche P., Jacques I., Grayon M., AL Dahouk S., Bouchon P., Denoeud F., Nöckler K., Neubauer H., Guilloteau L.A., Vergnaud G.: Evaluation and selection of tandem repeat loci for a Brucella MLVA typing assay. BMC Microbiol.6, 9 (2006).
Li Y.C., Korol A.B., Fahima T., Nevo E.: Microsatellites within genes: structure, function and evolution. Mol.Biol.Evol.21, 991–1007 (2003).
López S., Mayo B.: Identification and characterization of homofermentative mesophilic Lactobacillus strains isolated from artisan starter-free cheeses. Lett.Appl.Microbiol.25, 233–238 (1997).
O’Leary W.: Practical Handbook of Microbiology. CRC Press, Boca Raton 1989.
Pangallo D., Kaclíková E., Kuchta T., Drahovská H.: Detection of Listeria monocytogenes by polymerase chain reaction oriented to inlB gene. New Microbiol.24, 333–339 (2001).
Pangallo D., Karpíšková R., Turňa J., Kuchta T.: Typing of food-borne Listeria monocytogenes by the optimized repetitive extragenic palindrome-based polymerase chain reaction (REP-PCR). New Microbiol.25, 449–454 (2002).
Rijpens N.P., Herman L.F.: Molecular methods for identification and detection of bacterial food pathogens. J.AOAC Internat.85, 984–995 (2002).
Sánchez I., Seseña S., Poveda J.M., Cabezas L., Palop L.: Genetic diversity, dynamics, and activity of Lactobacillus community involved in traditional processing of artisanal Manchego cheese. Internat.J.Food Microbiol.107, 265–273 (2006).
Švec P., Nováková D., Žáčková L., Kukletová M., Sedláček I.: Evaluation of (GTG)5-PCR for rapid identification of Streptococcus mutans. Antonie van Leeuwenhoek94, 573–579 (2008).
Torriani S., Zapparoli G., Dellaglio F.: Use of PCR-based methods for rapid differentiation of Lactobacillus delbrueckii subsp. bulgaricus and L. delbrueckii subsp. lactis. Appl.Environ.Microbiol.65, 4351–4356 (1999).
Versalovic J., Koeuth T., Lupski J.R.: Distribution of repetitive DNA sequences in eubacteria and application to fingerprinting of bacterial genomes. Nucl.Acids Res.19, 6823–6831 (1991).
Volpini A.C., DE Azeredo Passos V.M., Romanha A.J.: Attempt to differentiate Leishmania (Leishmania) amazonensis, L. (L.) chagasi, Leishmania (Viannia) braziliensis and L. (V.) guyanensis using the SSR-PCR technique. Parasitol.Res.87, 1056–1059 (2001).
Wu K.S., Jones R., Danneberger L., Scolnik P.A.: Detection of microsatellite polymorphisms without cloning. Nucl.Acids Res.22, 3257–3258 (1994)
Zapparoli G., Torriani S., Dellaglio F.: Differentiation of Lactobacillus sanfranciscensis strains by randomly amplified polymorphic DNA and pulsed-field gel electrophoresis. FEMS Microbiol.Lett.166, 325–332 (1998).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Chebeňová, V., Berta, G., Kuchta, T. et al. Randomly-amplified microsatellite polymorphism for preliminary typing of lactic acid bacteria from Bryndza Cheese. Folia Microbiol 55, 598–602 (2010). https://doi.org/10.1007/s12223-010-0096-4
Received:
Revised:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12223-010-0096-4