Abstract
Seedlessness is a highly desirable trait in fresh fruit. Citrus varieties, such as Clementine mandarin and other related species, show parthenocarpic fruit development without seeds due to self-incompatibility. In spite of that, these fruits frequently contain seeds as a result of cross-pollination by insects with compatible pollen from other citrus cultivars grown nearby. To solve this problem using a biotechnological approach we aim at the destruction of ovules and/or seeds by directing the expression of a toxic gene using the promoter of an ovule and/or seed specific gene. With the purpose of isolating this kind of genes we constructed two cDNA libraries from ovules and seeds at different developmental stages of the Clementine mandarin (Citrus clementina cv. Clemenules). A total of 1,014 ESTs from the ovule library and 1,042 ESTs from the seed library were generated, with a novelty percentage of 27% and 36% among the Spanish Citrus Functional Genomic Project (CFGP) ESTs database, respectively. Quantitative PCR analysis confirmed nearly specific expression in ovule and/or seed of two genes, TRANSPARENT TESTA16 (CcTT16) and TRANSPARENT TESTA7 (CcTT7). Expression of these two genes is restricted to early seed development, and is localized in the embryo sac and endothelium. The promoters of those genes may be useful to genetically engineer citrus species to avoid seed formation in fruits of commercial varieties.





Similar content being viewed by others
References
Al-Shahrour F, Diaz-Uriarte R, Dopazo J (2005) Discovering molecular functions significantly related to phenotypes by combining gene expression data and biological information. Bioinformatics 21:2988–2993
Al-Shahrour F, Minguez P, Tarraga J, Montaner D, Alloza E, Vaquerizas JMM, Conde L, Blaschke C, Vera J, Dopazo J (2006) BABELOMICS: a systems biology perspective in the functional annotation of genome-scale experiments. Nucleic Acids Res 34:W472–W476
Alos E, Cercos M, Rodrigo MJ, Zacarias L, Talon M (2006) Regulation of color break in citrus fruits. Changes in pigment profiling and gene expression induced by gibberellins and nitrate, two ripening retardants. J Agric Food Chem 54:4888–4895
Bugos RC, Chiang VL, Zhang XH, Campbell ER, Podila GK, Campbell WH (1995) RNA isolation from plant tissues recalcitrant to extraction in guanidine. Biotechniques 19:734–737
Costantini E, Landi L, Silvestroni O, Pandolfini T, Spena A, Mezzetti B (2007) Auxin synthesis-encoding transgene enhances grape fecundity. Plant Physiol 143:1689–1694
Debeaujon I, Nesi N, Perez P, Devic M, Grandjean O, Caboche M, Lepiniec L (2003) Proanthocyanidin-accumulating cells in Arabidopsis testa: regulation of differentiation and role in seed development. Plant Cell 15:2514–2531
Dorcey E, Urbez C, Blazquez MA, Carbonell J, Perez-Amador MA (2009) Fertilization-dependent auxin response in ovules triggers fruit development through the modulation of gibberellin metabolism in Arabidopsis. Plant J 58:318–332
Debeaujon I, Peeters AJ, Léon-Kloosterziel KM, Koornneef M (2001) The TRANSPARENT TESTA12 gene of Arabidopsis encodes a multidrug secondary transporter-like protein required for flavonoid sequestration in vacuoles of the seed coat endothelium. Plant Cell 13:853–571
Forment J, Gadea J, Huerta L, Abizanda L, Agusti J, Alamar S, Alos E, Andres F, Arribas R, Beltran JP, Berbel A, Blazquez MA, Brumos J, Canas LA, Cercos M, Colmenero-Flores JM, Conesa A, Estables B, Gandia M, Garcia-Martinez JL, Gimeno J, Gisbert A, Gomez G, Gonzalez-Candelas L, Granell A, Guerra J, Lafuente MT, Madueno F, Marcos JF, Marques MC, Martinez F, Martinez-Godoy MA, Miralles S, Moreno P, Navarro L, Pallas V, Perez-Amador MA, Perez-Valle J, Pons C, Rodrigo I, Rodriguez PL, Royo C, Serrano R, Soler G, Tadeo F, Talon M, Terol J, Trenor M, Vaello L, Vicente O, Vidal C, Zacarias L, Conejero V (2005) Development of a citrus genome-wide EST collection and cDNA microarray as resources for genomic studies. Plant Mol Biol 57:375–391
Forment J, Gilabert F, Robles A, Conejero V, Nuez F, Blanca JM (2008) EST2uni: an open, parallel tool for automated EST analysis and database creation, with a data mining web interface and microarray expression data integration. BMC Bioinformatics 9:5
Fos M, Nuez F, Garcia-Martinez JL (2000) The gene pat-2, which induces natural parthenocarpy, alters the gibberellin content in unpollinated tomato ovaries. Plant Physiol 122:471–480
Fos M, Proaño K, Nuez F, Garcia-Martinez JL (2001) Role of gibberellins in parthenocarpic fruit development induced by the genetic system pat-3/pat-4 in tomato. Physiol Plant 111:545–550
Hashimoto JG, Beadles-Bohling AS, Wiren KM (2004) Comparison of RiboGreen and 18S rRNA quantitation for normalizing real-time RT-PCR expression analysis. Biotechniques 36:54–60
Haughn G, Chaudhury A (2005) Genetic analysis of seed coat development in Arabidopsis. Trends Plant Sci 10:472–477
Hartley RW (1988) Barnase and barstar. Expression of its cloned inhibitor permits expression of a cloned ribonuclease. J Mol Biol 202:913–915
Jackson DP (1992) In situ hybridization in plants. In: Bowles DJ, Gurr SJ, Pherenson M (eds) Molecular plant pathology: a practical approach. Oxford University Press, Oxford, UK, pp 163–174
Koltunow AM (1993) Isolation and construction of genes to control seed production in Citrus. In: Hyashi T, Omura M, Scott NS (eds) Techniques on gene diagnosis and breeding. Fruit Tree Research Station, Tsukuba, Japan, pp 101–108
Koltunow AM, Soltys K, Nito N, Mcclure S (1995) Anther, ovule, seed and nucellar embryo development in Citrus sinensis cv Valencia. Can J Bot 73:1567–1582
Koltunow AM, Brennan P, Bond JE, Barker SJ (1998) Evaluation of genes to reduce seed size in Arabidopsis and tobacco and their application to Citrus. Mol Breeding 4:235–251
Le BH, Cheng C, Bui AQ, Wagmaister JA, Henry KF, Pelletier J, Kwong L, Belmonte M, Kirkbride R, Horvath S, Drews GN, Fischer RL, Okamuro JK, Harada JJ, Goldberg RB (2010) Global analysis of gene activity during Arabidopsis seed development and identification of seed-specific transcription factors. Proc Natl Acad Sci USA 107:8063–8070
Li DD, Shi W, Deng XX (2002) Agrobacterium-mediated transformation of embryogenic calluses of Ponkan mandarin and the regeneration of plants containing the chimeric ribonuclease gene. Plant Cell Rep 21:153–156
Marques MC, Alonso-Cantabrana H, Forment J, Arribas R, Alamar S, Conejero V, Perez-Amador MA (2009) A new set of ESTs and cDNA clones from full-length and normalized libraries for gene discovery and functional characterization in citrus. BMC Genomics 10:428
Mezzetti B, Landi L, Pandolfini T, Spena A (2004) The DefH9-iaaM auxin-synthesizing gene increases plant fecundity and fruit production in strawberry and raspberry. BMC Biotechnology 4:4
Nesi N, Debeaujon I, Jond C, Stewart AJ, Jenkins GI, Caboche M, Lepiniec L (2002) The TRANSPARENT TESTA16 locus encodes the ARABIDOPSIS BSISTER MADS domain protein and is required for proper development and pigmentation of the seed coat. Plant Cell 14:2463–2479
Olimpieri I, Siligato F, Caccia R, Mariotti L, Ceccarelli N, Soressi GP, Mazzucato A (2007) Tomato fruit-set driven by pollination or by the parthenocarpic fruit allele are mediated by transcriptionally regulated gibberellin biosynthesis. Planta 226:877–888
Peer AW, Brown DE, Tague BW, Muday GK, Taiz L, Murphy AS (2001) Flavonoid accumulation patterns of transparent testa mutants of Arabidopsis. Plant Physiol 126:536–548
Rommens CM, Haring MA, Swords K, Davies HV, Belknap WR (2007) The intragenic approach as a new extension to traditional plant breeding. Trends Plant Sci 12:397–403
Rotino GL, Acciarri N, Sabatini E, Mennella G, Scalzo RL, Maestrelli A, Molesini B, Pandolfini T, Scalzo J, Mezzetti B, Spena A (2005) Open field trial of genetically modified parthenocarpic tomato: seedlessness and fruit quality. BMC Biotechnol 5:32
Schoenbohm C, Martens S, Eder C, Forkmann G, Weisshaar B (2000) Identification of the Arabidopsis thaliana flavonoid 3′-hydroxylase gene and functional expression of the encoded P450 enzyme. Biol Chem 381:749–753
Schwabe WW, Mills JJ (1981) Hormones and parthenocarpic fruit set: a literature survey. Hort Abstr 51:661–698
Singh DP, Filardo FF, Storey R, Jermakow AM, Yamaguchi S, Swain SM (2010) Overexpression of a gibberellin inactivation gene alters seed development, KNOX gene expression, and plant development in Arabidopsis. Physiol Plant 138:74–90
Talon M, Zacarias L, Primo-Millo E (1992) Gibberellins and parthenocarpic ability in developing ovaries of seedless mandarins. Plant Physiol 99:1575–1581
Zimmermann P, Hirsch-Hoffmann M, Hennig L, Gruissem W (2004) GENEVESTIGATOR: Arabidopsis microarray database and analysis toolbox. Plant Physiol 136:2621–2632
Zhulidov PA, Bogdanova EA, Shcheglov AS, Vagner LL, Khaspekov GL, Kozhemyako VB, Matz MV, Meleshkevitch E, Moroz LL, Lukyanov SA, Shagin DA (2004) Simple cDNA normalization using Kamchatka crab duplex-specific nuclease. Nucleic Acids Res 32:e37
Acknowledgements
The authors thank Dr. L. Navarro and J. Juarez, from the Instituto Valenciano de Investigaciones Agrarias (Generalitat Valenciana), for the use of the Clementine orchard and Fortune pollen; Dr. J Forment and the Genomics Facility at the IBMCP for sequence analysis and maintenance of to the CFGP database; and Drs. F Tadeo and J Carbonell for critical reading of the manuscript. This work was supported by grants from the Conselleria de Agricultura, Pesca y Alimentacion (Generalitat Valenciana) and Spanish Ministerio de Ciencia y Tecnologia (research grant GEN2001-4885-C05). A.G-L. received a PhD fellowship from the Conselleria de Agricultura, Pesca y Alimentacion.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by F. GMITTER
Electronic Supplementary Materials
Below is the link to the electronic supplementary material.
Fig. S1
Functional categorization of genes isolated from the ovule (758 unigenes, 684 with Arabidopsis ortholog) and seed (796 unigenes, 707 with Arabidopsis ortholog) libraries compared with the CFGP database (34,895 unigenes, 24,780 with Arabidopsis ortholog). Annotations were classified based on the GO Biological Process categories. The percentage rates of genes in each GO are shown (PDF 21 kb)
Fig. S2
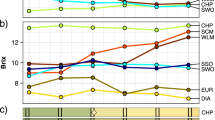
In silico Meta-Profile analysis of ovule gene expression using Geneinvestigator. Reproductive tissues and seed are highlighted by horizontal red rectangles. Five genes were identified (asterisks). The Arabidopsis ortholog gene to the citrus C37005H06 CcTT16 (At5g23260, microarray probe 249851_at) is highlighted by a vertical green rectangle. Orthologs for the other four selected genes (Fig. S4) are highlighted by a grey rectangle: At1g67820, probe 245194_at; At4g21190, probe 254427_at; At5g10510, probe 250426_at; and At4g30850, probe 253590_at) (PDF 208 kb)
Fig. S3
In silico Meta-Profile analysis of seed expression using Geneinvestigator. Reproductive tissues and seed are highlighted by horizontal red rectangles. Five genes were identified (asterisks). The Arabidopsis ortholog gene to the citrus C36004D04 CcTT7 (At3g49240, microarray probe 252305_at) is highlighted by a vertical green rectangle. Orthologs for the other four selected genes (Fig. S4) are highlighted by a grey rectangle: At5g14050, probe 250222_at; At5g66540, probe 247046_at; At3g14690, probe 258094_at; and At5g07990, probe 250558_at) (PDF 299 kb)
Fig. S4
Expression levels of seed (C36) and ovule (C37) genes selected by the in silico meta-profile analysis using Geneinvestigator in different tissues of C. clementine. L leaf, R root, I inflorescence, FB flower bud, P petiole, FP fruit pulp, S seed. Three independent samples were assayed with similar results. Mean ± SD of three technical replica are shown (PDF 21 kb)
Fig. S5
Sequence alignment of C. clementine C36004D04 CcTT7 with its orthologs Gossypium hirsutum (AAP31058.1), Populus trichocarpa (XP_002314004.1), Cyclamen persicum (ACX37698.1), Vitis vinífera (BAE47007.1), Vinca major (ACZ63205.1), Catharanthus roseus (CAA09850.1), Camellia sinensis (AAY23287.1), Rhododendron x pulchrum (BAH98132.1), Viola x wittrockiana (BAF93855.1), and Arabidopsis thaliana AtTT7 (NP_196416.1). Alignment was carried out with the Clustalw2 software at EMBL-EBI (http://www.ebi.ac.uk/Tools/clustalw2/index.html) (PDF 46 kb)
Fig. S6
Sequence alignment of C. clementine C37005H06 CcTT16 and its orthologs from Populus trichocarpa (XP_002318890.1), Arabidopsis thaliana AtTT16 (NP_974823.1), Brassica napus (ABY59774.1), Ricinus communis (XP_002513703.1), Vitis vinifera (XP_002276139.1), Antirrhinum majus DEFH21 (Q8RVL4.1), Zea mays ZMM17 (NP_001105130.1), Sorghum bicolor (XP_002453370.1), Oryza sativa (AAY46447.1), Gnetum gnemon GGM13 (Q9XGJ4.1), Petunia hibrida FBP24 (Q9ATE5.1), and Ginkgo biloba (BAD93174.1). Alignment was carried out with the Clustalw2 software at EMBL-EBI (http://www.ebi.ac.uk/Tools/clustalw2/index.html). Relevant motifs and domains are indicated (PDF 41 kb)
Fig. S7
Phylogenetic relationships of C36004D04 CcTT7 and related flavonoid 3′,5′-hydroxylase proteins (a) and C37005H06 CcTT16 and related MADS domain proteins (b). The consensus most parsimonious trees are shown (PDF 19 kb)
Table S1
(DOC 31.0 kb)
Table S2
(XLS 53.5 kb)
Table S3
(DOC 32.5 kb)
Rights and permissions
About this article
Cite this article
Garcia-Lor, A., Garcia-Martinez, J.L. & Perez-Amador, M.A. Identification of ovule and seed genes from Citrus clementina . Tree Genetics & Genomes 8, 227–235 (2012). https://doi.org/10.1007/s11295-011-0435-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11295-011-0435-x