Abstract
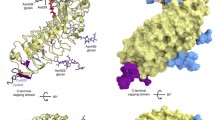
Leucine rich repeat (LRR) domain, characterized by a repetitive sequence pattern rich in leucine residues, is a universal protein-protein interaction motif present in all life forms. LRR repeats interrupted by sequences of 30–70 residues (termed island domain, ID) have been found in some plant LRR receptor-like kinases (RLKs) and animal Toll-like receptors (TLR7-9). Recent studies provide insight into how a single ID is structurally integrated into an LRR protein. However, structural information on an LRR protein with two IDs is lacking. The receptor-like protein kinase 2 (RPK2) is an LRR-RLK and has important roles in controlling plant growth and development by perception and transduction of hormone signal. Here we present the crystal structure of the extracellular LRR domain of RPK2 (RPK2-LRR) containing two IDs from Arabidopsis. The structure reveals that both of the IDs are helical and located at the central region of the single RPK2-LRR solenoid. One of them binds to the inner surface of the solenoid, whereas the other one mainly interacts with the lateral side. Unexpectedly, a long loop immediately following the N-terminal capping domain of RPK2-LRR is presented toward and sandwiched between the two IDs, further stabilizing their embedding to the LRR solenoid. A potential ligand binding site formed by the two IDs and the solenoid is located at the C-terminal side of RPK2-LRR. The structural information of RPK2-LRR broadens our understanding toward the large family of LRR proteins and provides insight into RPK2-mediated signaling.
Article PDF
Similar content being viewed by others
References
Kobe B, Kajava AV. The leucine-rich repeat as a protein recognition motif. Curr Opin Struct Biol, 2001, 11:725–732
Botos I, Segal DM, Davies DR. The structural biology of Toll-like receptors. Structure, 2011, 19:447–459
She J, Han Z, Zhou B, Chai J. Structural basis for differential recognition of brassinolide by its receptors. Protein Cell, 2013, 4:475–482
Liu P, Hu Z, Zhou B, Liu S, Chai J. Crystal structure of an LRR protein with two solenoids. Cell Res, 2013, 23:303–305
She J, Han Z, Kim TW, Wang J, Cheng W, Chang J, Shi S, Wang J, Yang M, Wang ZY, Chai J. Structural insight into brassinosteroid perception by BRI1. Nature, 2011, 474:472–476
Hothorn M, Belkhadir Y, Dreux M, Dabi T, Noel JP, Wilson IA, Chory J. Structural basis of steroid hormone perception by the receptor kinase BRI1. Nature, 2011, 474:467–471
Shiu SH, Bleecker AB. Plant receptor-like kinase gene family: diversity, function, and signaling. Science’s STKE, 2001, 2001:re22
Matsushima N, Mikami T, Tanaka T, Miyashita H, Yamada K, Kuroki Y. Analyses of non-leucine-rich repeat (non-LRR) regions intervening between LRRs in proteins. Biochim Biophys Acta, 2009, 1790:1217–1237
Schoof H, Lenhard M, Haecker A, Shinozaki K, Fukuda H, Sawa S. The stem cell population of Arabidopsis shoot meristems is maintained by a regulatory loop between the CLAVATA and WUSCHEL genes. Cell, 2000, 100:635–644
Brand U, Fletcher JC, Hobe M, Meyerowitz EM, Simon R. Dependence of stem cell fate in Arabidopsis on a feedback loop regulated by CLV3 activity. Science, 2000, 289:617–619
Stahl Y, Simon R. Plant primary meristems: shared functions and regulatory mechanisms. Curr Opin Plant Biol, 2010, 13:53–58
Gao X, Guo Y. CLE peptides in plants: proteolytic processing, structure-activity relationship, and ligand-receptor interaction. J Integr Plant Biol, 2012, 54:738–745
Miwa H, Kinoshita A, Fukuda H, Sawa S. Plant meristems: CLAVATA3/ESR-related signaling in the shoot apical meristem and the root apical meristem. J Plant Res, 2009, 122:31–39
Kinoshita A, Betsuyaku S, Osakabe Y, Mizuno S, Nagawa S, Stahl Y, Simon R, Yamaguchi-Shinozaki K, Fukuda H, Sawa S. RPK2 is an essential receptor-like kinase that transmits the CLV3 signal in Arabidopsis. Development, 2010, 137:3911–3920
Sawa S, Tabata R. RPK2 functions in diverged CLE signaling. Plant Signal Behav, 2011, 6:86–88
Guo Y, Ni J, Denver R, Wang X, Clark SE. Mechanisms of molecular mimicry of plant CLE peptide ligands by the parasitic nematode. Plant Physiol, 2011, 157:476–484
Replogle A, Wang J, Paolillo V, Smeda J, Kinoshita A, Durbak A, Tax FE, Wang X, Sawa S, Mitchum MG. Synergistic interaction of CLAVATA1, CLAVATA2, and RECEPTOR-LIKE PROTEIN KINASE 2 in cyst nematode parasitism of Arabidopsis. Mol Plant Microbe In, 2013, 26:87–96
Nodine MD, Yadegari R, Tax FE. RPK1 and TOAD2 are two receptor-like kinases redundantly required for Arabidopsis embryonic pattern formation. Dev Cell, 2007, 12:943–956
Nodine MD, Tax FE. Two receptor-like kinases required together for the establishment of Arabidopsis cotyledon primordia. Dev Biol, 2008, 314:161–170
Mizuno S, Osakabe Y, Maruyama K, Ito T, Osakabe K, Sato T, Shinozaki K, Yamaguchi-Shinozaki K. Receptor-like protein kinase 2 (RPK2) is a novel factor controlling anther development in Arabidopsis thaliana. Plant J, 2007, 50:751–766
Zhao D. Control of anther cell differentiation: a teamwork of receptor-like kinases. Sex Plant Reprod, 2009, 22:221–228
Otwinowski Z, Minor W. Processing of X-ray diffraction data collected in oscillation mode. Method Enzymol, 1997, 276:307–326
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr, 1994, D50:760–763
Emsley P, Cowtan KC. Model-building tools for molecular graphics. Acta Crystallogr D Biol Crystallogr, 2004, 60:2126–2132
Adams P, Grosse KR, Huang LW, Ioerger TR, McCoy AJ, Moriarty NW, Read RJ, Sacchettini JC, Sauter NK, Terwilliger TC. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr, 2002, D58:1948–1954
DeLano WL. PyMOL molecular viewer. 2002, http://www.pymol.org/
Kim TW, Wang ZY. Brassinosteroid signal transduction from receptor kinases to transcription factors. Annu Rev Plant Biol, 2010, 61:681–704
Sheard LB, Tan X, Mao H, Withers J, Ben-Nissan G, Hinds TR, Kobayashi Y, Hsu FF, Sharon M, Browse J, He SY, Rizo J, Howe GA, Zheng N. Jasmonate perception by inositol-phosphate-potentiated COI1-JAZ co-receptor. Nature, 2010, 468:400–405
Author information
Authors and Affiliations
Corresponding author
Additional information
This article is published with open access at link.springer.com
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 2.0 International License (https://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Song, W., Han, Z., Sun, Y. et al. Crystal structure of a plant leucine rich repeat protein with two island domains. Sci. China Life Sci. 57, 137–144 (2014). https://doi.org/10.1007/s11427-013-4586-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11427-013-4586-x