Abstract
Introduction
Autism spectrum disorders (ASD) is a group of neurodevelopmental disorders believed to have a multifactorial basis. Presently, diagnosis is based on behavioral and developmental signs in children before the age of 3 and no reliable clinical biomarkers are available for early detection.
Objectives
This study aimed to biochemically profile the cerebellum from post-mortem human brain from ASD sufferers (n = 11) and compare their profiles to that of age-matched controls (n = 11) with no known brain disorder.
Methods
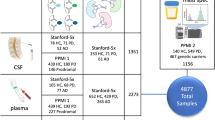
Using liquid chromatography combined with LTQ-Orbitrap mass spectrometry we detected 14,328 features in ESI+ mode in polar extracts of post-mortem brain.
Results
Of these only 37 were found to be statistically significantly different between ASD and controls (p < 0.05; fdr < 0.05). A panel of four features had a predictive power of 96.64 %, following statistical cross validation, for ASD detection. This model produced an AUC = 0.874 (CI 0.768–0.944) and a Fisher’s exact score of p = 4.50E−29.
Conclusion
Whilst at this time we were unable to chemically identify the four features of interest we believe that this study underscores the potential value of high resolution metabolomics for the study of ASD. Further characterization of the polar metabolome of post mortem ASD brains could lead to the identification of potential biomarkers and novel therapeutics for the disease. The development of accurate biomarkers could assist in the early detection of ASD and promote early intervention strategies to improve outcome.


Similar content being viewed by others
References
American Psychiatric Association (APA). (2013). Diagnostic and statistical manual of mental disorders: DSM-5 (5th ed.). Washington, DC: American Psychiatric Publishing.
Baio, J. (2012). Prevalence of autism spectrum disorders: Autism and developmental disabilities monitoring network, 14 sites, United States, 2008. Morbidity and Mortality Weekly Report. Surveillance Summaries, Vol. 61, No. 3. Atlanta, GA: Centers for Disease Control and Prevention.
Blatt, G. J. (2012). The neuropathology of autism. Scientifica (Cairo), 2012, 703675.
Chen, R., Jiao, Y., & Herskovits, E. H. (2011). Structural MRI in autism spectrum disorder. Pediatric Research, 69, 63R–68R.
Courchesne, E., Karns, C. M., Davis, H. R., Ziccardi, R., Carper, R. A., Tigue, Z. D., et al. (2001). Unusual brain growth patterns in early life in patients with autistic disorder: An MRI study. Neurology, 57, 245–254.
Currenti, S. A. (2010). Understanding and determining the etiology of autism. Cellular and Molecular Neurobiology, 30, 161–171.
Emond, P., Mavel, S., Aidoud, N., Nadal-Desbarats, L., Montigny, F., Bonnet-Brilhault, F., et al. (2013). GC-MS-based urine metabolic profiling of autism spectrum disorders. Analytical and Bioanalytical Chemistry, 405, 5291–5300.
Eriksson, L., Kettaneh-Wold, N., Trygg, J., Wikström, C., & Wold, S. (2006). Multi-and megavariate data analysis: Part I: basic principles and applications. MKS Umetrics AB.
Gowda, H., Ivanisevic, J., Johnson, C. H., Kurczy, M. E., Benton, H. P., Rinehart, D., et al. (2014). Interactive XCMS Online: Simplifying advanced metabolomic data processing and subsequent statistical analyses. Analytical Chemistry, 86, 6931–6939.
Graham, S. F., Chevallier, O. P., Roberts, D., Holscher, C., Elliott, C. T., & Green, B. D. (2013). Investigation of the human brain metabolome to identify potential markers for early diagnosis and therapeutic targets of Alzheimer’s disease. Analytical Chemistry, 85, 1803–1811.
Hallmayer, J., Cleveland, S., Torres, A., Phillips, J., Cohen, B., Torigoe, T., et al. (2011). Genetic heritability and shared environmental factors among twin pairs with autism. Archives of General Psychiatry, 68, 1095–1102.
Horai, H., Arita, M., Kanaya, S., Nihei, Y., Ikeda, T., Suwa, K., et al. (2010). MassBank: A public repository for sharing mass spectral data for life sciences. Journal of Mass Spectrometry, 45, 703–714.
Kaluzna-Czaplinska, J., Socha, E., & Rynkowski, J. (2010). Determination of homovanillic acid and vanillylmandelic acid in urine of autistic children by gas chromatography/mass spectrometry. Medical Science Monitor, 16, CR445–CR450.
Kuwabara, H., Yamasue, H., Koike, S., Inoue, H., Kawakubo, Y., Kuroda, M., et al. (2013). Altered metabolites in the plasma of autism spectrum disorder: A capillary electrophoresis time-of-flight mass spectroscopy study. PLoS One, 8, e73814.
Mavel, S., Nadal-Desbarats, L., Blasco, H., Bonnet-Brilhault, F., Barthelemy, C., Montigny, F., et al. (2013). 1H-13C NMR-based urine metabolic profiling in autism spectrum disorders. Talanta, 114, 95–102.
Ming, X., Stein, T. P., Barnes, V., Rhodes, N., & Guo, L. (2012). Metabolic perturbance in autism spectrum disorders: A metabolomics study. Journal of Proteome Research, 11, 5856–5862.
Nicholson, J. K., Connelly, J., Lindon, J. C., & Holmes, E. (2002). Metabonomics: A platform for studying drug toxicity and gene function. Nature Reviews Drug Discovery, 1, 153–161.
Nicholson, J. K., Lindon, J. C., & Holmes, E. (1999). ‘Metabonomics’: Understanding the metabolic responses of living systems to pathophysiological stimuli via multivariate statistical analysis of biological NMR spectroscopic data. Xenobiotica, 29, 1181–1189.
Pelphrey, K., Adolphs, R., & Morris, J. P. (2004). Neuroanatomical substrates of social cognition dysfunction in autism. Mental Retardation and Developmental Disabilities Research Reviews, 10, 259–271.
Ronald, A., Happe, F., Bolton, P., Butcher, L. M., Price, T. S., Wheelwright, S., et al. (2006). Genetic heterogeneity between the three components of the autism spectrum: A twin study. Journal of the American Academy of Child and Adolescent Psychiatry, 45, 691–699.
Rosenberg, R. E., Law, J. K., Yenokyan, G., McGready, J., Kaufmann, W. E., & Law, P. A. (2009). Characteristics and concordance of autism spectrum disorders among 277 twin pairs. Archives of Pediatrics and Adolescent Medicine, 163, 907–914.
Rossignol, D. A., & Frye, R. E. (2012). A review of research trends in physiological abnormalities in autism spectrum disorders: Immune dysregulation, inflammation, oxidative stress, mitochondrial dysfunction and environmental toxicant exposures. Molecular Psychiatry, 17, 389–401.
Sumner, L. W., Amberg, A., Barrett, D., Beale, M. H., Beger, R., Daykin, C. A., et al. (2007). Proposed minimum reporting standards for chemical analysis Chemical Analysis Working Group (CAWG) Metabolomics Standards Initiative (MSI). Metabolomics: Official Journal of the Metabolomic Society, 3, 211–221.
Sypert, G. W., & Alvord Jr, E. C. (1975). Cerebellar infarction: A clinicopathological study. Archives of Neurology, 32, 357–363.
Voineagu, I., & Yoo, H. J. (2013). Current progress and challenges in the search for autism biomarkers. Disease Markers, 35, 55–65.
West, P. R., Amaral, D. G., Bais, P., Smith, A. M., Egnash, L. A., Ross, M. E., et al. (2014). Metabolomics as a tool for discovery of biomarkers of autism spectrum disorder in the blood plasma of children. PLoS One, 9, e112445.
Wishart, D. S. (2009). Computational strategies for metabolite identification in metabolomics. Bioanalysis, 1, 1579–1596.
Wishart, D. S., Jewison, T., Guo, A. C., Wilson, M., Knox, C., Liu, Y., et al. (2013). HMDB 3.0–The human metabolome database in 2013. Nucleic Acids Research, 41, D801–D807.
Wishart, D. S., Knox, C., Guo, A. C., Eisner, R., Young, N., Gautam, B., et al. (2009). HMDB: A knowledgebase for the human metabolome. Nucleic Acids Research, 37, D603–D610.
Wishart, D. S., Tzur, D., Knox, C., Eisner, R., Guo, A. C., Young, N., et al. (2007). HMDB: The human metabolome database. Nucleic Acids Research, 35, D521–D526.
Xia, J., Mandal, R., Sinelnikov, I. V., Broadhurst, D., & Wishart, D. S. (2012). MetaboAnalyst 2.0–A comprehensive server for metabolomic data analysis. Nucleic Acids Research, 40, W127–W133.
Xia, J., Psychogios, N., Young, N., & Wishart, D. S. (2009). MetaboAnalyst: A web server for metabolomic data analysis and interpretation. Nucleic Acids Research, 37, W652–W660.
Xia, J., Sinelnikov, I. V., Han, B., & Wishart, D. S. (2015). MetaboAnalyst 3.0-making metabolomics more meaningful. Nucleic Acids Research, 43, W251–W257.
Yap, I. K., Angley, M., Veselkov, K. A., Holmes, E., Lindon, J. C., & Nicholson, J. K. (2010). Urinary metabolic phenotyping differentiates children with autism from their unaffected siblings and age-matched controls. Journal of Proteome Research, 9, 2996–3004.
Acknowledgments
We are exceedingly grateful to the University of Maryland Brain and Tissue Bank which is a Brain and Tissue Repository of the NIH NeuroBioBank for graciously providing the tissue samples for this study. Without their support this pilot project could not have been executed. In addition the authors would like to thank Dr. Rachel Hill (Queen’s University Belfast) for her help with organizing the raw data prior to both multivariate and univariate analyses.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Ethical approval
The NIH NeuroBioBank has full ethical approval for harvesting and storing PM human brain. In addition this study received full ethical approval from Beaumont Health Human Investigative Committee (Ref#2014-142).
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Fig. 1
The PCA scores plot of the complete data set to include controls (blue circles), ASD (red squares) and the pooled QC injections (black triangles). Supplementary material 1 (PDF 15 kb)
Supplementary File 1
All the fragmentation information at 3 NCE’s pertaining to the 4 ions of interest used to construct the predictive model in Fig. 2. Supplementary material 2 (XLSX 42 kb)
Supplementary Table 1
The details such as age, gender, race and post-mortem delay for each of the individual post-mortem brain samples. Supplementary material 3 (DOCX 16 kb)
Rights and permissions
About this article
Cite this article
Graham, S.F., Chevallier, O.P., Kumar, P. et al. High resolution metabolomic analysis of ASD human brain uncovers novel biomarkers of disease. Metabolomics 12, 62 (2016). https://doi.org/10.1007/s11306-016-0986-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11306-016-0986-9