Abstract
piRNAs (PIWI-interacting RNAs) are a class of small non-coding RNAs (ncRNAs) abundantly expressed in germline cells and involved in suppressing the transposon activity. Interestingly, recent studies have found piRNA expression in the central nervous system (CNS), yet the underlying biological significance remains largely unknown. In this study, we investigated the expression and function of piRNAs during the retinoic acid (RA)-mediated neuronal differentiation in NT2 cells, a human embryonal carcinoma cell line. We identified a cohort of differentially expressed piRNAs by microarray. Two piRNAs, DQ582359 and DQ596268, were increasingly upregulated during the RA-induced differentiation and involved in regulating the expression of neuronal markers, MAP2 and TUBB3. Furthermore, these piRNAs were found to associate with cold-shock domain (CSD)-containing RNA binding proteins, DIS3, DIS3L2, and YB-1. Markedly, overexpression of these piRNAs further enhanced the protein levels of MAP2 and TUBB3, potentially by downregulating DIS3, DIS3L2, and YB-1. Hence, our study has identified a novel somatic function of piRNAs in regulating neuronal gene expression. The interaction of piRNA with some CSD-containing proteins can be further explored to enhance neuronal differentiation to treat neurodegenerative diseases.







Similar content being viewed by others
Availability of Data and Material
Data will be made available upon request.
Code Availability
Not applicable.
References
Lunn JS, Sakowski SA, Hur J, Feldman EL (2011) Stem cell technology for neurodegenerative diseases. Ann Neurol 70(3):353–361. https://doi.org/10.1002/ana.22487
Barker RA, Drouin-Ouellet J, Parmar M (2015) Cell-based therapies for Parkinson disease-past insights and future potential. Nat Rev Neurol 11(9):492–503. https://doi.org/10.1038/nrneurol.2015.123
Subhramanyam CS, Hu Q (2017) Non-coding RNA in brain development and disorder. Curr Med Chem 24(18):1983–1997. https://doi.org/10.2174/0929867324666170124151436
Siomi MC, Sato K, Pezic D, Aravin AA (2011) PIWI-interacting small RNAs: the vanguard of genome defence. Nat Rev Mol Cell Biol 12(4):246–258. https://doi.org/10.1038/nrm3089
Aravin A, Gaidatzis D, Pfeffer S, Lagos-Quintana M, Landgraf P, Iovino N, Morris P, Brownstein MJ, Kuramochi-Miyagawa S, Nakano T, Chien M, Russo JJ, Ju J, Sheridan R, Sander C, Zavolan M, Tuschl T (2006) A novel class of small RNAs bind to MILI protein in mouse testes. Nature 442(7099):203–207. https://doi.org/10.1038/nature04916
Girard A, Sachidanandam R, Hannon GJ, Carmell MA (2006) A germline-specific class of small RNAs binds mammalian Piwi proteins. Nature 442(7099):199–202. https://doi.org/10.1038/nature04917
Grivna ST, Beyret E, Wang Z, Lin H (2006) A novel class of small RNAs in mouse spermatogenic cells. Genes Dev 20(13):1709–1714. https://doi.org/10.1101/gad.1434406
Lau NC, Seto AG, Kim J, Kuramochi-Miyagawa S, Nakano T, Bartel DP, Kingston RE (2006) Characterization of the piRNA complex from rat testes. Science 313(5785):363–367. https://doi.org/10.1126/science.1130164
Aravin AA, Sachidanandam R, Bourc’his D, Schaefer C, Pezic D, Toth KF, Bestor T, Hannon GJ (2008) A piRNA pathway primed by individual transposons is linked to de novo DNA methylation in mice. Mol Cell 31(6):785–799. https://doi.org/10.1016/j.molcel.2008.09.003
Huang XA, Yin H, Sweeney S, Raha D, Snyder M, Lin H (2013) A major epigenetic programming mechanism guided by piRNAs. Dev Cell 24(5):502–516. https://doi.org/10.1016/j.devcel.2013.01.023
Carmell MA, Girard A, van de Kant HJ, Bourc’his D, Bestor TH, de Rooij DG, Hannon GJ (2007) MIWI2 is essential for spermatogenesis and repression of transposons in the mouse male germline. Dev Cell 12(4):503–514. https://doi.org/10.1016/j.devcel.2007.03.001
Kuramochi-Miyagawa S, Watanabe T, Gotoh K, Totoki Y, Toyoda A, Ikawa M, Asada N, Kojima K, Yamaguchi Y, Ijiri TW, Hata K, Li E, Matsuda Y, Kimura T, Okabe M, Sakaki Y, Sasaki H, Nakano T (2008) DNA methylation of retrotransposon genes is regulated by Piwi family members MILI and MIWI2 in murine fetal testes. Genes Dev 22(7):908–917. https://doi.org/10.1101/gad.1640708
Pezic D, Manakov SA, Sachidanandam R, Aravin AA (2014) piRNA pathway targets active LINE1 elements to establish the repressive H3K9me3 mark in germ cells. Genes Dev 28(13):1410–1428. https://doi.org/10.1101/gad.240895.114
Lau NC, Robine N, Martin R, Chung WJ, Niki Y, Berezikov E, Lai EC (2009) Abundant primary piRNAs, endo-siRNAs, and microRNAs in a Drosophila ovary cell line. Genome Res 19(10):1776–1785. https://doi.org/10.1101/gr.094896.109
He X, Chen X, Zhang X, Duan X, Pan T, Hu Q, Zhang Y, Zhong F, Liu J, Zhang H, Luo J, Wu K, Peng G, Luo H, Zhang L, Li X, Zhang H (2015) An Lnc RNA (GAS5)/SnoRNA-derived piRNA induces activation of TRAIL gene by site-specifically recruiting MLL/COMPASS-like complexes. Nucleic Acids Res 43(7):3712–3725. https://doi.org/10.1093/nar/gkv214
Zhong F, Zhou N, Wu K, Guo Y, Tan W, Zhang H, Zhang X, Geng G, Pan T, Luo H, Zhang Y, Xu Z, Liu J, Liu B, Gao W, Liu C, Ren L, Li J, Zhou J, Zhang H (2015) A SnoRNA-derived piRNA interacts with human interleukin-4 pre-mRNA and induces its decay in nuclear exosomes. Nucleic Acids Res 43(21):10474–10491. https://doi.org/10.1093/nar/gkv954
Qiu W, Guo X, Lin X, Yang Q, Zhang W, Zhang Y, Zuo L, Zhu Y, Li CR, Ma C, Luo X (2017) Transcriptome-wide piRNA profiling in human brains of Alzheimer’s disease. Neurobiol Aging 57:170–177. https://doi.org/10.1016/j.neurobiolaging.2017.05.020
Lee EJ, Banerjee S, Zhou H, Jammalamadaka A, Arcila M, Manjunath BS, Kosik KS (2011) Identification of piRNAs in the central nervous system. RNA 17(6):1090–1099. https://doi.org/10.1261/rna.2565011
Rajasethupathy P, Antonov I, Sheridan R, Frey S, Sander C, Tuschl T, Kandel ER (2012) A role for neuronal piRNAs in the epigenetic control of memory-related synaptic plasticity. Cell 149(3):693–707. https://doi.org/10.1016/j.cell.2012.02.057
Ghosheh Y, Seridi L, Ryu T, Takahashi H, Orlando V, Carninci P, Ravasi T (2016) Characterization of piRNAs across postnatal development in mouse brain. Sci Rep 6:25039. https://doi.org/10.1038/srep25039
Phay M, Kim HH, Yoo S (2018) Analysis of piRNA-like small non-coding RNAs present in axons of adult sensory neurons. Mol Neurobiol 55(1):483–494. https://doi.org/10.1007/s12035-016-0340-2
Gonzalez J, Qi H, Liu N, Lin H (2015) Piwi is a key regulator of both somatic and germline stem cells in the Drosophila testis. Cell Rep 12(1):150–161. https://doi.org/10.1016/j.celrep.2015.06.004
Subhramanyam CS, Cao Q, Wang C, Heng ZSL, Zhou Z, Hu Q (2020) Role of PIWI-like 4 in modulating neuronal differentiation from human embryonal carcinoma cells. RNA Biol 17(11):1613–1624. https://doi.org/10.1080/15476286.2020.1757896
Mei Y, Wang Y, Kumari P, Shetty AC, Clark D, Gable T, MacKerell AD, Ma MZ, Weber DJ, Yang AJ, Edelman MJ, Mao L (2015) A piRNA-like small RNA interacts with and modulates p-ERM proteins in human somatic cells. Nat Commun 6:7316. https://doi.org/10.1038/ncomms8316
Ju Lee H, Bartsch D, Xiao C, Guerrero S, Ahuja G, Schindler C, Moresco JJ, Yates JR 3rd, Gebauer F, Bazzi H, Dieterich C, Kurian L, Vilchez D (2017) A post-transcriptional program coordinated by CSDE1 prevents intrinsic neural differentiation of human embryonic stem cells. Nat Commun 8(1):1456. https://doi.org/10.1038/s41467-017-01744-5
Faehnle CR, Walleshauser J, Joshua-Tor L (2014) Mechanism of Dis3l2 substrate recognition in the Lin28-let-7 pathway. Nature 514(7521):252–256. https://doi.org/10.1038/nature13553
Fotovati A, Abu-Ali S, Wang PS, Deleyrolle LP, Lee C, Triscott J, Chen JY, Franciosi S, Nakamura Y, Sugita Y, Uchiumi T, Kuwano M, Leavitt BR, Singh SK, Jury A, Jones C, Wakimoto H, Reynolds BA, Pallen CJ, Dunn SE (2011) YB-1 bridges neural stem cells and brain tumor-initiating cells via its roles in differentiation and cell growth. Cancer Res 71(16):5569–5578. https://doi.org/10.1158/0008-5472.CAN-10-2805
Segalla S, Pivetti S, Todoerti K, Chudzik MA, Giuliani EC, Lazzaro F, Volta V, Lazarevic D, Musco G, Muzi-Falconi M, Neri A, Biffo S, Tonon G (2015) The ribonuclease DIS3 promotes let-7 miRNA maturation by degrading the pluripotency factor LIN28B mRNA. Nucleic Acids Res 43(10):5182–5193. https://doi.org/10.1093/nar/gkv387
Ustianenko D, Hrossova D, Potesil D, Chalupnikova K, Hrazdilova K, Pachernik J, Cetkovska K, Uldrijan S, Zdrahal Z, Vanacova S (2013) Mammalian DIS3L2 exoribonuclease targets the uridylated precursors of let-7 miRNAs. RNA 19(12):1632–1638. https://doi.org/10.1261/rna.040055.113
Mihailovich M, Militti C, Gabaldon T, Gebauer F (2010) Eukaryotic cold shock domain proteins: highly versatile regulators of gene expression. BioEssays 32(2):109–118. https://doi.org/10.1002/bies.200900122
Schneider C, Anderson JT, Tollervey D (2007) The exosome subunit Rrp44 plays a direct role in RNA substrate recognition. Mol Cell 27(2):324–331. https://doi.org/10.1016/j.molcel.2007.06.006
Malecki M, Viegas SC, Carneiro T, Golik P, Dressaire C, Ferreira MG, Arraiano CM (2013) The exoribonuclease Dis3L2 defines a novel eukaryotic RNA degradation pathway. EMBO J 32(13):1842–1854. https://doi.org/10.1038/emboj.2013.63
Izumi H, Imamura T, Nagatani G, Ise T, Murakami T, Uramoto H, Torigoe T, Ishiguchi H, Yoshida Y, Nomoto M, Okamoto T, Uchiumi T, Kuwano M, Funa K, Kohno K (2001) Y box-binding protein-1 binds preferentially to single-stranded nucleic acids and exhibits 3’–>5’ exonuclease activity. Nucleic Acids Res 29(5):1200–1207
Przyborski SA, Morton IE, Wood A, Andrews PW (2000) Developmental regulation of neurogenesis in the pluripotent human embryonal carcinoma cell line NTERA-2. Eur J Neurosci 12(10):3521–3528
Pleasure SJ, Page C, Lee VM (1992) Pure, postmitotic, polarized human neurons derived from NTera 2 cells provide a system for expressing exogenous proteins in terminally differentiated neurons. J Neurosci 12(5):1802–1815
Pleasure SJ, Lee VM (1993) NTera 2 cells: a human cell line which displays characteristics expected of a human committed neuronal progenitor cell. J Neurosci Res 35(6):585–602. https://doi.org/10.1002/jnr.490350603
Kirino Y, Mourelatos Z (2007) Mouse Piwi-interacting RNAs are 2’-O-methylated at their 3’ termini. Nat Struct Mol Biol 14(4):347–348. https://doi.org/10.1038/nsmb1218
Ohara T, Sakaguchi Y, Suzuki T, Ueda H, Miyauchi K, Suzuki T (2007) The 3’ termini of mouse Piwi-interacting RNAs are 2’-O-methylated. Nat Struct Mol Biol 14(4):349–350. https://doi.org/10.1038/nsmb1220
Tian Y, Simanshu DK, Ma JB, Patel DJ (2011) Structural basis for piRNA 2’-O-methylated 3’-end recognition by Piwi PAZ (Piwi/Argonaute/Zwille) domains. Proc Natl Acad Sci U S A 108(3):903–910. https://doi.org/10.1073/pnas.1017762108
Dong ZW, Shao P, Diao LT, Zhou H, Yu CH, Qu LH (2012) RTL-P: a sensitive approach for detecting sites of 2’-O-methylation in RNA molecules. Nucleic Acids Res 40(20):e157. https://doi.org/10.1093/nar/gks698
Smith B, Treadwell J, Zhang D, Ly D, McKinnell I, Walker PR, Sikorska M (2010) Large-scale expression analysis reveals distinct microRNA profiles at different stages of human neurodevelopment. PLoS One 5(6):e11109
Tomecki R, Kristiansen MS, Lykke-Andersen S, Chlebowski A, Larsen KM, Szczesny RJ, Drazkowska K, Pastula A, Andersen JS, Stepien PP, Dziembowski A, Jensen TH (2010) The human core exosome interacts with differentially localized processive RNases: hDIS3 and hDIS3L. EMBO J 29(14):2342–2357. https://doi.org/10.1038/emboj.2010.121
Lubas M, Damgaard CK, Tomecki R, Cysewski D, Jensen TH, Dziembowski A (2013) Exonuclease hDIS3L2 specifies an exosome-independent 3’-5’ degradation pathway of human cytoplasmic mRNA. EMBO J 32(13):1855–1868. https://doi.org/10.1038/emboj.2013.135
Labno A, Tomecki R (1863) Dziembowski A (2016) Cytoplasmic RNA decay pathways - enzymes and mechanisms. Biochim Biophys Acta 12:3125–3147. https://doi.org/10.1016/j.bbamcr.2016.09.023
Kiss DL, Andrulis ED (2010) Genome-wide analysis reveals distinct substrate specificities of Rrp6, Dis3, and core exosome subunits. RNA 16(4):781–791. https://doi.org/10.1261/rna.1906710
Przyborski SA, Smith S, Wood A (2003) Transcriptional profiling of neuronal differentiation by human embryonal carcinoma stem cells in vitro. Stem Cells 21(4):459–471. https://doi.org/10.1634/stemcells.21-4-459
Haile Y, Fu W, Shi B, Westaway D, Baker G, Jhamandas J, Giuliani F (2014) Characterization of the NT2-derived neuronal and astrocytic cell lines as alternative in vitro models for primary human neurons and astrocytes. J Neurosci Res 92(9):1187–1198. https://doi.org/10.1002/jnr.23399
Rouget C, Papin C, Boureux A, Meunier AC, Franco B, Robine N, Lai EC, Pelisson A, Simonelig M (2010) Maternal mRNA deadenylation and decay by the piRNA pathway in the early Drosophila embryo. Nature 467(7319):1128–1132. https://doi.org/10.1038/nature09465
Dziembowski A, Lorentzen E, Conti E, Séraphin B (2007) A single subunit, Dis3, is essentially responsible for yeast exosome core activity. Nat Struct Mol Biol 14(1):15–22. https://doi.org/10.1038/nsmb1184
Schaeffer D, Tsanova B, Barbas A, Reis FP, Dastidar EG, Sanchez-Rotunno M, Arraiano CM, van Hoof A (2009) The exosome contains domains with specific endoribonuclease, exoribonuclease and cytoplasmic mRNA decay activities. Nat Struct Mol Biol 16(1):56–62. https://doi.org/10.1038/nsmb.1528
Hou D, Ruiz M, Andrulis ED (2012) The ribonuclease Dis3 is an essential regulator of the developmental transcriptome. BMC Genomics 13:359. https://doi.org/10.1186/1471-2164-13-359
Chang HM, Triboulet R, Thornton JE, Gregory RI (2013) A role for the Perlman syndrome exonuclease Dis3l2 in the Lin28-let-7 pathway. Nature 497(7448):244–248. https://doi.org/10.1038/nature12119
Mori F, Tanji K, Miki Y, Toyoshima Y, Sasaki H, Yoshida M, Kakita A, Takahashi H, Wakabayashi K (2018) Immunohistochemical localization of exoribonucleases (DIS3L2 and XRN1) in intranuclear inclusion body disease. Neurosci Lett 662:389–394. https://doi.org/10.1016/j.neulet.2017.10.061
Pashler AL, Towler BP, Jones CI, Newbury SF (2016) The roles of the exoribonucleases DIS3L2 and XRN1 in human disease. Biochem Soc Trans 44(5):1377–1384. https://doi.org/10.1042/BST20160107
Tong LM, Fong H, Huang Y (2015) Stem cell therapy for Alzheimer’s disease and related disorders: current status and future perspectives. Exp Mol Med 47:e151. https://doi.org/10.1038/emm.2014.124
Kikuchi T, Morizane A, Doi D, Magotani H, Onoe H, Hayashi T, Mizuma H, Takara S, Takahashi R, Inoue H, Morita S, Yamamoto M, Okita K, Nakagawa M, Parmar M, Takahashi J (2017) Human iPS cell-derived dopaminergic neurons function in a primate Parkinson’s disease model. Nature 548(7669):592–596. https://doi.org/10.1038/nature23664
Acknowledgements
We would like to thank Ms. Michelle Mok and Ms. Wang Xianhui (Department of Biological Sciences, National University of Singapore (NUS)) for their assistance in the mass spectrometry analysis; Dr. Deepan Balakrishnan (Department of Biological Sciences, NUS) for his help in generating the heat map; and Dr. Stepanka Vanacova (Central European Institute of Technology, Masaryk University, Czech Republic) for the kind gift of the DIS3L2 overexpression plasmid. We also want to thank Mr. Sylvester Wong Shu Ming for helping with imaging and bioinformatics support and Mr. Lal Zo Thanga for administrative assistance. The GTEx data described in this manuscript were obtained from the GTEx Portal (https://www.gtexportal.org/home/gene/LINC01015; https://www.gtexportal.org/home/gene/LINC00905), dbGaP accession number phs000424.v8.p2. The Genotype-Tissue Expression (GTEx) Project was supported by the Common Fund of the Office of the Director of the National Institutes of Health, and by NCI, NHGRI, NHLBI, NIDA, NIMH, and NINDS.
Funding
This study was supported by the National University of Singapore Start-up Grant R-181–000-155–133, Ministry of Education Tier 1 Grant R-181–000-179–114, and National University Health System Seed Fund R-181–000-192–114. C.S.S. and C.W. were supported by the NUS Postgraduate Scholarships.
Author information
Authors and Affiliations
Contributions
C.S.S. performed most of the experiments in the study. Q.C., C.W., Z.S.L.H., and Z.Z. provided technical assistance in cell culture, Northern blotting, and imaging. Q.H. initiated and managed the project. C.S.S. and Q.H. analyzed the data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
12035_2021_2678_MOESM1_ESM.pdf
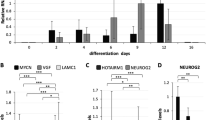
Supplementary file1 Supplementary Fig. 1 Profiling of RA-mediated neuronal gene expression in NT2 cells. qPCR was performed to show the suppression of ESC markers (a) and the induction of neuronal markers (b) at different time points of RA treatment. d: days of RA treatment. Bars: mean · SD; n=3. (c) Western blot was performed to profile the protein levels of ESC and neuronal markers at different time points of RA treatment. (PDF 139 KB)
12035_2021_2678_MOESM2_ESM.pdf
Supplementary file2 Supplementary Fig. 2 Immunofluorescent staining of MAP2 in 15-day RA-treated NT2 cells. NT2 cells were treated with RA for 15 days and then immunostained for MAP2. The cells were counterstained with Hoechst 33342 dye. Scale bar: 5µm. (PDF 96 KB)
12035_2021_2678_MOESM3_ESM.pdf
Supplementary file3 Supplementary Fig. 3 Profiling of piRNA expression in non-treated and RA-treated NT2 cells. (a) Box plot shows the normalized intensities of piRNA array data. (b) Scatter plot shows the differentially expressed piRNAs between indicated samples. (c) Bar graphs illustrate the number of upregulated and downregulated piRNAs from comparisons between non-treated and RA-treated cells as indicated. Ctrl: non-treated cells; 6dRA: 6 days of RA treatment; 15dRA: 15 days of RA treatment; DE: differentially expressed. (PDF 140 KB)
12035_2021_2678_MOESM4_ESM.pdf
Supplementary file4 Supplementary Fig. 4 Interaction of piRNAs with PIWI protein. HA-tagged PIWIL4 protein was overexpressed in 293T cells and piRNA mimics were used to pull down the interacting protein. Western blot with anti-HA antibody shows the association of piR1 and piR2 with PIWIL4. C: control Vector; PL4: HA-tagged PIWIL4 overexpression; piR1: pull-down sample with piR1 probe; piR2: pull-down sample with piR2 probe; piNC: pull-down sample with the scrambled sequence probe. (PDF 94 KB)
12035_2021_2678_MOESM5_ESM.pdf
Supplementary file5 Supplementary Fig. 5 Secondary antibody control for immunofluorescent staining in NT2 cells. The cells were immunostained with anti-Rabbit Alexa 555 secondary antibody with or without pre-labeling with rabbit anti-YB1 antibody, or with anti-mouse Alexa 546 secondary antibody with or without pre-labeling with mouse anti-DIS3 antibody. Scale bar: 20µm. (PDF 109 KB)
12035_2021_2678_MOESM6_ESM.pdf
Supplementary file6 Supplementary Fig. 6 DIS3L2 overexpression affects piRNA-mediated neuronal gene expression. piR1 was concurrently overexpressed along with its interacting partner, DIS3L2 in RA-treated NT2 cells. The expression levels of MAP2 and DIS3L2 were then assessed by Western blot. Ctrl: control Vector; DIS3L2: pEGFP-Flag-HA-DIS3L2 overexpression plasmid; piNC: scrambled sequence. (PDF 93 KB)
Rights and permissions
About this article
Cite this article
Subhramanyam, C.S., Cao, Q., Wang, C. et al. piRNAs Interact with Cold-Shock Domain-Containing RNA Binding Proteins and Regulate Neuronal Gene Expression During Differentiation. Mol Neurobiol 59, 1285–1300 (2022). https://doi.org/10.1007/s12035-021-02678-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12035-021-02678-2