Abstract
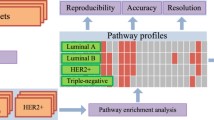
Selecting differentially expressed genes (DEGs) is one of the most important tasks in microarray applications for studying multi-factor diseases including cancers. However, the small samples typically used in current microarray studies may only partially reflect the widely altered gene expressions in complex diseases, which would introduce low reproducibility of gene lists selected by statistical methods. Here, by analyzing seven cancer datasets, we showed that, in each cancer, a wide range of functional modules have altered gene expressions and thus have high disease classification abilities. The results also showed that seven modules are shared across diverse cancers, suggesting hints about the common mechanisms of cancers. Therefore, instead of relying on a few individual genes whose selection is hardly reproducible in current microarray experiments, we may use functional modules as functional signatures to study core mechanisms of cancers and build robust diagnostic classifiers.
Article PDF
Similar content being viewed by others
References
Tusher V G, Tibshirani R, Chu G. Significance analysis of microarrays applied to the ionizing radiation response. Proc Natl Acad Sci USA, 2001, 98: 5116–5121, 1:CAS:528:DC%2BD3MXjt1Ons7w%3D, 10.1073/pnas.091062498, 11309499
Jeffery I B, Higgins D G, Culhane A C. Comparison and evaluation of methods for generating differentially expressed gene lists from microarray data. BMC Bioinformatics, 2006, 7: 359, 10.1186/1471-2105-7-359, 16872483
Chen J J, Wang S J, Tsai C A, et al. Selection of differentially expressed genes in microarray data analysis. Pharmacogenomics J, 2006
Cui X, Churchill G A. Statistical tests for differential expression in cDNA microarray experiments. Genome Biol, 2003, 4: 210, 10.1186/gb-2003-4-4-210, 12702200
Pavlidis P, Li Q, Noble W S. The effect of replication on gene expression microarray experiments. Bioinformatics, 2003, 19: 1620–1627, 1:CAS:528:DC%2BD3sXnt1Gqu7s%3D, 10.1093/bioinformatics/btg227, 12967957
Zhang M, Yao C, Guo Z, et al. Apparently low reproducibility of true differential expression discoveries in microarray studies. Bioinformatics, 2008, 24: 2057–2063, 1:CAS:528:DC%2BD1cXhtFWhurrK, 10.1093/bioinformatics/btn365, 18632747
Guo L, Lobenhofer E K, Wang C, et al. Rat toxicogenomic study reveals analytical consistency across microarray platforms. Nat Biotechnol, 2006, 24: 1162–1169, 1:CAS:528:DC%2BD28XptlSlsLc%3D, 10.1038/nbt1238, 17061323
Shi L, Reid L H, Jones W D, et al. The MicroArray Quality Control (MAQC) project shows inter- and intraplatform reproducibility of gene expression measurements. Nat Biotechnol, 2006, 24: 1151–1161, 1:CAS:528:DC%2BD28XptlSlsb4%3D, 10.1038/nbt1239, 16964229
Allison D B, Cui X, Page G P, et al. Microarray data analysis: from disarray to consolidation and consensus. Nat Rev Genet, 2006, 7: 55–65, 1:CAS:528:DC%2BD2MXhtlanu73F, 10.1038/nrg1749, 16369572
Wolfe C J, Kohane I S, Butte A J. Systematic survey reveals general applicability of “guilt-by-association” within gene coexpression networks. BMC Bioinformatics, 2005, 6: 227, 10.1186/1471-2105-6-227, 16162296
Breitling R, Armengaud P, Amtmann A, et al. Rank products: a simple, yet powerful, new method to detect differentially regulated genes in replicated microarray experiments. FEBS Lett, 2004, 573: 83–92, 1:CAS:528:DC%2BD2cXmvFyrtr4%3D, 10.1016/j.febslet.2004.07.055, 15327980
Yang D, Li Y, Xiao H, et al. Gaining confidence in biological interpretation of the microarray data: the functional consistence of the significant GO categories. Bioinformatics, 2008, 24: 265–271, 1:CAS:528:DC%2BD1cXptlWhtg%3D%3D, 10.1093/bioinformatics/btm558, 18006543
Khalil I G, Hill C. Systems biology for cancer. Curr Opin Oncol, 2005, 17: 44–48, 1:STN:280:DC%2BD2cnjtFKnug%3D%3D, 10.1097/01.cco.0000150951.38222.16, 15608512
Morley M, Molony C M, Weber T M, et al. Genetic analysis of genome-wide variation in human gene expression. Nature, 2004, 430: 743–747, 1:CAS:528:DC%2BD2cXmsVGls7k%3D, 10.1038/nature02797, 15269782
Guo Z, Zhang T, Li X, et al. Towards precise classification of cancers based on robust gene functional expression profiles. BMC Bioinformatics, 2005, 6: 58, 10.1186/1471-2105-6-58, 15774002
Barrett T, Troup D B, Wilhite S E, et al. NCBI GEO: mining tens of millions of expression profiles-database and tools update. Nucleic Acids Res, 2007, 35: 760–765, 10.1093/nar/gkl887
Garber M E, Troyanskaya O G, Schluens K, et al. Diversity of gene expression in adenocarcinoma of the lung. Proc Natl Acad Sci USA, 2001, 98: 13784–13789, 1:CAS:528:DC%2BD3MXovVynt7o%3D, 10.1073/pnas.241500798, 11707590
Bhattacharjee A, Richards W G, Staunton J, et al. Classification of human lung carcinomas by mRNA expression profiling reveals distinct adenocarcinoma subclasses. Proc Natl Acad Sci USA, 2001, 98: 13790–13795, 1:CAS:528:DC%2BD3MXovVynt7s%3D, 10.1073/pnas.191502998, 11707567
Galamb O, Gyorffy B, Sipos F, et al. Inflammation, adenoma and cancer: objective classification of colon biopsy specimens with gene expression signature. Dis Markers, 2008, 25: 1–16, 1:CAS:528:DC%2BD1cXhtVWksbvO, 18776587
Sabates-Bellver J, Van der Flier L G, de Palo M, et al. Transcriptome profile of human colorectal adenomas. Mol Cancer Res, 2007, 5: 1263–1275, 1:CAS:528:DC%2BD1cXjtVCgtA%3D%3D, 10.1158/1541-7786.MCR-07-0267, 18171984
Lapointe J, Li C, Higgins J P, et al. Gene expression profiling identifies clinically relevant subtypes of prostate cancer. Proc Natl Acad Sci USA, 2004, 101: 811–816, 1:CAS:528:DC%2BD2cXhsVWhu7o%3D, 10.1073/pnas.0304146101, 14711987
Chen X, Leung S Y, Yuen S T, et al. Variation in gene expression patterns in human gastric cancers. Mol Biol Cell, 2003, 14: 3208–3215, 1:CAS:528:DC%2BD3sXmvFWksr0%3D, 10.1091/mbc.E02-12-0833, 12925757
Chen X, Higgins J, Cheung S T, et al. Novel endothelial cell markers in hepatocellular carcinoma. Mod Pathol, 2004, 17: 1198–1210, 1:CAS:528:DC%2BD2cXovFOqtr0%3D, 10.1038/modpathol.3800167, 15154008
Rhodes D R, Chinnaiyan A M. Integrative analysis of the cancer transcriptome. Nat Genet, 2005, 37: 31–37, 10.1038/ng1570
Diehn M, Sherlock G, Binkley G, et al. SOURCE: a unified genomic resource of functional annotations, ontologies, and gene expression data. Nucleic Acids Res, 2003, 31: 219–223, 1:CAS:528:DC%2BD3sXhvFSnurg%3D, 10.1093/nar/gkg014, 12519986
Ashburner M, Ball C A, Blake J A, et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet, 2000, 25: 25–29, 1:CAS:528:DC%2BD3cXjtFSlsbc%3D, 10.1038/75556, 10802651
Dudoit S, Fridlyand J, Speed T. Comparison of discrimination methods for the classification of tumours using gene expression data. J Am Statisti Associ, 2002, 97: 77–87, 1:CAS:528:DC%2BD28Xht1SqtL7I, 10.1198/016214502753479248
Zhang H, Yu C Y, Singer B. Cell and tumor classification using gene expression data: construction of forests. Proc Natl Acad Sci USA, 2003, 100: 4168–4172, 1:CAS:528:DC%2BD3sXivFWjsrY%3D, 10.1073/pnas.0230559100, 12642676
Alexa A, Rahnenfuhrer J, Lengauer T. Improved scoring of functional groups from gene expression data by decorrelating GO graph structure. Bioinformatics, 2006, 22: 1600–1607, 1:CAS:528:DC%2BD28XmsVentro%3D, 10.1093/bioinformatics/btl140, 16606683
Falcon S, Gentleman R. Using GOstats to test gene lists for GO term association. Bioinformatics, 2007, 23: 257–258, 1:CAS:528:DC%2BD2sXotFGntA%3D%3D, 10.1093/bioinformatics/btl567, 17098774
Hanahan D, Weinberg R A. The hallmarks of cancer. Cell, 2000, 100: 57–70, 1:CAS:528:DC%2BD3cXks1CktA%3D%3D, 10.1016/S0092-8674(00)81683-9, 10647931
Olivier S, Robe P, Bours V. Can NF-kappaB be a target for novel and efficient anti-cancer agents? Biochem Pharmacol, 2006, 72: 1054–1068, 1:CAS:528:DC%2BD28XhtVagtr%2FO, 10.1016/j.bcp.2006.07.023, 16973133
Long Y M, Ye S, Rong J, et al. Nuclear factor kappa B: a marker of chemotherapy for human stage IV gastric carcinoma. World J Gastroenterol, 2008, 14: 4739–4744, 10.3748/wjg.14.4739, 18720533
Hartwell L H, Kastan M B. Cell cycle control and cancer. Science, 1994, 266: 1821–1828, 1:CAS:528:DyaK2MXislCjsr8%3D, 10.1126/science.7997877, 7997877
Rosengren S, Henson P M, Worthen G S. Migration-associated volume changes in neutrophils facilitate the migratory process in vitro. Am J Physiol, 1994, 267: 1623–1632
Saadoun S, Papadopoulos M C, Watanabe H, et al. Involvement of aquaporin-4 in astroglial cell migration and glial scar formation. J Cell Sci, 2005, 118: 5691–5698, 1:CAS:528:DC%2BD28XosFemtw%3D%3D, 10.1242/jcs.02680, 16303850
Larue L, Bellacosa A. Epithelial-mesenchymal transition in development and cancer: role of phosphatidylinositol 3′ kinase/AKT pathways. Oncogene, 2005, 24: 7443–7454, 1:CAS:528:DC%2BD2MXht1Wis7vE, 10.1038/sj.onc.1209091, 16288291
Jeanes A, Gottardi C J, Yap A S. Cadherins and cancer: how does cadherin dysfunction promote tumor progression? Oncogene, 2008, 27: 6920–6929, 1:CAS:528:DC%2BD1cXhsVWhu73O, 10.1038/onc.2008.343, 19029934
Hynes R O. Metastatic potential: generic predisposition of the primary tumor or rare, metastatic variants—or both? Cell, 2003, 113: 821–823, 1:CAS:528:DC%2BD3sXlsVSjtb8%3D, 10.1016/S0092-8674(03)00468-9, 12837240
Hallahan A R, Pritchard J I, Chandraratna R A, et al. BMP-2 mediates retinoid-induced apoptosis in medulloblastoma cells through a paracrine effect. Nat Med, 2003, 9: 1033–1038, 1:CAS:528:DC%2BD3sXlvV2ju7c%3D, 10.1038/nm904, 12872164
Chuang H Y, Lee E, Liu Y T, et al. Network-based classification of breast cancer metastasis. Mol Syst Biol, 2007, 3: 140, 10.1038/msb4100180, 17940530
Taylor I W, Linding R, Warde-Farley D, et al. Dynamic modularity in protein interaction networks predicts breast cancer outcome. Nat Biotechnol, 2009, 27: 199–204, 1:CAS:528:DC%2BD1MXht1ehsbs%3D, 10.1038/nbt.1522, 19182785
Zhang M, Zhang L, Zou J, et al. Evaluating reproducibility of differential expression discoveries in microarray studies by considering correlated molecular changes. Bioinformatics, 2009, 25: 1662–1668, 1:CAS:528:DC%2BD1MXnsFygtb4%3D, 10.1093/bioinformatics/btp295, 19417058
Segal E, Friedman N, Koller D, et al. A module map showing conditional activity of expression modules in cancer. Nat Genet, 2004, 36: 1090–1098, 1:CAS:528:DC%2BD2cXnvFamtbw%3D, 10.1038/ng1434, 15448693
Campagne F, Skrabanek L. Mining expressed sequence tags identifies cancer markers of clinical interest. BMC Bioinformatics, 2006, 7: 481, 10.1186/1471-2105-7-481, 17078886
Author information
Authors and Affiliations
Corresponding author
Additional information
Contributed equally to this work
This article is published with open access at Springerlink.com
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Yao, C., Zhang, M., Zou, J. et al. Functional modules with disease discrimination abilities for various cancers. Sci. China Life Sci. 54, 189–193 (2011). https://doi.org/10.1007/s11427-010-4129-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11427-010-4129-7