Abstract
Background
It is well documented that over-expression of thec-myc proto-oncogene occurs in the vast majority of mouse thymic lymphomas induced by γ-irradiation, evidencing the importance of this gene in T-cell lymphomagenesis. However, it remains unknown whether elevated levels ofc-myc expression are driven by extrac-myc copy numbers.
Materials and methods
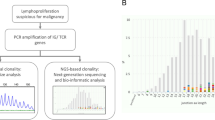
Here we use a quantitative test on the basis of real-time PCR to determine the cellular copy number of c-myc in a set of 14 g-radiation-induced thymic lymphomas obtained from (C57BL/6J x BALB/cJ) F1 hybrid mice with increased mRNA c-myc expression.
Results
Since 5 out of 14 (35.7%) cases had no extra copy numbers of c-myc, gene amplification was obviously not the cause of c-myc over-expression in these tumours. In the remaining 9 tumours, c-myc over-expression was also accompanied with extra DNA copy numbers. Therefore, c-myc amplification might be a consequence of the genomic instability subsequent to the up-regulation of c-myc. However, linear regression analysis showed a lack of correlation between increasing DNA copy numbers and mRNA over expression of c-myc in these tumours (r =0.029, p=0.94).
Conclusion
De-regulation of c-myc does not necessarity imply amplification of this gene in these tumours. This report is, to our knowledge, the first one comparing c-myc amplification with expression in lymphomas of the T-cell lineage.
Similar content being viewed by others
References
Vennstrom B, Sheiness D, Zabielski J, Bishop JM. Isolation and characterization ofc-myc, a cellular homolog of the oncogene (v-myc) of avian myelocytomatosis virus strain 29. J Virol. 1982;42:773–9.
Dalla-Favera R, Bregni M, Erikson J, Patterson D, Gallo RC, Croce CM. Humanc-myc onc gene is located on the region of chromosome 8 that is translocated in Burkitt lymphoma cells. Proc Natl Acad Sci USA. 1982;79:7824–7.
Spencer CA, Groudine M. Control ofc-myc regulation in normal and neoplastic cells. Adv Cancer Res. 1991;56:1–48.
McMorrow LE, Newcomb EW, Pellicer A. Identification of a specific marker chromosome early in tumour development in gamma-irradiated C57BL/6J mice. Leukemia. 1988;2:115–9.
Muto M, Chen Y, Kubo E, Mita K. Analysis of early initiating event(s) in radiation-induced thymic lymphomagenesis. Jpn J Cancer Res. 1996;87:247–57.
Vasmel WL, Matthews EA, Gillis CP, et al. Distinct chromosomal abnormalities in murine leukemia virus-induced T- and B-cell lymphomas. Int J Cancer. 1989;43:1112–9.
Gaudet F, Hodgson JG, Eden A, et al. Induction of tumours in mice by genomic hypomethylation. Science. 2003;300:489–92.
Liyanage M, Weaver Z, Barlow C, et al. Abnormal rearrangement within the alpha/delta T-cell receptor locus in lymphomas fromAtm-deficient mice. Blood. 2000;96:1940–6.
Silva S, Babonits M, Wiener F, Klein G. Further studies on chromosome 15 trisomy in murine T-cell lymphomas: mapping of the relevant chromosome segment. Int J Cancer. 1988;41:738–43.
Banerjee M, Wiener F, Spira J, et al. Mapping of thec-myc, pvt-1 and immunoglobulin kappa genes in relation to the mouse plasmacytoma-associated variant (6;15) translocation breakpoint. EMBO J. 1985;4:3183–8.
López-Nieva P, Santos J, Fernández-Piqueras J. Altered expression ofNotch1, Notch2, c-myc andIkaros in γ-radiation induced mouse thymic lymphomas. Carcinogenesis, 2004;25:1299–304.
Stewart M, Cameron E, Campbell M, et al. conditional expression and oncogenicity ofc-myc linked to a CD2 gene dominant control region. Int J Cancer. 1993;53:1023–30.
Higuchi R, Fockler C, Dollinger G, Watson R. Kinetic PCR analysis: real-time monitoring of DNA amplification reactions. Biotechnology. 1993;11:1026–30.
Mocellin S, Rossi CR, Pilati P, Nitti D, Marincola FM. Quantitative real-time PCR: a powerful ally in cancer research. Trends Mol Med. 2003;9:189–95.
Santos J, Pérez de Castro I, Herranz M, Pellicer A, Fernández-Piqueras J. Allelic losses on chromosome 4 suggest the existence of a candidate tumour suppressor gene region of about 0.6 cM in gamma-radiation-induced mouse primary thymic lymphomas. Oncogene. 1996;12:669–76.
Wittwer CT, Herrmann MG, Moss AA, Rasmussen RP. Continuous fluorescence monitoring of rapid cycle DNA amplification. Biotechniques. 1997;22:130–9.
Santos J, Herranz M, Pérez de Castro I, Pellicer A, Fernández-Piqueras J. A new candidate site for a tumour suppressor gene involved in mouse thymic lymphomagenesis is located on the distal part of chromosome 4. Oncogene. 1998;17:925–9.
Meléndez B, Santos J, Fernández-Piqueras J. Loss of heterozygosity at the proximal-mid part of mouse chromosome 4 defines two novel tumour suppressor gene loci in T-cell lymphomas. Oncogene. 1999;18:4166–9.
Bieche I, Olivi M, Champeme MH, Vidaud D, Lidereau R, Vidaud M. Novel approach to quantitative polymerase chain reaction using real-time detection: application to the detection of gene amplification in breast cancer. Int J Cancer. 1998;78:661–6.
Rochlitz CF, Herrmann R, de Kant E. Over-expression and amplification ofc-myc during progression of human colorectal cancer. Oncology, 1996;53:448–54.
Herms J, Neidt I, Luscher B, et al.c-myc expression in medulloblastoma and its pronostic value. Int J Cancer. 2000;89: 395–402.
Rothberg PG, Erisman MD, Diehl RE, Rovigatti UG, Astrin SM. Structure and expression of the oncogenec-myc in fresh tumour material from patients with hematopoietic malignancies. Mol Cell Biol. 1984;4:1096–103.
Green DR, Evan GI. A matter of life and death. Cancer Cell. 2002;1:19–30
Gray JW, Collins C. Genome changes and gene expression in human solid tumours. Carcinogenesis. 2000;21:443–52.
Marinkovic D, Marinkovic T, Mahr B, Hess J, Wirth T. Reversible lymphomagenesis in conditionallyc-myc expressing mice. Int J Cancer. 2004;20:336–42.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Santos, J., Vaquero, C., Reyes, J. et al. Lack of correlation between DNA copy number and mRNA expression levels ofc-myc in γ-radiation-induced mouse thymic lymphomas by using quantitative real-time PCR. Clin Transl Oncol 8, 349–353 (2006). https://doi.org/10.1007/s12094-006-0181-y
Received:
Revised:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/s12094-006-0181-y