Abstract
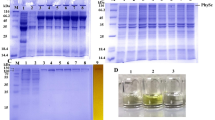
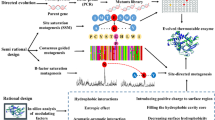
To improve the temperature characteristics of AoXyn11A, a mesophilic glycoside hydrolase family (GHF) 11 xylanase from Aspergillus oryzae CICC40186, its N-terminal and “cord” regions were selected to be substituted by means of the computer-aided analysis and calculation. In brief, one mutant, named ATX11A41, possessing the lowest root-mean-square deviation (RMSD) value was designed based on the molecular dynamics (MD) simulation by substituting the N-terminal 41 amino acids of AoXyn11A with the corresponding 42 ones of pXYL11, a thermophilic GHF11 xylanase from Thermobifida fusca. On the basis of the primary structure alignment of pXYL11 with ATX11A41 (or AoXyn11A), another mutant, named ATX11A41/cord, was designed by substituting the cord region (93GTYNPGSGG101) of ATX11A41 with the corresponding one (93GTYRPTG99) of pXYL11. Both mutant-encoding genes, ATx11A41 and ATx11A41/cord, were constructed as designed theoretically by a megaprimer PCR technique and were expressed in Pichia pastoris GS115. The specific activities of recombinant (re) AoXyn11A, ATX11A41, and ATX11A41/cord were 2916.7, 2667.6, and 2457.0 U/mg, respectively. The analysis of temperature characteristics displayed that the temperature optimum (Topt) of reATX11A41 or reATX11A41/cord was 65 °C, which was 15 °C higher than that of reAoXyn11A. The thermal inactivation half-life (t1/2) values of reATX11A41 and reATX11A41/cord at 60 °C were 55 and 83 min, respectively, whereas that of reAoXyn11A was only 18 min at 50 °C. The melting temperature (Tm) values of reAoXyn11A, reATX11A41, and reATX11A41/cord were 54.2, 66.7, and 71.9 °C, respectively. In conclusion, the above findings indicated that the substitution of both the N-terminal and cord regions of a mesophilic AoXyn11A greatly contributed to its improved temperature characteristics.







Similar content being viewed by others
References
Trevizano, L. M., Ventorim, R. Z., de Rezende, S. T., Junior, F. P. S., & Guimarães, V. M. (2012). Thermostability improvement of Orpinomyces sp. xylanase by directed evolution. Journal of Molecular Catalysis B: Enzymatic, 81, 12–18. https://doi.org/10.1016/j.molcatb.2012.04.021.
Fu, X. Y., Zhao, W., Xiong, A. S., Tian, Y. S., & Peng, R. H. (2011). High expression of recombinant Streptomyces sp. S38 xylanase in Pichia pastoris by codon optimization and analysis of its biochemical properties. Molecular Biology Reports, 38(8), 4991–4997. https://doi.org/10.1007/s11033-010-0644-7.
Conejo-Saucedo, U., Cano-Camacho, H., Villa-Rivera, M. G., Lara-Marquez, A., López-Romero, E., & Zavala-Páramo, M. G. (2017). Protein homology modeling, docking, and phylogenetic analyses of an endo-1,4-β-xylanase GH11 of Colletotrichum lindemuthianum. Mycological Progress, 16(6), 577–591. https://doi.org/10.1007/s11557-017-1291-3.
Paës, G., Berrin, J. G., & Beaugrand, J. (2012). GH11 xylanases: structure/function/properties relationships and applications. Biotechnology Advances, 30(3), 564–592. https://doi.org/10.1016/j.biotechadv.2011.10.003.
Zhang, H. M., Li, J. F., Wang, J. Q., Yang, Y. J., & Wu, M. C. (2014). Determinants for the improved thermostability of a mesophilic family 11 xylanase predicted by computational methods. Biotechnology for Biofuels, 7(1), 3. https://doi.org/10.1186/1754-6834-7-3.
Robledo, A., Aguilar, C. N., Belmares-Cerda, R. E., Flores-Gallegos, A. C., Contreras-Esquivel, J. C., Montañez, J. C., & Mussatto, S. I. (2016). Production of thermostable xylanase by thermophilic fungal strains isolated from maize silage. CyTA-Journal of Food, 14(2), 302–308. https://doi.org/10.1080/19476337.2015.1105298.
Kumar, S., Tsai, C. J., & Nussinov, R. (2000). Factors enhancing protein thermostability. Protein Engineering, 13(3), 179–191. https://doi.org/10.1093/protein/13.3.179.
Yu, H. R., & Huang, H. (2014). Engineering proteins for thermostability through rigidifying flexible sites. Biotechnology Advances, 32(2), 308–315. https://doi.org/10.1016/j.biotechadv.2013.10.012.
Wang, Q., & Xia, T. (2008). Importance of C-terminal region for thermostability of GH11 xylanase from Streptomyces lividans. Applied Biochemistry and Biotechnology, 144(3), 273–282. https://doi.org/10.1007/s12010-007-8016-z.
Yin, X., Li, J. F., Wang, J. Q., Tang, C. D., & Wu, M. C. (2013). Enhanced thermostability of a mesophilic xylanase by N-terminal replacement designed by molecular dynamics simulation. Journal of the Science of Food and Agriculture, 93(12), 3016–3023. https://doi.org/10.1002/jsfa.6134.
Dumon, C., Varvak, A., Wall, M. A., Flint, J. E., Lewis, R. J., Lakey, J. H., Morland, C., Luginbühl, P., Healey, S., Todaro, T., DeSantis, G., Sun, M., Parra-Gessert, L., Tan, X., Weiner, D. P., & Gilbert, H. J. (2008). Engineering hyperthermostability into a GH11 xylanase is mediated by subtle changes to protein structure. Journal of Biological Chemistry, 283(33), 22557–22564. https://doi.org/10.1074/jbc.M800936200.
Hakulinen, N., Turunen, O., Janis, J., Leisola, M., & Rouvinen, J. (2003). Three-dimensional structures of thermophilic β-1,4-xylanases from Chaetomium thermophilum and Nonomuraea flexuosa: comparison of twelve xylanases in relation to their thermal stability. European Journal of Biochemistry, 270(7), 1399–1412. https://doi.org/10.1046/j.1432-1033.2003.03496.x.
Li, J. F., Gao, S. J., Liu, X. T., Gong, Y. Y., Chen, Z. F., Wei, X. H., Zhang, H. M., & Wu, M. C. (2013). Modified pPIC9K vector-mediated expression of a family 11 xylanase gene, Aoxyn11A, from Aspergillus oryzae in Pichia pastoris. Annals of Microbiology, 63(3), 1109–1120. https://doi.org/10.1007/s13213-012-0568-7.
Cheng, Y. F., Yang, C. H., & Liu, W. H. (2005). Cloning and expression of Thermobifida xylanase gene in the methylotrophic yeast Pichia pastoris. Enzyme Microbial Technology, 37(5), 541–546. https://doi.org/10.1016/j.enzmictec.2005.04.006.
Reetz, M. T., & Carballeira, J. D. (2007). Iterative saturation mutagenesis (ISM) for rapid directed evolution of functional enzymes. Nature Protocols, 2(4), 891–903. https://doi.org/10.1038/nprot.2007.72.
Xie, Z. H., & Shi, X. J. (2009). Fast and almost 100% efficiency site-directed mutagenesis by the megaprimer PCR method. Progress in Biochemistry and Biophysics, 36(11), 1490–1494. https://doi.org/10.3724/SP.J.1206.2009.00139.
Jang, M. K., Lee, S. W., Lee, D. G., Kim, N. Y., Yu, K. H., Jang, H. J., Kim, S., Kim, A., & Lee, S. H. (2010). Enhancement of the thermostability of a recombinant β-agarase, AgaB, from Zobellia galactanivorans by random mutagenesis. Biotechnology Letters, 32(7), 943–949. https://doi.org/10.1007/s10529-010-0237-5.
Dong, Y. H., Li, J. F., Hu, D., Yin, X., Wang, C. J., Tang, S. H., & Wu, M. C. (2016). Replacing a piece of loop-structure in the substrate-binding groove of Aspergillus usamii β-mannanase, AuMan5A, to improve its enzymatic properties by rational design. Applied Microbiology and Biotechnology, 100(9), 3989–3998. https://doi.org/10.1007/s00253-015-7224-7.
Le, Q. A., Joo, J. C., Yoo, Y. J., & Kim, Y. H. (2012). Development of thermostable Candida antarctica lipase B through novel in silico design of disulfide bridge. Biotechnology and Bioengineering, 109(4), 867–876. https://doi.org/10.1002/bit.24371.
Badieyan, S., Bevan, D. R., & Zhang, C. M. (2012). Study and design of stability in GH5 cellulases. Biotechnology and Bioengineering, 109(1), 31–44. https://doi.org/10.1002/bit.23280.
Radestock, S., & Gohlke, H. (2008). Exploiting the link between protein rigidity and thermostability for data-driven protein engineering. Engineering in Life Sciences, 8(5), 507–522. https://doi.org/10.1002/elsc.200800043.
Niesen, F. H., Berglund, H., & Vedadi, M. (2007). The use of differential scanning fluorimetry to detect ligand interactions that promote protein stability. Nature Protocols, 2(9), 2212–2221. https://doi.org/10.1038/nprot.2007.321.
Zhang, S., Zhang, K., Chen, X. Z., Chu, X., Sun, F., & Dong, Z. Y. (2010). Five mutations in N-terminus confer thermostability on mesophilic xylanase. Biochemical and Biophysical Research Communications, 395(2), 200–206. https://doi.org/10.1016/j.bbrc.2010.03.159.
You, C., Huang, Q., Xue, H. P., Xu, Y., & Lu, H. (2010). Potential hydrophobic interaction between two cysteines in interior hydrophobic region improves thermostability of a family 11 xylanase from Neocallimastix patriciarum. Biotechnology and Bioengineering, 105(5), 861–870. https://doi.org/10.1002/bit.22623.
Tina, K. G., Bhadra, R., & Srinivasan, N. (2007). PIC: Protein Interactions Calculator. Nucleic Acids Research, 35(Web Server), W473–W476. https://doi.org/10.1093/nar/gkm423.
Georis, J., Esteves, F. D., Lamotte-Brasseur, J., Bougnet, V., Devreese, B., Giannotta, F., et al. (2000). An additional aromatic interaction improves the thermostability and thermophilicity of a mesophilic family 11 xylanase: structural basis and molecular study. Protein Science, 9, 466–475.
Lee, J. H., Heo, S. Y., Lee, J. W., Yoon, K. H., Kim, Y. H., & Nam, S. W. (2009). Thermostability and xylan-hydrolyzing property of endoxylanase expressed in yeast Saccharomyces cerevisiae. Biotechnology and Bioprocess Engineering, 14(5), 639–644. https://doi.org/10.1007/s12257-009-0014-2.
Anbarasan, S., Jänis, J., Paloheimo, M., Laitaoja, M., Vuolanto, M., Karimäki, J., et al. (2010). Effect of glycosylation and additional domains on the thermostability of a family 10 xylanase produced by Thermopolyspora flexuosa. Applied and Environmental Microbiology, 76(1), 356–360. https://doi.org/10.1128/AEM.00357-09.
Thompson, M. J., & Eisenberg, D. (1999). Transproteomic evidence of a loop-deletion mechanism for enhancing protein thermostability. Journal of Molecular Biology, 290(2), 595–604. https://doi.org/10.1006/jmbi.1999.2889.
Hawwa, R., Aikens, J., Turner, R. J., Santarsiero, B. D., & Mesecar, A. D. (2009). Structural basis for thermostability revealed through the identification and characterization of a highly thermostable phosphotriesterase-like lactonase from Geobacillus stearothermophilus. Archives of Biochemistry and Biophysics, 488(2), 109–120. https://doi.org/10.1016/j.abb.2009.06.005.
Xue, H. P., Zhou, J. G., You, C., Huang, Q., & Lu, H. (2012). Amino acid substitutions in the N-terminus, cord and α-helix domains improved the thermostability of a family 11 xylanase XynR8. Journal of Industrial Microbiology and Biotechnology, 39, 1279–1288.
Donald, J. E., Kulp, D. W., & DeGrado, W. F. (2011). Salt bridges: geometrically specific, designable interactions. Proteins, 79, 898–915.
Funding
This work was financially supported by the National Natural Science Foundation of China (21676117) and the Postgraduate Innovation Training Project of Jiangsu (KYLX16_0804). The authors are grateful to Prof. Xianzhang Wu (School of Biotechnology, Jiangnan University, Jiangsu, China) for providing the technical assistance.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
ESM 1
(DOCX 342 kb)
Rights and permissions
About this article
Cite this article
Li, C., Li, J., Wang, R. et al. Substituting Both the N-Terminal and “Cord” Regions of a Xylanase from Aspergillus oryzae to Improve Its Temperature Characteristics. Appl Biochem Biotechnol 185, 1044–1059 (2018). https://doi.org/10.1007/s12010-017-2681-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12010-017-2681-3