Abstract
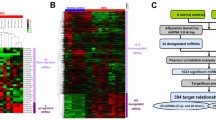
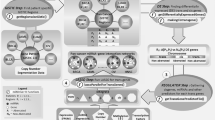
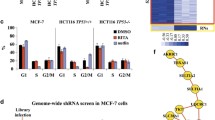
Novel computational systems biology tools such as common targets analysis, common regulators analysis, pathway discovery, and transcriptomic-based hotspot discovery provide new opportunities in understanding of apoptosis molecular mechanisms. In this study, after measuring the global contribution of microRNAs in the course of apoptosis by Affymetrix platform, systems biology tools were utilized to obtain a comprehensive view on the role of microRNAs in apoptosis process. Network analysis and pathway discovery highlighted the crosstalk between transcription factors and microRNAs in apoptosis. Within the transcription factors, PRDM1 showed the highest upregulation during the course of apoptosis, with more than 9-fold expression increase compared to non-apoptotic condition. Within the microRNAs, MIR1208 showed the highest expression in non-apoptotic condition and downregulated by more than 6 fold during apoptosis. Common regulators algorithm showed that TNF receptor is the key upstream regulator with a high number of regulatory interactions with the differentially expressed microRNAs. BCL2 and AKT1 were the key downstream targets of differentially expressed microRNAs. Enrichment analysis of the genomic locations of differentially expressed microRNAs led us to the discovery of chromosome bands which were highly enriched (p < 0.01) with the apoptosis-related microRNAs, such as 13q31.3, 19p13.13, and Xq27.3 This study opens a new avenue in understanding regulatory mechanisms and downstream functions in the course of apoptosis as well as distinguishing genomic-enriched hotspots for apoptosis process.





Similar content being viewed by others
References
Agarwal, V., Bell, G. W., Nam, J.-W., & Bartel, D. P. (2015). Predicting effective microRNA target sites in mammalian mRNAs. Elife, 4, e05005.
Alanazi, I., Ebrahimie, E., Hoffmann, P., & Adelson, D. L. (2013). Combined gene expression and proteomic analysis of EGF induced apoptosis in A431 cells suggests multiple pathways trigger apoptosis. Apoptosis, 18, 1291–1305.
Alanazi, I., Hoffmann, P., & Adelson, D. L. (2015). MicroRNAs are part of the regulatory network that controls EGF induced apoptosis, including elements of the JAK/STAT pathway, in A431 Cells. PloS one, 10, e0120337.
Alisoltani, A., Fallahi, H., Ebrahimi, M., Ebrahimi, M., & Ebrahimie, E. (2014). Prediction of potential cancer-risk regions based on transcriptome data: Towards a comprehensive view. PLoS One, 9, e96320.
Alisoltani, A., Shiran, B., Fallahi, H., & Ebrahimie, E. (2015). Gene regulatory network in almond (Prunus dulcis Mill.) in response to frost stress. Tree Genetics & Genomes, 11, 1–15.
Bakhtiarizadeh, M. R., Moradi-Shahrbabak, M., & Ebrahimie, E. (2013). Underlying functional genomics of fat deposition in adipose tissue. Gene, 521, 122–128.
Bakhtiarizadeh, M. R., Moradi-Shahrbabak, M., & Ebrahimie, E. (2014). Transcriptional regulatory network analysis of the over-expressed genes in adipose tissue. Genes & Genomics, 36, 105–117.
Bolton, K. L., Tyrer, J., Song, H., Ramus, S. J., Notaridou, M., Jones, C., et al. (2010). Common variants at 19p13 are associated with susceptibility to ovarian cancer. Nature Genetics, 42, 880–884.
Brabletz, S., & Brabletz, T. (2010). The ZEB/miR-200 feedback loop—a motor of cellular plasticity in development and cancer? EMBO Reports, 11, 670–677.
Chang, T.-C., Zeitels, L. R., Hwang, H.-W., Chivukula, R. R., Wentzel, E. A., Dews, M., et al. (2009). Lin-28B transactivation is necessary for Myc-mediated let-7 repression and proliferation. Proceedings of the National Academy of Sciences, 106, 3384–3389.
Ebrahimie, M., Esmaeili, F., Cheraghi, S., Houshmand, F., Shabani, L., & Ebrahimie, E. (2014). Efficient and simple production of insulin-producing cells from embryonal carcinoma stem cells using mouse neonate pancreas extract, as a natural inducer. PloS One, 9, e90885.
Elmore, S. (2007). Apoptosis: A review of programmed cell death. Toxicologic Pathology, 35, 495–516.
Fruzangohar, M., Ebrahimie, E., & Adelson, D. L. (2014). Application of global transcriptome data in gene ontology classification and construction of a gene ontology interaction network. bioRxiv.
Fruzangohar, M., Ebrahimie, E., Ogunniyi, A. D., Mahdi, L. K., Paton, J. C., & Adelson, D. L. (2013). Comparative GO: A web application for comparative gene ontology and gene ontology-based gene selection in bacteria. PLoS One, 8, e58759.
Gregory, P. A., Bracken, C. P., Smith, E., Bert, A. G., Wright, J. A., Roslan, S., et al. (2011). An autocrine TGF-β/ZEB/miR-200 signaling network regulates establishment and maintenance of epithelial-mesenchymal transition. Molecular Biology of the Cell, 22, 1686–1698.
Gupta, S. (2003). Molecular signaling in death receptor and mitochondrial pathways of apoptosis (Review). International Journal of Oncology, 22, 15–20.
He, X., He, L., & Hannon, G. J. (2007). The guardian’s little helper: microRNAs in the p53 tumor suppressor network. Cancer Research, 67, 11099–11101.
Hosseinpour, B., Bakhtiarizadeh, M. R., Khosravi, P., & Ebrahimie, E. (2013). Predicting distinct organization of transcription factor binding sites on the promoter regions: a new genome-based approach to expand human embryonic stem cell regulatory network. Gene, 531, 212–219.
Hosseinpour, B., HajiHoseini, V., Kashfi, R., Ebrahimie, E., & Hemmatzadeh, F. (2012). Protein interaction network of Arabidopsis thaliana female gametophyte development identifies novel proteins and relations. PLoS One, 7, e49931.
Jin, Y., Chen, Z., Liu, X., & Zhou, X. (2013). Evaluating the microRNA targeting sites by luciferase reporter gene assay. MicroRNA Protocols, 117–127.
Johnston, R. J, Jr, Chang, S., Etchberger, J. F., Ortiz, C. O., & Hobert, O. (2005). MicroRNAs acting in a double-negative feedback loop to control a neuronal cell fate decision. Proceedings of the National Academy of Sciences of the United States of America, 102, 12449–12454.
Kalaitzidis, D., & Gilmore, T. D. (2005). Transcription factor cross-talk: the estrogen receptor and NF-kappa B. Trends in Endocrinology and Metabolism, 16, 46–52.
Kang, M. H., & Reynolds, C. P. (2009). Bcl-2 inhibitors: Targeting mitochondrial apoptotic pathways in cancer therapy. Clinical Cancer Research, 15, 1126–1132.
Kargarfard, F., Sami, A., & Ebrahimie, E. (2015). Knowledge discovery and sequence-based prediction of pandemic influenza using an integrated classification and association rule mining (CBA) algorithm. Journal of Biomedical Informatics, 57, 181–188.
Khaksefidi, R. E., Mirlohi, S., Khalaji, F., Fakhari, Z., Shiran, B., Fallahi, H., Rafiei, F., Budak, H., & Ebrahimie, E. (2015). Differential expression of seven conserved microRNAs in response to abiotic stress and their regulatory network in Helianthus annuus. Frontiers in Plant Science, 6.
Khan, A. A., Betel, D., Miller, M. L., Sander, C., Leslie, C. S., & Marks, D. S. (2009). Transfection of small RNAs globally perturbs gene regulation by endogenous microRNAs. Nature Biotechnology, 27, 549–555.
Kim, H. H., Kuwano, Y., Srikantan, S., Lee, E. K., Martindale, J. L., & Gorospe, M. (2009). HuR recruits let-7/RISC to repress c-Myc expression. Genes & Development, 23, 1743–1748.
Kim, N. H., Kim, H. S., Kim, N.-G., Lee, I., Choi, H.-S., Li, X.-Y., et al. (2011). p53 and miRNA-34 are suppressors of canonical Wnt signaling. Science Signaling, 4, ra71.
Kim, P. K. M., Armstrong, M., Liu, Y., Yan, P., Bucher, B., Zuckerbraun, B. S., et al. (2004). IRF-1 expression induces apoptosis and inhibits tumor growth in mouse mammary cancer cells in vitro and in vivo. Oncogene, 23, 1125–1135.
Liang, P., Stein, S., Thomas, E. K., Herzog, B., Westfall, M. D., Rocheleau, J. V., et al. (2004). NDRG1 is necessary for p53-dependent apoptosis. Journal of Biological Chemistry, 279, 48930–48940.
Lin, C.-C., Chen, Y.-J., Chen, C.-Y., Oyang, Y.-J., Juan, H.-F., & Huang, H.-C. (2012). Crosstalk between transcription factors and microRNAs in human protein interaction network. BMC Systems Biology, 6, 18.
Mahdi, L. K., Deihimi, T., Zamansani, F., Fruzangohar, M., Adelson, D. L., Paton, J. C., et al. (2014). A functional genomics catalogue of activated transcription factors during pathogenesis of pneumococcal disease. BMC Genomics, 15, 769.
Mahdi, L. K., Ebrahimie, E., Adelson, D. L., Paton, J. C., & Ogunniyi, A. D. (2013). A transcription factor contributes to pathogenesis and virulence in Streptococcus pneumoniae. PLoS One, 8, e70862.
Mansouri, A., Esmaeili, F., Nejatpour, A., Houshmand, F., Shabani, L., & Ebrahimie, E. (2014). Differentiation of P19 embryonal carcinoma stem cells into insulin‐producing cells promoted by pancreas‐conditioned medium. Journal of Tissue Engineering and Regenerative Medicine.
Mo, J. S., Jung, J., Yoon, J. H., Hong, J. A., Kim, M. Y., Ann, E. J., et al. (2010). DJ-1 modulates the p38 mitogen-activated protein kinase pathway through physical interaction with apoptosis signal-regulating kinase 1. Journal of Cellular Biochemistry, 110, 229–237.
Nielsen, C. B., Shomron, N., Sandberg, R., Hornstein, E., Kitzman, J., & Burge, C. B. (2007). Determinants of targeting by endogenous and exogenous microRNAs and siRNAs. RNA, 13, 1894–1910.
Nikitin, A., Egorov, S., Daraselia, N., & Mazo, I. (2003). Pathway studio—the analysis and navigation of molecular networks. Bioinformatics, 19, 2155–2157.
Panahi, B., Mohammadi, S. A. & Ebrahimie, E. (2013). Identification of miRNAs and their potential targets in halophyte plant Thellungiella halophila. BioTechnologia. Journal of Biotechnology Computational Biology and Bionanotechnology, 94.
Panahi, B., Mohammadi, S. A., Khaksefidi, R. E., Mehrabadi, J. F., & Ebrahimie, E. (2015). Genome-wide analysis of alternative splicing events in Hordeum vulgare: Highlighting retention of intron-based splicing and its possible function through network analysis. FEBS Letters, 589, 3564–3575.
Pressinotti, N. C., Klocker, H., Schafer, G., Luu, V. D., Ruschhaupt, M., Kuner, R., Steiner, E., Poustka, A., Bartsch, G., & Sultmann, H. (2009). Differential expression of apoptotic genes PDIA3 and MAP3K5 distinguishes between low- and high-risk prostate cancer. Molecular Cancer, 8.
Qin, F. Z., Patel, R., Yan, C., & Liu, W. M. (2006). NADPH oxidase is involved in angiotensin II-induced apoptosis in H9C2 cardiac muscle cells: Effects of apocynin. Free Radical Biology and Medicine, 40, 236–246.
Qiu, C. X., Wang, J. A., Yao, P. Y., Wang, E., & Cui, Q. H. (2010). microRNA evolution in a human transcription factor and microRNA regulatory network. BMC Systems Biology, 4.
Rokavec, M., Li, H., Jiang, L. & Hermeking, H. (2014). The p53/miR-34 axis in development and disease. Journal of Molecular Cell Biology.
Shamloo-Dashtpagerdi, R., Razi, H., Aliakbari, M., Lindlöf, A., Ebrahimi, M., & Ebrahimie, E. (2015). A novel pairwise comparison method for in silico discovery of statistically significant cis-regulatory elements in eukaryotic promoter regions: Application to Arabidopsis. Journal of Theoretical Biology, 364, 364–376.
Subramanian, A., Tamayo, P., Mootha, V. K., Mukherjee, S., Ebert, B. L., Gillette, M. A., et al. (2005). Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proceedings of the National Academy of Sciences of the United States of America, 102, 15545–15550.
Sun, X. H., Yang, Y. Z., & Wang, H. C. (2008). IdI induces apoptosis through inhibition of RORgammat expression. Bmc Immunology, 9.
Thomassen, M., Tan, Q. H., & Kruse, T. A. (2008) Gene expression meta-analysis identifies metastatic pathways and transcription factors in breast cancer. Bmc Cancer, 8.
Tsang, J., Zhu, J., & van Oudenaarden, A. (2007). MicroRNA-mediated feedback and feedforward loops are recurrent network motifs in mammals. Molecular Cell, 26, 753–767.
Walsh, B., Pearl, A., Suchy, S., Tartaglio, J., Visco, K., & Phelan, S. A. (2009). Overexpression of Prdx6 and resistance to peroxide-induced death in Hepa1-6 cells: Prdx suppression increases apoptosis. Redox Report : Communications in Free Radical Research, 14, 275–284.
Wiley, S. R., Schooley, K., Smolak, P. J., Din, W. S., Huang, C.-P., Nicholl, J. K., et al. (1995). Identification and characterization of a new member of the TNF family that induces apoptosis. Immunity, 3, 673–682.
Wright, K. L., Desai, S., Maurin, M., Smith, M. A., Bolick, S. C. E., Dessureault, S., et al. (2010). PRDM1 is required for mantle cell lymphoma response to bortezomib. Molecular Cancer Research, 8, 907–918.
Xu, D., Perez, R. E., Rezaiekhaligh, M. H., Bourdi, M., & Truog, W. E. (2009). Knockdown of ERp57 increases BiP/GRP78 induction and protects against hyperoxia and tunicamycin-induced apoptosis. American Journal of Physiology-Lung Cellular and Molecular Physiology, 297, L44–L51.
Yui, S., Nakatani, Y., & Mikami, M. (2003). Calprotectin (S100A8/S100A9), an inflammatory protein complex from neutrophils with a broad apoptosis-inducing activity. Biological & Pharmaceutical Bulletin, 26, 753–760.
Zhang, T., Zhang, X., Ding, K., Yang, K., Zhang, Z., & Xu, Y. (2010). PIM-1 gene RNA interference induces growth inhibition and apoptosis of prostate cancer cells and suppresses tumor progression in vivo. Journal of Surgical Oncology, 101, 513–519.
Acknowledgments
Computational analysis of this research was supported by School of Biological Sciences Science of The University of Adelaide.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Ibrahim Alanazi and Esmaeil Ebrahimie have been contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Alanazi, I.O., Ebrahimie, E. Computational Systems Biology Approach Predicts Regulators and Targets of microRNAs and Their Genomic Hotspots in Apoptosis Process. Mol Biotechnol 58, 460–479 (2016). https://doi.org/10.1007/s12033-016-9938-x
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12033-016-9938-x