Abstract
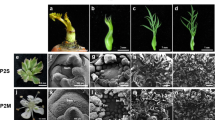
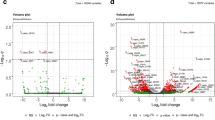
CMS is induced by the coordinated expression of certain mitochondrial and nuclear genes in flower development. Mitochondrial genes regulate manifestation of CMS, whereas nuclear genes regulate fertility phenotype and thus affect negatively. In this article, the buds of newly bred Ogura CMS of non-heading Chinese cabbage and its maintainer line as plant materials, genes differentially expressed transcripts were analyzed by cDNA-AFLP. Seventeen differently expressed genes were found in new Ogura CMS and nine genes in maintainer line. These genes were involved in energy metabolism, signal transduction, flower development, stress-related metabolism, transcription, etc. Expression patterns of three genes encoding BrCAM6, BrANK, BrTUB3 were verified by qRT-PCR in different organs and various stage flower buds of CMS and its maintainer line. The results revealed that two genes related to signal transduction, BrCAM6 and BrANK, were highly expressed in stamens and microspores of CMS than in maintainer line. As believed, these two genes involved in signal transduction of male sterile in CMS line. In comparison, BrTUB3 gene was accumulated in stamens and was expressed in significantly lower level in CMS line than in maintainer line. It expressed significantly lower in CMS than maintainer line after tetrad stage. This expression profile suggests that BrTUB3 played an important role in the development of the pollen, and may be closely related to male sterility.



Similar content being viewed by others
Abbreviations
- CMS:
-
Cytoplasmic male sterility
- cDNA-AFLP:
-
cDNA-amplified fragment length polymorphism
- qRT-PCR:
-
quantitative Real-time RT-PCR
- BrCAM6 :
-
Calcium-ion-binding 6
- BrANK :
-
Ankyrin repeat family protein
- BrTUB3 :
-
Tubulin alpha-3
- GAPDH :
-
Glyceraldehyde-3-phosphate dehydrogenase
Reference:
Artavanis-Tsakonas S, Deladakis C, Fehon RG (1991) The Notch locus and the cell biology of neuroblast segregation. Ann Rev Cell Biol 7:427–452
Bachem CWB, Van der Hoeven RS, deBruijin SM et al (1996) Visualization of differential gene expression using a novel method of RNA fingerprinting based on AFLP: analysis of gene expression during tuber development. Plant J 9:745–753
Balk J, Leaver C (2001) The PET1-CMS mitochondrial mutation in sunflower is associated with premature programmed cell death and cytochrome c release. Plant Cell 13:1803–1818
Breeden L, Nasmyth K (1987) Cell cycle control of the yeast HO gene: cis- and trans-acting regulators. Cell 48:389–397
Butow RA, Avadhani NG (2004) Mitochondrial signaling: the retrograde response. Mol Cell 14:1–15
Carlsson J, Lagercrantz U, Sundstrom J et al (2007) Microarray analysis reveals altered expression of a large number of nuclear genes in developing cytoplasmic male sterile Brassica napus flowers. Plant J 49:452–462
Carpenter JL, Ploense SE, Snustad DP, Silflow CD (1992) Preferential expression of an α-tubulin gene of Arabidopsis in pollen. Plant Cell 4:557–571
Carpenter JL, Kopczak SD, Snustad DP (1993) Semi-constitutive expression of an Arabidopsis thaliana α-tubulin gene. Plant Mol Biol 21:937–942
Ditt RF, Nester EW, Comai L (2001) Plant gene expression response to Agrobacterium tumefaciens. Proc Natl Acad Sci USA 98:10954–10959
Durrant WE, Rowland O, Piedras P, Hammond-Kosack KE, Jones JD (2000) cDNA-AFLP reveals a striking overlap in race-specific resistance and wound response gene expression profiles. Plant Cell 12:963–977
Fujii S, Komatsu S, Toriyama K (2007) Retrograde regulation of nuclear gene expression in CW-CMS of rice. Plant Mol Biol 63:405–417
Hanson MR, Bentolila S (2004) Interactions of mitochondrial and nuclear genes that affect male gametophyte development. Plant Cell 16:S154–S169
Hou XL, Cao SC, He YK (2004) Creation of a new germplasm of CMS non-heading Chinese cabbage. Acta Hortic 637:75–81
Krishnasamy S, Makaroff CA (1994) Organ-specific reduction in the abundance of a mitochondrial protein accompanies fertility restoration in cytoplasmic male-sterile radish. Plant Mol Biol 32:879–890
Linke B, Nothnagel T, Borner T (2003) Flower development in carrot CMS plants: mitochondria affect the expression of MADS box genes homologous to GLOBOSA and DEFICIENS. Plant J 34:27–37
Lou P, Kang JG, Zhang GY et al (2007) Transcript profiling of a dominant male sterile mutant (Ms-cd1) in cabbage during flower bud development. Plant Sci 172:111–119
Murai K, Takumi S, Koga H, Ogihara Y (2002) Pistillody, homeotic transformation of stamens into pistil-like structures, caused by nuclear–cytoplasm interaction in wheat. Plant J 29:169–181
Ogura H (1968) Studies on the new male sterility in Japanese radish, with special reference to the utilization of this sterility towards the practical raising of hybrid seeds. Mem Fac Agr Kagoshima Univ 6:39–78
Teixeira RT, Farbos I, Glimelius K (2005) Expression levels of meristem identity and homeotic genes are modified by nuclear–mitochondrial interactions in alloplasmic male-sterile lines of Brassica napus. Plant J 42:731–742
Watanabe T, Noritake J, Kaibuchi K (2005) Regulation of microtubule in cell migration. Trends Cell Biol 15:76–83
Xu X, Jiang CZ, Donnelly L, Reid MS (2007) Functional analysis of a RING domain ankyrin repeat protein that is highly expressed during flower senescence. J Exp Bot 58:3623–3630
Ye XL, Edward Y, Xu SX et al (2003) Microtubule Structure and Male Sterility in a Gene Cytoplasmic Male Sterile Line of Rice, Zhen Shan97A. Acta Bot Sin 45:183–192
Yuan JY, Hou XL, Zhang CW, Ye F (2007) Active oxygen metabolism in the floral buds and leaves of the new cytoplasm male sterile (CMS) line and its maintainer line of non-heading Chinese cabbage. Front Agric China 2:1–5
Zhang JY, Li Y, Shi GJ et al (2009) Characterization of α-tubulin gene distinctively presented in a cytoplasmic male sterile and its maintainer line of non-heading Chinese cabbage. J Sci Food Agric 89:274–280
Zubko MK (2004) Mitochondrial tuning fork in nuclear homeotic functions. Trends Plant Sci 9:61–64
Acknowledgments
We wish to thank Dr. Zengcui Zhang (North Dakota State University, USA) and Dr. Chowda Reddy (Southern Crop Protection and Food Research Centre, Canada) for critical reading the manuscript. This research was supported by the national natural science foundation of China(30671420) and Scientific research Innovations Program of Graduate students in General higher learning institutions of Jiangsu province in 2008 (CX08B_154Z).
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by L. A. Kleczkowski.
Rights and permissions
About this article
Cite this article
Zhang, C., Qi, L., Hou, X. et al. Differential gene expression analysis of a new Ogura CMS line and its maintainer in non-heading Chinese cabbage by cDNA-AFLP. Acta Physiol Plant 32, 781–787 (2010). https://doi.org/10.1007/s11738-010-0463-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11738-010-0463-4