Abstract
Purpose
Hepatocellular carcinoma (HCC) is one of the leading causes of cancer-related death worldwide with lack of effective systemic chemotherapy. In this study, we aimed to evaluate the value of ATPase family AAA domain-containing protein 2 (ATAD2) as a biomarker and potential therapeutic target for HCC.
Methods
The expression of ATAD2 was tested in different HCC patient cohorts by immunohistochemistry and comparative transcriptional analysis. The co-expression of ATAD2 and proliferation markers was compared during liver regeneration and malignancy with different bioinformatics tools. The cellular effects of ATAD2 inactivation in liver malignancy was tested on cell cycle, apoptosis, and colony formation ability as well as tumor formation using RNA interference. The genes affected by ATAD2 inactivation in three different HCC cell lines were identified by global gene expression profiling and bioinformatics tools.
Results
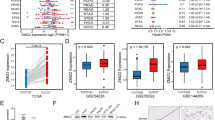
ATAD2 overexpression is closely correlated with HCC tumor stage. There was gradual increase from dysplasia, well-differentiated and poorly-differentiated HCC, respectively. We also observed transient upregulation of ATAD2 expression during rat liver regeneration in parallel to changes in Ki-67 expression. ATAD2 knockdown resulted in apoptosis and decreased cell survival in vitro and decreased tumor formation in some HCC cell lines. However, three other HCC cell lines tested were not affected. Similarly, gene expression response to ATAD2 inactivation in different HCC cell lines was highly heterogeneous.
Conclusions
ATAD2 is a potential proliferation marker for liver regeneration and HCC. It may also serve as a therapeutic target despite heterogeneous response of malignant cells.







Similar content being viewed by others
Availability of Data and Material
The contributions for the study are included in the article/Online Resources. Raw microarray data that support the findings of this study and further information are available upon request from corresponding author.
References
Cattaneo M, Morozumi Y, Perazza D, Boussouar F, Jamshidikia M, Rousseaux S, Verdel A, Khochbin S. Lessons from yeast on emerging roles of the ATAD2 protein family in gene regulation and genome organization. Mol Cells. 2014;37:851–6. https://doi.org/10.14348/molcells.2014.0258.
Morozumi Y, Boussouar F, Tan M, Chaikuad A, Jamshidikia M, Colak G, He H, Nie L, Petosa C, De Dieuleveult M, et al. ATAD2 is a generalist facilitator of chromatin dynamics in embryonic stem cells. J Mol Cell Biol. 2016;8:349–62. https://doi.org/10.1093/jmcb/mjv060.
Koo SJ, Fernández-Montalván AE, Badock V, Ott CJ, Holton SJ, von Ahsen O, Toedling J, Vittori S, Bradner JE, Gorjánácz M. ATAD2 is an epigenetic reader of newly synthesized histone marks during DNA replication. Oncotarget. 2016;7:70323–35. https://doi.org/10.18632/oncotarget.11855.
Cho C, Jang J, Kang Y, Watanabe H, Uchihashi T, Kim SJ, Kato K, Lee JY, Song JJ. Structural basis of nucleosome assembly by the Abo1 AAA+ ATPase histone chaperone. Nat Commun. 2019;10:1–13. https://doi.org/10.1038/s41467-019-13743-9.
Shahnejat-Bushehri S, Ehrenhofer-Murray AE. The ATAD2/ANCCA homolog Yta7 cooperates with Scm3HJURP to deposit Cse4CENP-A at the centromere in yeast. Proc Natl Acad Sci U S A. 2020;117:5386–93. https://doi.org/10.1073/pnas.1917814117.
Zou JX, Revenko AS, Li LB, Gemo AT, Chen HW. ANCCA, an estrogen-regulated AAA+ ATPase coactivator for ERα, is required for coregulator occupancy and chromatin modification. Proc Natl Acad Sci U S A. 2007;104:18067–72. https://doi.org/10.1073/pnas.0705814104.
Ciró M, Prosperini E, Quarto M, Grazini U, Walfridsson J, McBlane F, Nucifero P, Pacchiana G, Capra M, Christensen J, et al. ATAD2 is a novel cofactor for MYC, overexpressed and amplified in aggressive tumors. Cancer Res. 2009;69:8491–8. https://doi.org/10.1158/0008-5472.CAN-09-2131.
Caron C, Lestrat C, Marsal S, Escoffier E, Curtet S, Virolle V, Barbry P, Debernardi A, Brambilla C, Brambilla E, et al. Functional characterization of ATAD2 as a new cancer/testis factor and a predictor of poor prognosis in breast and lung cancers. Oncogene. 2010;29:5171–81. https://doi.org/10.1038/onc.2010.259.
Yildiz G, Arslan-Ergul A, Bagislar S, Konu O, Yuzugullu H, Gursoy-Yuzugullu O, Ozturk N, Ozen C, Ozdag H, Erdal E, Karademir S, et al. Genome-wide transcriptional reorganization associated with senescence-to-immortality switch during human hepatocellular carcinogenesis. PLoS One. 2013;8. https://doi.org/10.1371/journal.pone.0064016.
Hussain M, Zhou Y, Song Y, Hameed HMA, Jiang H, Tu Y, Zhang J. ATAD2 in cancer: a pharmacologically challenging but tractable target. Expert Opin Ther Targets. 2018;22:85–96. https://doi.org/10.1080/14728222.2018.1406921.
Yuzugullu H, Benhaj K, Ozturk N, Senturk S, Celik E, Toylu A, Tasdemir N, Yilmaz M, Erdal E, Akcali KC, et al. Canonical Wnt signaling is antagonized by noncanonical Wnt5a in hepatocellular carcinoma cells. Mol Cancer. 2009;8:90. https://doi.org/10.1186/1476-4598-8-90.
Torbenson MS, Ng IOL, Park YN, Roncalli M, Sakamoto M. Hepatocellular carcinoma. In: WHO Classification of Tumors Editorial Board. Digestive system tumours. WHO classification of tumours series. 5th ed. Lyon: Int Agen Res Cancer. 2019:229–39.
Goldman MJ, Craft B, Hastie M, Repečka K, McDade F, Kamath A, Banerjee A, Luo Y, Rogers D, Brooks AN, et al. Visualizing and interpreting cancer genomics data via the Xena platform. Nat Biotechnol. 2020;38:675–8. https://doi.org/10.1038/s41587-020-0546-8.
Gautier L, Cope L, Bolstad BM, Irizarry RA. Affy-analysis of Affymetrix GeneChip data at the probe level. Bioinformatics. 2004;20:307–15. https://doi.org/10.1093/bioinformatics/btg405.
Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, Smyth GK. Limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015;43: e47. https://doi.org/10.1093/nar/gkv007.
Yu G, Wang LG, Han Y, He QY. ClusterProfiler: an R package for comparing biological themes among gene clusters. Omi A J Integr Biol. 2012;16:284–7. https://doi.org/10.1089/omi.2011.0118.
Edgar R, Domrachev M, Lash AE. Gene Expression Omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res. 2002;30:207–10. https://doi.org/10.1093/nar/30.1.207.
Brewer JW, Hendershot LM, Sherr CJ, Diehl JA. Mammalian unfolded protein response inhibits cyclin D1 translation and cell-cycle progression. Proc Natl Acad Sci U S A. 1999;96:8505–10. https://doi.org/10.1073/pnas.96.15.8505.
Banerjee A, Lang JY, Hung MC, Sengupta K, Banerjee SK, Baksi K, Banerjee DK. Unfolded protein response is required in nu/nu mice microvasculature for treating breast tumor with tunicamycin. J Biol Chem. 2011;286:29127–38. https://doi.org/10.1074/jbc.M110.169771.
Yang J, Huang J, Luo L, Chen Z, Guo Y, Guo L. Significance of PRO2000/ANCCA expression, a novel proliferation-associated protein in hepatocellular carcinoma. Cancer Cell Int. 2014;14:1–7. https://doi.org/10.1186/1475-2867-14-33.
Huang J, Yang J, Lei Y, Gao H, Wei T, Luo L, Zhang F, Chen H, Zeng Q, Guo L. An ANCCA/PRO2000-miR-520a-E2F2 regulatory loop as a driving force for the development of hepatocellular carcinoma. Oncogenesis. 2016;5:e229. https://doi.org/10.1038/oncsis.2016.22.
Hwang HW, Ha SY, Bang H, Park CK. ATAD2 as a poor prognostic marker for hepatocellular carcinoma after curative resection. Cancer Res Treat. 2015;47:853–61. https://doi.org/10.4143/crt.2014.177.
Kaita KDE, Pettigrew N, Minuk GY. Hepatic regeneration in humans with various liver disease as assessed by Ki-67 staining of formalin-fixed paraffin-embedded liver tissue. Liver. 1997;17:13–6. https://doi.org/10.1111/j.1600-0676.1997.tb00772.x.
Gerlach C, Sakkab DY, Scholzen T, Daßler R, Alison MR, Gerdes J. Ki-67 expression during rat liver regeneration after partial hepatectomy. Hepatology. 1997;26:573–8. https://doi.org/10.1053/jhep.1997.v26.pm0009303485.
Gerdes J, Schwab U, Lemke H, Stein H. Production of a mouse monoclonal antibody reactive with a human nuclear antigen associated with cell proliferation. Int J Cancer. 1983;31:13–20. https://doi.org/10.1002/ijc.2910310104.
Cuylen S, Blaukopf C, Politi AZ, Muller-Reichert T, Neumann B, Poser I, Ellenberg J, Hyman AA, Gerlich DW. Ki-67 acts as a biological surfactant to disperse mitotic chromosomes. Nature. 2016;535:308–12. https://doi.org/10.1038/nature18610.
Bassik MC, Kampmann M. Knocking out the door to tunicamycin entry. Proc Natl Acad Sci U S A. 2011;108:11731–2. https://doi.org/10.1073/pnas.1109035108.
Lai S-S, Zhao D-D, Cao P, Lu K, Luo O-Y, Chen W-B, Liu J, Jiang E-Z, Yu Z-H, Lee G, et al. PP2Acα positively regulates the termination of liver regeneration in mice through the AKT/GSK3β/Cyclin D1 pathway. J Hepatol. 2016;64:352–60. https://doi.org/10.1016/j.jhep.2015.09.025.
Oki T, Nishimura K, Kitaura J, Togami K, Maehara A, Izawa K, Sakaue-Sawano A, Niida A, Miyano S, Aburatani H, Kiyonari H, et al. A novel cell-cycle-indicator, mVenus-p27K -, identifies quiescent cells and visualizes G0-G1 transition. Sci Rep. 2014;4. https://doi.org/10.1038/srep04012.
Sreekumar R, Emaduddin M, Al-Saihati H, Moutasim K, Chan J, Spampinato M, Bhome R, Yuen HM, Mescoli C, Vitale A, Cillo U, et al. Protein kinase C inhibitors override ZEB1-induced chemoresistance in HCC. Cell Death Dis. 2019;10. https://doi.org/10.1038/s41419-019-1885-6.
Sayan AE, Sayan BS, Findikli N, Ozturk M. Acquired expression of transcriptionally active p73 in hepatocellular carcinoma cells. Oncogene. 2001;20:5111–7. https://doi.org/10.1038/sj.onc.1204669.
Lu WJ, Chua MS, So SK. Suppression of ATAD2 inhibits hepatocellular carcinoma progression through activation of p53-and p38-mediated apoptotic signaling. Oncotarget. 2015;6:41722–35. https://doi.org/10.18632/oncotarget.6152.
Wu G, Liu H, He H, Wang Y, Lu X, Yu Y, Xia S, Meng X, Liu Y. MiR-372 down-regulates the oncogene ATAD2 to influence hepatocellular carcinoma proliferation and metastasis. BMC Cancer. 2014;14:1–11. https://doi.org/10.1186/1471-2407-14-107.
Acknowledgements
This work was part of PhD theses of H.Y. (Bilkent University, Ankara, Turkey). We would like to thank Izmir Biomedicine and Genome Center’s core facilities especially Optical Imaging and IBG Bioinformatics Platform (IBG-BIP) for their support during data collection and analysis.
Funding
This study was supported by the funds from TUBITAK (109S191, 111T558), Turkish Academy of Sciences, Dokuz Eylul University, and İzmir Biomedicine and Genome Center.
Author information
Authors and Affiliations
Contributions
Study conception and design: MO; experiments: UE, HY, CO, OY, PK, and EB. Bioinformatics and statistical analyses: GK, HU, and UE; evaluation of the data: MO, NA, HA, FY, and PBK; drafting of the manuscript: MO and UE. All authors were involved in finalization of the manuscript and approved the submitted version.
Corresponding author
Ethics declarations
Ethics Approval
Study with clinical samples was approved by Ege University Ethics Committee. Animal experiments were approved by Bilkent University Animal Ethics Committee. (Decision No: 2006/1; Decision date: 10/5/2006).
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Ekin, U., Yuzugullu, H., Ozen, C. et al. Evaluation of ATAD2 as a Potential Target in Hepatocellular Carcinoma. J Gastrointest Canc 52, 1356–1369 (2021). https://doi.org/10.1007/s12029-021-00732-9
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12029-021-00732-9