Abstract
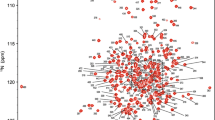
Cyclophilins regulate protein folding, transport and signalling through catalysis of proline isomerization, and are ubiquitously expressed in both prokaryotes and eukaryotes. Cpr3 is the yeast mitochondrial cyclophilin and it is structurally and biophysically uncharacterized so far. Yeast cyclophilin gene cpr3 is essential for the lactate metabolism. Here, we report 1H, 13C, and 15N chemical shift assignments of Cpr3 protein determined by various 2D and 3D heteronuclear NMR experiments at pH 6.5, and temperature 298 K.


Similar content being viewed by others
References
Alfano C, Babon J, Kelly G, Curry S, Conte MR (2003) Resonance assignment and secondary structure of an N-terminal fragment of the human La protein. J Biomol NMR 27:93–94
Camilloni C, Sahakyan AB, Holliday MJ, Isern NG, Zhang F, Eisenmesser EZ, Vendruscolo M (2014) Cyclophilin A catalyzes proline isomerization by an electrostatic handle mechanism. Proc Natl Acad Sci U S A. 111(28):10203–10208
Chi CN, Vögeli B, Bibow S, Strotz D, Orts J, Güntert P, Riek R (2015) A Structural ensemble for the enzyme cyclophilin reveals an orchestrated mode of action at atomic resolution. Angew Chem 54(40):11657–11661
Cornilescu G, Delaglio F, Bax A (1999) Protein backbone angle restraints from searching a database for chemical shift and sequence homology. J Biomol NMR 13:289–302
Davis ES, Becker A, Heitman J, Hall MN, Brennan MB (1992) A yeast cyclophilin gene essential for lactate metabolism at high temperature. Proc Natl Acad Sci U S A 89(23):11169–11173
Eisenmesser EZ, Bosco DA, Akke M, Kern D (2002) Enzyme dynamics during catalysis. Science 295(5559):1520
Keller R (2004) The computer aided resonance assignment tutorial CANTINA. Verlag, Goldau, Switzerland
Kofron JL, Kuzmic P, Kishore V, Colón-Bonilla E, Rich DH (1991) Determination of kinetic constants for peptidyl prolyl cis-trans isomerases by an improved spectrophotometric assay. Biochemistry 30(25):6127–6134
Nigro P, Pompilio G, Capogrossi MC (2013) Cyclophilin A: a key player for human disease. Cell Death Dis 4:e888
Shen Y, Delaglio F, Cornilescu G, Bax A (2009) TALOS+ : a hybrid method for predicting protein backbone torsion angles from NMR chemical shifts. J Biomol NMR 44:213–223
Wang P, Heitman J (2005) The cyclophilins. Genome Biol 6(7):226
Zhao Y, Ke H (1996) Mechanistic implication of crystal structures of the cyclophilin-dipeptide complexes. Biochemistry 35(23):7362–7368
Zhao Y, Chen Y, Schutkowski M, Fischer G, Ke H (1997) Cyclophilin A complexed with afragment of HIV-1 gag protein: insights into HIV-1 infectious activity. Structure 5(1):139–146
Acknowledgments
We are thankful to RIFC, IRCC, NMR facility (750 MHz) at IIT Bombay.V.K.S is the recipients of research fellowships from DAE, Mumbai, India. J.S.S and D.T. are recipients of research fellowships from MHRD, IIT Bombay, Mumbai, India.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Shukla, V.K., Singh, J.S., Trivedi, D. et al. NMR assignments of mitochondrial cyclophilin Cpr3 from Saccharomyces cerevisiae . Biomol NMR Assign 10, 203–206 (2016). https://doi.org/10.1007/s12104-016-9667-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12104-016-9667-x