Abstract
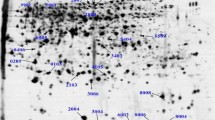
Platycladus orientalis (L.) Franco is widely used for afforestation in arid and semi-arid areas due to its high drought tolerance. To better understand the mechanism involved in drought tolerance in this important tree, responses to drought stress have been studied in 1-year-old P. orientalis via water withholding. Several physiological parameters were evaluated in four drought-treated groups. The root and leaf proteomes of two-dimensional electrophoresis (2-DE) gels were obtained, and a total of 162 proteins with significant quantitative variations and 1.5-fold differences in proteins were selected and identified by matrix-assisted laser desorption/ionization-time-of-flight/time-of-flight (MALDI-TOF/TOF) tandem mass spectrometry (MS/MS). The majority of identified proteins were classified into functional categories, including stress response/defense, carbohydrate metabolic process, nitrogen metabolism, proteolysis, and photosynthesis. The proteome results revealed that a series of strategies were employed to survive in the drought environment, such as the maintenance of protein stability, activation, and folding; reactive oxygen species (ROS) detoxification; the regulation of cell osmotic conditions and cell wall integrity; energy metabolism; and stabilization of the cell skeleton. One of the most prominent findings in this study was the number of bark protein-like proteins and heat shock proteins detected in both root and leaf tissues in drought stress conditions. In addition, 12 differentially expressed proteins were selected for quantitative reverse transcription PCR analysis; not all of the protein expression levels were consistent with the mRNA expression levels. Our data provide a comprehensive picture of the root and leaf responses under varying watering regimes, which could be beneficial for further research and for understanding highly complex drought stress responses.








Similar content being viewed by others
References
Ali GM, Komatsu S (2006) Proteomic analysis of rice leaf sheath during drought stress. J Proteome Res 5:396–403
Alvim FC, Carolino SM, Cascardo JC, Nunes CC, Martinez CA, Otoni WC, Fontes EP (2001) Enhanced accumulation of BiP in transgenic plants confers tolerance to water stress. Plant Physiol 126:1042–1054
Amor Y, Haigler CH, Johnson S, Wainscott M, Delmer DP (1995) A membrane-associated form of sucrose synthase and its potential role in synthesis of cellulose and callose in plants. Proc Natl Acad Sci U S A 92:9353–9357
Anderson CM, Kohorn BD (2001) Inactivation of Arabidopsis SIP1 leads to reduced levels of sugars and drought tolerance. J Plant Physiol 158:1215–1219
Arndt S, Clifford S, Wanek W, Jones H, Popp M (2001) Physiological and morphological adaptations of the fruit tree Ziziphus rotundifolia in response to progressive drought stress. Tree Physiol 21:705–715
Arnon DI (1949) Copper enzymes in isolated chloroplasts. Polyphenoloxidase in Beta vulgaris. Plant Physiol 24:1–15
Ashoub A, Beckhaus T, Berberich T, Karas M, Brüggemann W (2013) Comparative analysis of barley leaf proteome as affected by drought stress. Planta 237:771–781
Babiychuk E, Kushnir S, Belles-Boix E, Van Montagu M, Inzé D (1995) Arabidopsis thaliana NADPH oxidoreductase homologs confer tolerance of yeasts toward the thiol-oxidizing drug diamide. J Biol Chem 270:26224–26231
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Candiano G, Bruschi M, Musante L, Santucci L, Ghiggeri GM, Carnemolla B, Orecchia P, Zardi L, Righetti PG (2004) Blue silver: a very sensitive colloidal Coomassie G-250 staining for proteome analysis. Electrophoresis 25:1327–1333
Carvalho M (2008) Drought stress and reactive oxygen species. Plant Signal Behav 3:156–165
Chang E, Shi S, Liu J, Cheng T, Xue L, Yang X, Yang W, Lan Q, Jiang Z (2012) Selection of reference genes for quantitative gene expression studies in Platycladus orientalis (Cupressaceae) using real-time PCR. PLoS One 7:e33278
Chen S, Harmon AC (2006) Advances in plant proteomics. Proteomics 6:5504–5516
Cho EK, Choi YJ (2009) A nuclear-localized HSP70 confers thermoprotective activity and drought-stress tolerance on plants. Biotechnol Lett 31:597–606
Coleman GD, Banados MP, Chen TH (1994) Poplar bark storage protein and a related wound-induced gene are differentially induced by nitrogen. Plant Physiol 106:211–215
Coleman HD, Ellis DD, Gilbert M, Mansfield SD (2006) Up-regulation of sucrose synthase and UDP-glucose pyrophosphorylase impacts plant growth and metabolism. Plant Biotechnol J 4:87–101
Crane R, Craig R, Murray R, Dunand-Sauthier I, Humphrey T, Norbury C (2000) A fission yeast homolog of Int-6, the mammalian oncoprotein and eIF3 subunit, induces drug resistance when overexpressed. Mol Biol Cell 11:3993–4003
Deeba F, Pandey AK, Ranjan S, Mishra A, Singh R, Sharma Y, Shirke PA, Pandey V (2012) Physiological and proteomic responses of cotton (Gossypium herbaceum L.) to drought stress. Plant Physiol Biochem 53:6–18
Durand TC, Sergeant K, Renaut J, Planchon S, Hoffmann L, Carpin S, Label P, Morabito D, Hausman JF (2011) Poplar under drought: comparison of leaf and cambial proteomic responses. J Proteome 74:1396–1410
Echevarría-Zomeño S, Ariza D, Jorge I, Lenz C, Del Campo A, Jorrín JV, Navarro RM (2009) Changes in the protein profile of Quercus ilex leaves in response to drought stress and recovery. J Plant Physiol 166:233–245
Gazanchian A, Hajheidari M, Sima NK, Salekdeh GH (2007) Proteome response of Elymus elongatum to severe water stress and recovery. J Exp Bot 58:291–300
German MA, Asher I, Petreikov M, Dai N, Schaffer AA, Granot D (2004) Cloning, expression and characterization of LeFRK3, the fourth tomato (Lycopersicon esculentum Mill.) gene encoding fructokinase. Plant Sci 166:285–291
Ghabooli M, Khatabi B, Ahmadi FS, Sepehri M, Mirzaei M, Amirkhani A, Jorrín-Novo JV, Salekdeh GH (2013) Proteomics study reveals the molecular mechanisms underlying water stress tolerance induced by Piriformospora indica in barley. J Proteome 94:289–301
Glaser E, Eriksson A, Sjöling S (1994) Bifunctional role of the bc1 complex in plants mitochondrial bc1 complex catalyses both electron transport and protein processing. FEBS Lett 346:83–87
Hajheidari M, Abdollahian-Noghabi M, Askari H, Heidari M, Sadeghian SY, Ober ES, Hosseini Salekdeh G (2005) Proteome analysis of sugar beet leaves under drought stress. Proteomics 5:950–960
He C, Zhang J, Duan A, Zheng S, Sun H, Fu L (2008) Proteins responding to drought and high-temperature stress in Populus× euramericana cv.‘74/76’. Trees 22:803–813
Horn R, Chudobova I, Hänsel U, Herwartz D, Koskull-Döring P, Schillberg S (2013) Simultaneous treatment with tebuconazole and abscisic acid induces drought and salinity stress tolerance in Arabidopsis thaliana by maintaining key plastid protein levels. J Proteome Res 12:1266–1281
Hu X, Liu R, Li Y, Wang W, Tai F, Xue R, Li C (2010) Heat shock protein 70 regulates the abscisic acid-induced antioxidant response of maize to combined drought and heat stress. Plant Growth Regul 60:225–235
Johnson SM, Lim F-L, Finkler A, Fromm H, Slabas AR, Knight MR (2014) Transcriptomic analysis of Sorghum bicolor responding to combined heat and drought stress. BMC Genomics 15:456
Jorge I, Navarro RM, Lenz C, Ariza D, Jorrín J (2006) Variation in the holm oak leaf proteome at different plant developmental stages, between provenances and in response to drought stress. Proteomics 6:S207–S214
Jungblut PR, Holzhütter HG, Apweiler R, Schlüter H (2008) The speciation of the proteome. Chem Cent J 2:1
Khan MN, Sakata K, Komatsu S (2015) Proteomic analysis of soybean hypocotyl during recovery after flooding stress. J Proteome 121:15–27
Kim SG, Kim ST, Wang Y, Kim SK, Lee CH, Kim KK, Kim JK, Lee SY, Kang KY (2010) Overexpression of rice isoflavone reductase-like gene (OsIRL) confers tolerance to reactive oxygen species. Physiol Plant 138:1–9
Kurepa J, Toh-e A, Smalle JA (2008) 26S proteasome regulatory particle mutants have increased oxidative stress tolerance. Plant J 53:102–114
Lers A, Burd S, Lomaniec E, Droby S, Chalutz E (1998) The expression of a grapefruit gene encoding an isoflavone reductase-like protein is induced in response to UV irradiation. Plant MolBiol 36:847–856
Li H, Wang Z, Zhou X, Cheng Y, Xie Z, Manley JL, Feng Y (2013) Far upstream element-binding protein 1 and RNA secondary structure both mediate second-step splicing repression. Proc Natl Acad Sci U S A 110:E2687–E2695
Luciano P, Geli V (1996) The mitochondrial processing peptidase: function and specificity. Experientia 52:1077–1082
Luo S, Ishida H, Makino A, Mae T (2002) Fe2+-catalyzed site-specific cleavage of the large subunit of ribulose 1, 5-bisphosphate carboxylase close to the active site. J Biol Chem 277:12382–12387
Moschou PN, Delis ID, Paschalidis KA, Roubelakis-Angelakis KA (2008) Transgenic tobacco plants overexpressing polyamine oxidase are not able to cope with oxidative burst generated by abiotic factors. Physiol Plant 133:140–156
Munné-Bosch S, Peñuelas J (2004) Drought-induced oxidative stress in strawberry tree (Arbutus unedo L.) growing in Mediterranean field conditions. Plant Sci 166:1105–1110
Nakaminami K, Matsui A, Nakagami H, Minami A, Nomura Y, Tanaka M, Morosawa T, Ishida J, Takahashi S, Uemura M, Shirasu K, Seki M (2014) Analysis of differential expression patterns of mRNA and protein during cold-acclimation and de-acclimation in Arabidopsis. Mol Cell Proteomics 13:3602–3611
Oland K (1959) Nitrogenous reserves of apple trees. Physiol Plant 12:594–648
O'mahony PJ, Oliver MJ (1999) The involvement of ubiquitin in vegetative desiccation tolerance. Plant Mol. Biol 41:657–667
Pokalsky A, Hiatt W, Ridge N, Rasmussen R, Houck C, Shewmaker C (1989) Structure and expression of elongation factor 1α in tomato. Nucleic Acids Res 17:4661–4673
Ptushkina M, Malys N, McCarthy JE (2004) eIF4E isoform 2 in Schizosaccharomyces pombe is a novel stress-response factor. EMBO Rep 5:311–316
Riccardi F, Gazeau P, de Vienne D, Zivy M (1998) Protein changes in response to progressive water deficit in maize quantitative variation and polypeptide identification. Plant Physiol 117:1253–1263
Schmittgen TD, Livak KJ (2008) Analyzing real-time PCR data by the comparative CT method. Nat Protoc 3:1101–1108
Sergeant K, Spieß N, Renaut J, Wilhelm E, Hausman JF (2011) One dry summer: a leaf proteome study on the response of oak to drought exposure. J Proteome 74:1385–1395
Sung DY, Guy CL (2003) Physiological and molecular assessment of altered expression of Hsc70-1 in Arabidopsis. Evidence for pleiotropic consequences. Plant Physiol 132:979–987
Tezara W, Mitchell V, Driscoll S, Lawlor D (1999) Water stress inhibits plant photosynthesis by decreasing coupling factor and ATP. Nature 401:914–917
Thompson JE, Hopkins MT, Taylor C, Wang T-W (2004) Regulation of senescence by eukaryotic translation initiation factor 5A: implications for plant growth and development. Trends Plant Sci 9:174–179
Tobin AK, Yamaya T (2001) Cellular compartmentation of ammonium assimilation in rice and barley. J Exp Bot 52:591–604
Valdés AE, Irar S, Majada JP, Rodríguez A, Fernández B, Pagès M (2013) Drought tolerance acquisition in Eucalyptus globulus (Labill.): a research on plant morphology, physiology and proteomics. J Proteome 79:263–276
Valero-Galván J, González-Fernández R, Navarro-Cerrillo RM, Gil-Pelegrín E, Jorrín-Novo JV (2013) Physiological and proteomic analyses of drought stress response in Holm oak provenances. J Proteome Res 12:5110–5123
Villar-Salvador P, Planelles R, Oliet J, Peñuelas-Rubira JL, Jacobs DF, González M (2004) Drought tolerance and transplanting performance of holm oak (Quercus ilex) seedlings after drought hardening in the nursery. Tree Physiol 24:1147–1155
Washburn MP, Koller A, Oshiro G, Ulaszek RR, Plouffe D, Deciu C, Winzeler E, Yates JR (2003) Protein pathway and complex clustering of correlated mRNA and protein expression analyses in Saccharomyces cerevisiae. Proc Natl Acad Sci U S A 100:3107–3112
Wetzel S, Demmers C, Greenwood J (1989) Seasonally fluctuating bark proteins are a potential form of nitrogen storage in three temperate hardwoods. Planta 178:275–281
Xiao X, Yang F, Zhang S, Korpelainen H, Li C (2009) Physiological and proteomic responses of two contrasting Populus cathayana populations to drought stress. Physiol Plant 136:150–168
Yan S, Tang Z, Su W, Sun W (2005) Proteomic analysis of salt stress-responsive proteins in rice root. Proteomics 5:235–244
Yates SA, Swain MT, Hegarty MJ, Chernukin I, Lowe M, Allison GG, Ruttink T, Abberton MT, Jenkins G, Skøt L (2014) De novo assembly of red clover transcriptome based on RNA-Seq data provides insight into drought response, gene discovery and marker identification. BMC Genomics 15:453
Yer EN, Baloglu MC, Ziplar UT, Ayan S, Unver T (2016) Drought-responsive Hsp70 Gene analysis in Populus at genome-wide level. Plant Mol Biol Rep 34:483–500
Yoda H, Yamaguchi Y, Sano H (2003) Induction of hypersensitive cell death by hydrogen peroxide produced through polyamine degradation in tobacco plants. Plant Physiol 132:1973–1981
Zadražnik T, Hollung K, Egge-Jacobsen W, Meglič V, Šuštar-Vozlič J (2013) Differential proteomic analysis of drought stress response in leaves of common bean (Phaseolus vulgaris L.) J Proteome 78:254–272
Zhang X, Garreton V, Chua N-H (2005) The AIP2 E3 ligase acts as a novel negative regulator of ABA signaling by promoting ABI3 degradation. Genes Dev 19:1532–1543
Zhang X, Liu S, Takano T (2008) Overexpression of a mitochondrial ATP synthase small subunit gene (AtMtATP6) confers tolerance to several abiotic stresses in Saccharomyces cerevisiae and Arabidopsis thaliana. Biotechnol Lett 30:1289–1294
Zhang S, Chen F, Peng S, Ma W, Korpelainen H, Li C (2010) Comparative physiological, ultrastructural and proteomic analyses reveal sexual differences in the responses of Populus cathayana under drought stress. Proteomics 10:2661–2677
Zhang S, Zhang L, Chai Y, Wang F, Li Y, Su L, Zhao Z (2015) Physiology and proteomics research on the leaves of ancient Platycladus orientalis (L.) during winter. J. Proteomics 126:263–278
Zhang S, Zhang L, Zhao Z, Li Y, Zhou K, Su L, Zhou Q (2016) Root transcriptome sequencing and differentially expressed drought-responsive genes in the Platycladus orientalis (L.) Tree Genet Genomes 12:79
Zhu J-K (2001) Plant salt tolerance. Trends Plant Sci 6:66–71
Zhu J-K (2002) Salt and drought stress signal transduction in plants. Annu Rev Plant Biol 53:247–273
Acknowledgements
This work was financially supported by the National Forestry Industry Research Special Funds for Public Welfare Projects (China) (201404302).
Author information
Authors and Affiliations
Contributions
ZZ developed and supervised the work. The preparation of plant material, 2-DE gel analysis, gene-expression analysis, and data analysis were conducted by SZ and LLZ. KKZ and YML helped in the sample collection and qRT-PCR experiment. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing interests.
Additional information
Communicated by C. Chen
Data archiving statement
The local database search file (raw data from the Illumina sequencing of P. orientalis) of protein identification has been uploaded in NCBI SRA under accession number (SRX1717969~SRX1717974).
Rights and permissions
About this article
Cite this article
Zhang, S., Zhang, L., Zhou, K. et al. Changes in protein profile of Platycladus orientalis (L.) roots and leaves in response to drought stress. Tree Genetics & Genomes 13, 76 (2017). https://doi.org/10.1007/s11295-017-1159-3
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11295-017-1159-3