Abstract
Background
The long non-coding RNA CRNDE has emerged as an important regulator in carcinogenesis and cancer progression. While CRNDE has previously been found to be the most highly upregulated lncRNA in glioma, detailed information on its roles in regulating cancer cell growth remains limited.
Objective
In the present study, we aimed at exploring the functional roles and underlying mechanisms of CRNDE in glioma.
Methods
We applied microarray data analysis to determine the prognostic significance of CRNDE in glioma patients and its correlation with epidermal growth factor receptor (EGFR) activation. EGFR inhibition was used to confirm the role of EGFR in regulating CRNDE expression. Functional studies were performed upon CRNDE silencing to explore its role in gliomagenesis.
Results
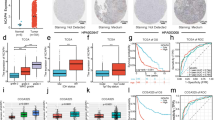
We confirm that CRNDE acts as an oncogene that is highly up-regulated in glioma, and high CRNDE expression correlates with poor prognosis in glioma patients. We further demonstrate that the expression of CRNDE correlates with EGFR activation. EGF and EGFR tyrosine kinase inhibitor (TKI) enhance and block the up-regulation of CRNDE expression, respectively, suggesting that EGFR signaling may positively regulate CRNDE expression. Functional assays show that CRNDE depletion inhibits glioma cell growth both in vitro and in vivo, and is associated with induced cellular apoptosis with decreased Bcl2/Bax ratio.
Conclusions
Our findings suggest that the aberrant expression of CRNDE may be mediated by activated EGFR signaling and play significant roles in gliomagenesis.






Similar content being viewed by others
References
Wilusz JE, Sunwoo H, Spector DL. Long noncoding RNAs: functional surprises from the RNA world. Genes Dev. 2009;23(13):1494–504. doi:10.1101/gad.1800909.
Kung JT, Colognori D, Lee JT. Long noncoding RNAs: past, present, and future. Genetics. 2013;193(3):651–69. doi:10.1534/genetics.112.146704.
Rinn JL, Chang HY. Genome regulation by long noncoding RNAs. Annu Rev Biochem. 2012;81:145–66. doi:10.1146/annurev-biochem-051410-092902.
Gibb EA, Brown CJ, Lam WL. The functional role of long non-coding RNA in human carcinomas. Mol Cancer. 2011;10:38. doi:10.1186/1476-4598-10-38.
Prensner JR, Chinnaiyan AM. The emergence of lncRNAs in cancer biology. Cancer Discov. 2011;1(5):391–407. doi:10.1158/2159-8290.CD-11-0209.
Mercer TR, Dinger ME, Mattick JS. Long non-coding RNAs: insights into functions. Nat Rev Genet. 2009;10(3):155–9. doi:10.1038/nrg2521.
Wang KC, Chang HY. Molecular mechanisms of long noncoding RNAs. Mol Cell. 2011;43(6):904–14. doi:10.1016/j.molcel.2011.08.018.
Skroblin P, Mayr M. "going long": long non-coding RNAs as biomarkers. Circ Res. 2014;115(7):607–9. doi:10.1161/Circresaha.114.304839.
Yarmishyn AA, Kurochkin IV. Long noncoding RNAs: a potential novel class of cancer biomarkers. Front Genet. 2015;6:145. doi:10.3389/fgene.2015.00145.
Qi P, Du X. The long non-coding RNAs, a new cancer diagnostic and therapeutic gold mine. Mod Pathol. 2013;26(2):155–65. doi:10.1038/modpathol.2012.160.
Spizzo R, Almeida MI, Colombatti A, Calin GA. Long non-coding RNAs and cancer: a new frontier of translational research? Oncogene. 2012;31(43):4577–87. doi:10.1038/onc.2011.621.
Omuro A, DeAngelis LM. Glioblastoma and other malignant gliomas: a clinical review. JAMA. 2013;310(17):1842–50. doi:10.1001/jama.2013.280319.
Wen PY, Kesari S. Malignant gliomas in adults. N Engl J Med. 2008;359(5):492–507. doi:10.1056/NEJMra0708126.
Schwartzbaum JA, Fisher JL, Aldape KD, Wrensch M. Epidemiology and molecular pathology of glioma. Nat Clin Pract Neurol. 2006;2(9):494–503. doi:10.1038/ncpneuro0289.
Mrugala MM. Advances and challenges in the treatment of glioblastoma: a clinician’s perspective. Discov Med. 2013;15(83):221–30.
Wick W, Platten M, Weller M. New (alternative) temozolomide regimens for the treatment of glioma. Neuro-Oncology. 2009;11(1):69–79. doi:10.1215/15228517-2008-078.
Huse JT, Holland E, DeAngelis LM. Glioblastoma: molecular analysis and clinical implications. Annu Rev Med. 2013;64:59–70. doi:10.1146/annurev-med-100711-143028.
Zhang X, Sun S, Pu JK, Tsang AC, Lee D, Man VO, et al. Long non-coding RNA expression profiles predict clinical phenotypes in glioma. Neurobiol Dis. 2012;48(1):1–8. doi:10.1016/j.nbd.2012.06.004.
Graham LD, Pedersen SK, Brown GS, Ho T, Kassir Z, Moynihan AT, et al. Colorectal Neoplasia differentially expressed (CRNDE), a novel Gene with elevated expression in colorectal adenomas and adenocarcinomas. Genes Cancer. 2011;2(8):829–40. doi:10.1177/1947601911431081.
Ellis BC, Molloy PL, Graham LD. CRNDE: a long non-coding RNA involved in CanceR, neurobiology, and DEvelopment. Front Genet. 2012;3:270. doi:10.3389/fgene.2012.00270.
Wang Y, Li J, Zhang Y, Yin H, Han B. CRNDE, a long-noncoding RNA, promotes glioma cell growth and invasion through mTOR signaling. Cancer Lett. 2015;367(2):122–8. doi:10.1016/j.canlet.2015.03.027.
Zheng J, Li XD, Wang P, Liu XB, Xue YX, Hu Y, et al. CRNDE affects the malignant biological characteristics of human glioma stem cells by negatively regulating miR-186. Oncotarget. 2015;6(28):25339–55. doi:10.18632/oncotarget.4509.
Louis DN, Ohgaki H, Wiestler OD, Cavenee WK, Burger PC, Jouvet A, et al. The 2007 WHO classification of tumours of the central nervous system. Acta Neuropathol. 2007;114(2):97–109. doi:10.1007/s00401-007-0243-4.
Sun S, Lee D, Ho AS, Pu JK, Zhang XQ, Lee NP, et al. Inhibition of prolyl 4-hydroxylase, beta polypeptide (P4HB) attenuates temozolomide resistance in malignant glioma via the endoplasmic reticulum stress response (ERSR) pathways. Neuro-Oncology. 2013;15(5):562–77. doi:10.1093/neuonc/not005.
Wong ST, Zhang XQ, Zhuang JT, Chan HL, Li CH, Leung GK. MicroRNA-21 inhibition enhances in vitro chemosensitivity of temozolomide-resistant glioblastoma cells. Anticancer Res. 2012;32(7):2835–41.
Seike M, Goto A, Okano T, Bowman ED, Schetter AJ, Horikawa I, et al. MiR-21 is an EGFR-regulated anti-apoptotic factor in lung cancer in never-smokers. Proc Natl Acad Sci USA. 2009;106(29):12085–90. doi:10.1073/pnas.0905234106.
Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(T)(−Delta Delta C) method. Methods. 2001;25(4):402–8. doi:10.1006/meth.2001.1262.
Vichai V, Kirtikara K. Sulforhodamine B colorimetric assay for cytotoxicity screening. Nat Protoc. 2006;1(3):1112–6. doi:10.1038/nprot.2006.179.
Riccardi C, Nicoletti I. Analysis of apoptosis by propidium iodide staining and flow cytometry. Nat Protoc. 2006;1(3):1458–61. doi:10.1038/nprot.2006.238.
Adams JM, Cory S. The Bcl-2 apoptotic switch in cancer development and therapy. Oncogene. 2007;26(9):1324–37. doi:10.1038/sj.onc.1210220.
Si ML, Zhu S, Wu H, Lu Z, Wu F, Mo YY. miR-21-mediated tumor growth. Oncogene. 2007;26(19):2799–803. doi:10.1038/sj.onc.1210083.
Ellis BC, Graham LD, Molloy PL. CRNDE, a long non-coding RNA responsive to insulin/IGF signaling, regulates genes involved in central metabolism. Biochim Biophys Acta. 2014;1843(2):372–86. doi:10.1016/j.bbamcr.2013.10.016.
Huang PH, Xu AM, White FM. Oncogenic EGFR signaling networks in glioma. Sci Signal. 2009;2(87):re6. doi:10.1126/scisignal.287re6.
Hatanpaa KJ, Burma S, Zhao D, Habib AA. Epidermal growth factor receptor in glioma: signal transduction, neuropathology, imaging, and radioresistance. Neoplasia. 2010;12(9):675–84.
Kadonaga JT. Regulation of RNA polymerase II transcription by sequence-specific DNA binding factors. Cell. 2004;116(2):247–57.
Jaenisch R, Bird A. Epigenetic regulation of gene expression: how the genome integrates intrinsic and environmental signals. Nat Genet. 2003;33:245–54. doi:10.1038/ng1089.
Yilmaz A, Grotewold E. Components and mechanisms of regulation of gene expression. Methods Mol Biol. 2010;674:23–32. doi:10.1007/978-1-60761-854-6_2.
Zhang XQ, Leung GK. Long non-coding RNAs in glioma: functional roles and clinical perspectives. Neurochem Int. 2014;77:78–85. doi:10.1016/j.neuint.2014.05.008.
Eisele G, Weller M. Targeting apoptosis pathways in glioblastoma. Cancer Lett. 2013;332(2):335–45. doi:10.1016/j.canlet.2010.12.012.
Steinbach JP, Weller M. Apoptosis in gliomas: molecular mechanisms and therapeutic implications. J Neuro-Oncol. 2004;70(2):247–56. doi:10.1007/s11060-004-2753-4.
Krakstad C, Chekenya M. Survival signalling and apoptosis resistance in glioblastomas: opportunities for targeted therapeutics. Mol Cancer. 2010;9:135. doi:10.1186/1476-4598-9-135.
Kelly PN, Strasser A. The role of Bcl-2 and its pro-survival relatives in tumourigenesis and cancer therapy. Cell Death Differ. 2011;18(9):1414–24. doi:10.1038/cdd.2011.17.
Martinou JC, Youle RJ. Mitochondria in apoptosis: Bcl-2 family members and mitochondrial dynamics. Dev Cell. 2011;21(1):92–101. doi:10.1016/j.devcel.2011.06.017.
Gross A, McDonnell JM, Korsmeyer SJ. BCL-2 family members and the mitochondria in apoptosis. Genes Dev. 1999;13(15):1899–911.
Adams JM, Cory S. Bcl-2-regulated apoptosis: mechanism and therapeutic potential. Curr Opin Immunol. 2007;19(5):488–96. doi:10.1016/j.coi.2007.05.004.
Zheng J, Liu XB, Wang P, Xue YX, Ma J, Qu CB, et al. CRNDE promotes malignant progression of glioma by attenuating miR-384/PIWIL4/STAT3 Axis. Mol Ther. 2016;24(7):1199–215.
Liu T, Zhang X, Yang YM, Du LT, Wang CX. Increased expression of the long noncoding RNA CRNDE-h indicates a poor prognosis in colorectal cancer, and is positively correlated with IRX5 mRNA expression. OncoTargets Ther. 2016;9:1437–48. doi:10.2147/OTT.S98268.
Szafron LM, Balcerak A, Grzybowska EA, Pienkowska-Grela B, Podgorska A, Zub R, et al. The putative oncogene, CRNDE, is a negative prognostic factor in ovarian cancer patients. Oncotarget. 2015;6(41):43897–910. doi:10.18632/oncotarget.6016.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
None.
Conflict of Interest
The authors declare no conflict of interest.
Electronic supplementary material
ESM 1
(DOCX 62 kb)
Rights and permissions
About this article
Cite this article
Kiang, K.MY., Zhang, XQ., Zhang, G.P. et al. CRNDE Expression Positively Correlates with EGFR Activation and Modulates Glioma Cell Growth. Targ Oncol 12, 353–363 (2017). https://doi.org/10.1007/s11523-017-0488-3
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11523-017-0488-3