Abstract
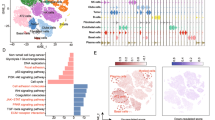
Adenosine-to-inosine (A-to-I) RNA editing is a widespread posttranscriptional modification that has been shown to play an important role in tumorigenesis. Here, we evaluated a total of 19,316 RNA editing sites in the tissues of 80 lung adenocarcinoma (LUAD) patients from our Nanjing Lung Cancer Cohort (NJLCC) and 486 LUAD patients from the TCGA database. The global RNA editing level was significantly increased in tumor tissues and was highly heterogeneous across patients. The high RNA editing level in tumors was attributed to both RNA (ADAR1 expression) and DNA alterations (mutation load). Consensus clustering on RNA editing sites revealed a new molecular subtype (EC3) that was associated with the poorest prognosis of LUAD patients. Importantly, the new classification was independent of classic molecular subtypes based on gene expression or DNA methylation. We further proposed a simplified model including eight RNA editing sites to accurately distinguish the EC3 subtype in our patients. The model was further validated in the TCGA dataset and had an area under the curve (AUC) of the receiver operating characteristic curve of 0.93 (95%CI: 0.91–0.95). In addition, we found that LUAD cell lines with the EC3 subtype were sensitive to four chemotherapy drugs. These findings highlighted the importance of RNA editing events in the tumorigenesis of LUAD and provided insight into the application of RNA editing in the molecular subtyping and clinical treatment of cancer.
Similar content being viewed by others
References
Agarwal, V., Bell, G.W., Nam, J.W., and Bartel, D.P. (2015). Predicting effective microRNA target sites in mammalian mRNAs. eLife 4, e05005.
Amin, E.M., Liu, Y., Deng, S., Tan, K.S., Chudgar, N., Mayo, M.W., Sanchez-Vega, F., Adusumilli, P.S., Schultz, N., and Jones, D.R. (2017). The RNA-editing enzyme ADAR promotes lung adenocarcinoma migration and invasion by stabilizing FAK. Sci Signal 10, eaah3941.
Bass, B.L. (2002). RNA editing by adenosine deaminases that act on RNA. Annu Rev Biochem 71, 817–846.
Bray, F., Ferlay, J., Soerjomataram, I., Siegel, R.L., Torre, L.A., and Jemal, A. (2018). Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 68, 394–424.
Cancer Genome Atlas Research, N. (2014). Comprehensive molecular profiling of lung adenocarcinoma. Nature 511, 543–550.
Chan, T.H.M., Qamra, A., Tan, K.T., Guo, J., Yang, H., Qi, L., Lin, J.S., Ng, V.H.E., Song, Y., Hong, H., et al. (2016). ADAR-mediated RNA editing predicts progression and prognosis of gastric cancer. Gastroenterology 151, 637–650.e10.
Chen, F., Zhang, Y., Parra, E., Rodriguez, J., Behrens, C., Akbani, R., Lu, Y., Kurie, J.M., Gibbons, D.L., Mills, G.B., et al. (2017a). Multiplatform-based molecular subtypes of non-small-cell lung cancer. Oncogene 36, 1384–1393.
Chen, J., Wang, L., Wang, F., Liu, J., and Bai, Z. (2020). Genomic identification of RNA editing through integrating omics datasets and the clinical relevance in hepatocellular carcinoma. Front Oncol 10, 37.
Chen, L., Li, Y., Lin, C.H., Chan, T.H.M., Chow, R.K.K., Song, Y., Liu, M., Yuan, Y.F., Fu, L., Kong, K.L., et al. (2013). Recoding RNA editing of AZIN1 predisposes to hepatocellular carcinoma. Nat Med 19, 209–216.
Chen, Y.B., Liao, X.Y., Zhang, J.B., Wang, F., Qin, H.D., Zhang, L., Shugart, Y.Y., Zeng, Y.X., and Jia, W.H. (2017b). ADAR2 functions as a tumor suppressor via editing IGFBP7 in esophageal squamous cell carcinoma. Int J Oncol 50, 622–630.
Dobin, A., Davis, C.A., Schlesinger, F., Drenkow, J., Zaleski, C., Jha, S., Batut, P., Chaisson, M., and Gingeras, T.R. (2013). Star: Ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21.
Ellrott, K., Bailey, M.H., Saksena, G., Covington, K.R., Kandoth, C., Stewart, C., Hess, J., Ma, S., Chiotti, K.E., McLellan, M., et al. (2018). Scalable open science approach for mutation calling of tumor exomes using multiple genomic pipelines. Cell Syst 6, 271–281.e7.
Friedman, J., Hastie, T., and Tibshirani, R. (2010). Regularization paths for generalized linear models via coordinate descent. J Stat Soft 33, 1–22.
Fumagalli, D., Gacquer, D., Rothé, F., Lefort, A., Libert, F., Brown, D., Kheddoumi, N., Shlien, A., Konopka, T., Salgado, R., et al. (2015). Principles governing A-to-I RNA editing in the breast cancer transcriptome. Cell Rep 13, 277–289.
Galeano, F., Rossetti, C., Tomaselli, S., Cifaldi, L., Lezzerini, M., Pezzullo, M., Boldrini, R., Massimi, L., Di Rocco, C.M., Locatelli, F., et al. (2013). ADAR2-editing activity inhibits glioblastoma growth through the modulation of the CDC14B/Skp2/p21/p27 axis. Oncogene 32, 998–1009.
Geeleher, P., Zhang, Z., Wang, F., Gruener, R.F., Nath, A., Morrison, G., Bhutra, S., Grossman, R.L., and Huang, R.S. (2017). Discovering novel pharmacogenomic biomarkers by imputing drug response in cancer patients from large genomics studies. Genome Res 27, 1743–1751.
Group, P.T.C., Calabrese, C., Davidson, N.R., Demircioğlu, D., Fonseca, N. A., He, Y., Kahles, A., Lehmann, K.V., Liu, F., Shiraishi, Y., et al. (2020). Genomic basis for RNA alterations in cancer. Nature 578, 129–136.
Han, L., Diao, L., Yu, S., Xu, X., Li, J., Zhang, R., Yang, Y., Werner, H.M. J., Eterovic, A.K., Yuan, Y., et al. (2015). The genomic landscape and clinical relevance of A-to-I RNA editing in human cancers. Cancer Cell 28, 515–528.
Heale, B.S.E., Keegan, L.P., McGurk, L., Michlewski, G., Brindle, J., Stanton, C.M., Caceres, J.F., and O’Connell, M.A. (2009). Editing independent effects of adars on the miRNA/siRNA pathways. EMBO J 28, 3145–3156.
Jevremovic, D., Billadeau, D.D., Schoon, R.A., Dick, C.J., and Leibson, P. J. (2001). Regulation of NK cell-mediated cytotoxicity by the adaptor protein 3BP2. J Immunol 166, 7219–7228.
Kent, W.J. (2002). BLAT—The BLAST-like alignment tool. Genome Res 12, 656–664.
Liao, Y., Smyth, G.K., and Shi, W. (2014). FeatureCounts: An efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930.
Liu, J., Lichtenberg, T., Hoadley, K.A., Poisson, L.M., Lazar, A.J., Cherniack, A.D., Kovatich, A.J., Benz, C.C., Levine, D.A., Lee, A.V., et al. (2018). An integrated TCGA pan-cancer clinical data resource to drive high-quality survival outcome analytics. Cell 173, 400–416.e11.
Lortet-Tieulent, J., Soerjomataram, I., Ferlay, J., Rutherford, M., Weiderpass, E., and Bray, F. (2014). International trends in lung cancer incidence by histological subtype: Adenocarcinoma stabilizing in men but still increasing in women. Lung Cancer 84, 13–22.
Nakamura, H., and Saji, H. (2014). A worldwide trend of increasing primary adenocarcinoma of the lung. Surg Today 44, 1004–1012.
Nishikura, K. (2010). Functions and regulation of RNA editing by adar deaminases. Annu Rev Biochem 79, 321–349.
Nishikura, K. (2016). A-to-I editing of coding and non-coding RNAs by ADARs. Nat Rev Mol Cell Biol 17, 83–96.
Oakes, E., Anderson, A., Cohen-Gadol, A., and Hundley, H.A. (2017). Adenosine deaminase that acts on RNA 3 (ADAR3) binding to glutamate receptor subunit B pre-mRNA inhibits RNA editing in glioblastoma. J Biol Chem 292, 4326–4335.
Osmani, L., Askin, F., Gabrielson, E., and Li, Q.K. (2018). Current WHO guidelines and the critical role of immunohistochemical markers in the subclassification of non-small cell lung carcinoma (NSCLC): Moving from targeted therapy to immunotherapy. Semin Cancer Biol 52, 103–109.
Park, E., Williams, B., Wold, B.J., and Mortazavi, A. (2012). RNA editing in the human encode RNA-seq data. Genome Res 22, 1626–1633.
Paz-Yaacov, N., Bazak, L., Buchumenski, I., Porath, H.T., Danan-Gotthold, M., Knisbacher, B.A., Eisenberg, E., and Levanon, E.Y. (2015). Elevated RNA editing activity is a major contributor to transcriptomic diversity in tumors. Cell Rep 13, 267–276.
Paz, N., Levanon, E.Y., Amariglio, N., Heimberger, A.B., Ram, Z., Constantini, S., Barbash, Z.S., Adamsky, K., Safran, M., Hirschberg, A., et al. (2007). Altered adenosine-to-inosine RNA editing in human cancer. Genome Res 17, 1586–1595.
Peng, X., Xu, X., Wang, Y., Hawke, D.H., Yu, S., Han, L., Zhou, Z., Mojumdar, K., Jeong, K.J., Labrie, M., et al. (2018). A-to-I RNA editing contributes to proteomic diversity in cancer. Cancer Cell 33, 817–828.e7.
Peng, Z., Cheng, Y., Tan, B.C.M., Kang, L., Tian, Z., Zhu, Y., Zhang, W., Liang, Y., Hu, X., Tan, X., et al. (2012). Comprehensive analysis of RNA-seq data reveals extensive RNA editing in a human transcriptome. Nat Biotechnol 30, 253–260.
Picardi, E., D’Erchia, A.M., Lo Giudice, C., and Pesole, G. (2017). Rediportal: A comprehensive database of A-to-I RNA editing events in humans. Nucleic Acids Res 45, D750–D757.
Piskol, R., Ramaswami, G., and Li, J.B. (2013). Reliable identification of genomic variants from RNA-seq data. Am J Hum Genet 93, 641–651.
Qin, Y.R., Qiao, J.J., Chan, T.H.M., Zhu, Y.H., Li, F.F., Liu, H., Fei, J., Li, Y., Guan, X.Y., and Chen, L. (2014). Adenosine-to-inosine RNA editing mediated by ADARs in esophageal squamous cell carcinoma. Cancer Res 74, 840–851.
Ramaswami, G., Lin, W., Piskol, R., Tan, M.H., Davis, C., and Li, J.B. (2012). Accurate identification of human Alu and non-Alu RNA editing sites. Nat Methods 9, 579–581.
Rosseel, Y. (2012). Lavaan: An R package for structural equation modeling and more. Version 0.5-12 (beta). J Stat Soft 48, 1–36.
Serrano-Candelas, E., Ainsua-Enrich, E., Navinés-Ferrer, A., Rodrigues, P., García-Valverde, A., Bazzocco, S., Macaya, I., Arribas, J., Serrano, C., Sayós, J., et al. (2018). Silencing of adaptor protein SH3BP2 reduces KIT/PDGFRA receptors expression and impairs gastrointestinal stromal tumors growth. Mol Oncol 12, 1383–1397.
Shigeyasu, K., Okugawa, Y., Toden, S., Miyoshi, J., Toiyama, Y., Nagasaka, T., Takahashi, N., Kusunoki, M., Takayama, T., Yamada, Y., et al. (2018). AZIN1 RNA editing confers cancer stemness and enhances oncogenic potential in colorectal cancer. JCI Insight 3.
Silvestris, D.A., Picardi, E., Cesarini, V., Fosso, B., Mangraviti, N., Massimi, L., Martini, M., Pesole, G., Locatelli, F., and Gallo, A. (2019). Dynamic inosinome profiles reveal novel patient stratification and gender-specific differences in glioblastoma. Genome Biol 20, 33.
Song, Y., An, O., Ren, X., Chan, T.H.M., Tay, D.J.T., Tang, S.J., Han, J., Hong, H.Q., Ng, V.H.E., Ke, X., et al. (2021). RNA editing mediates the functional switch of COPA in a novel mechanism of hepatocarcinogenesis. J Hepatol 74, 135–147.
Tingley, D., Yamamoto, T., Hirose, K., Keele, L., Imai, K. (2014). Mediation: R package for causal mediation analysis. J Stat Soft 59.
Tomaselli, S., Galeano, F., Alon, S., Raho, S., Galardi, S., Polito, V.A., Presutti, C., Vincenti, S., Eisenberg, E., Locatelli, F., et al. (2015). Modulation of microRNA editing, expression and processing by ADAR2 deaminase in glioblastoma. Genome Biol 16, 5.
Vargas, A.J., and Harris, C.C. (2016). Biomarker development in the precision medicine era: Lung cancer as a case study. Nat Rev Cancer 16, 525–537.
Wang, C., Yin, R., Dai, J., Gu, Y., Cui, S., Ma, H., Zhang, Z., Huang, J., Qin, N., Jiang, T., et al. (2018). Whole-genome sequencing reveals genomic signatures associated with the inflammatory microenvironments in chinese NSCLC patients. Nat Commun 9, 2054.
Wilkerson, M.D., and Hayes, D.N. (2010). ConsensusClusterPlus: A class discovery tool with confidence assessments and item tracking. Bioinformatics 26, 1572–1573.
Yu, G., Wang, L.G., Han, Y., and He, Q.Y. (2012). ClusterProfiler: An R package for comparing biological themes among gene clusters. OMICS 16, 284–287.
Zhang, M., Fritsche, J., Roszik, J., Williams, L.J., Peng, X., Chiu, Y., Tsou, C.C., Hoffgaard, F., Goldfinger, V., Schoor, O., et al. (2018). RNA editing derived epitopes function as cancer antigens to elicit immune responses. Nat Commun 9, 3919.
Acknowledgements
This work was supported by the National Natural Science Foundation of China (81922061, 82072579, 81521004, 81973123 and 81871885), the National Key Research and Development Project (2017YFC0907905), and Research Unit of Prospective Cohort of Cardiovascular Diseases and Cancer, Chinese Academy of Medical Sciences (2019RU038). We are grateful to the patients who participated in this study and all the researchers, clinicians and technical and administrative staff who have made this work possible.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Compliance and ethics The author(s) declare that they have no conflict of interest. All participants signed an informed consent form that was approved by the local internal review boards or ethics committees (Nanjing, China).
Supporting Information
Rights and permissions
About this article
Cite this article
Wang, C., Huang, M., Chen, C. et al. Identification of A-to-I RNA editing profiles and their clinical relevance in lung adenocarcinoma. Sci. China Life Sci. 65, 19–32 (2022). https://doi.org/10.1007/s11427-020-1928-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11427-020-1928-0