Abstract
Glycopeptide antibiotics (GPAs) are a family of non-ribosomal peptide natural products with polypeptide skeleton characteristics, which are considered the last resort for treating severe infections caused by multidrug-resistant Gram-positive pathogens. Over the past few years, an increasing prevalence of Gram-positive resistant strain “superbugs” has emerged. Therefore, more efforts are needed to study and modify the GPAs to overcome the challenge of superbugs. In this mini-review, we provide an overview of the complex biosynthetic gene clusters (BGCs), the ingenious crosslinking and tailoring modifications, the new GPA derivatives, the discoveries of new natural GPAs, and the new applications of GPAs in antivirus and anti-Gram-negative bacteria. With the development and interdisciplinary integration of synthetic biology, next-generation sequencing (NGS), and artificial intelligence (AI), more GPAs with new chemical structures and action mechanisms will constantly be emerging.
Graphical abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
GPAs are a family of natural and semisynthetic glycosylated non-ribosomal peptides that comprise tricyclic or tetracyclic polypeptide scaffold and display antibacterial activity against Gram-positive organisms through binding to the C-terminal D-alanyl-D-alanine of the lipid II bacterial cell-wall precursor via H-bonds, preventing crosslinking of the peptidoglycan and ultimately inhibit the synthesis of cell wall (Butler et al. 2014; Vimberg et al. 2019). However, recent research discovered that GPA oritavancin inhibited bacterial activity by jointly blocking the biosynthesis of wall teichoic acid and peptidoglycan (Singh et al. 2017). In addition, the new actional mechanism of the type V GPAs complestatin, GP6738, and corbomycin was discovered, which presented that these antibiotics could inhibit the action of autolysins by binding to peptidoglycan and overcome the D-Ala-D-Lac GPA resistance (Culp et al. 2020). Therefore, these discoveries of new action mechanisms provide the possibility for the development of new GPAs.
Generally, the antibacterial spectrum of GPAs excludes Gram-negative species due to their physiochemical properties precluding transit through the outer membrane and blocking target to the lipid II in Gram-negative pathogens (Schaenzer and Wright 2020). In fact, it had been demonstrated that the vancomycin analogues with the characteristics of lipophilic and cationic modification could strongly destroy bacterial membranes and thereby overcome the inherent resistance of Gram-negative pathogens (Yarlagadda et al. 2016). Research showed that the cold stress cultivation made Gram-negative Escherichia coli susceptible to glycopeptide antibiotics by altering outer membrane integrity (Stokes et al. 2016). In addition, many studies had confirmed the inhibitory effect of glycopeptide antibiotics on viruses, including Ebola, MERS-Cov, and SARS-Cov virus (Acharya et al. 2022; Zhou et al. 2016).
GPAs are the last line of defense against Gram-positive drug-resistant bacteria. However, with the clinical application of first- and second-generation GPAs, drug-resistant bacteria have emerged. Antibiotic resistance continues to be a severe threat to modern medicine. Resistance to antibiotics occurs through a variety of molecular strategies. In particular, resistance to glycopeptide antibiotics manifests through the activity of enzymes such as dipeptidases VanH, VanA, and VanX that chemically alter elements of the cell envelope required for antibiotic binding (Frasch et al. 2015; Schaenzer and Wright 2020). In addition, resistance to GPAs can also be performed through intermediate strategies which are found in Vancomycin-intermediate S. aureus (VISA) (Hu et al. 2016).
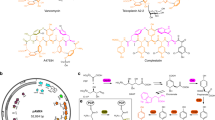
GPAs are divided into five distinct structural subclasses (I–V) according to the substituents and the type of residues at positions 1 and 3 of the polypeptide (Butler et al. 2014) (Fig. 1). Type I GPAs, represented by vancomycin, contain three side-chain crosslinked rings, including an A-B ring, a C-O-D ring, and a D-O-E ring. Type II GPAs (exemplified by avoparcin) possess the same crosslinked structure as Type I GPAs, unlike Type I, AA1 and AA3 residues of Type II GPAs are modified by aromatic amino acids rather than aliphatic groups. Type III and IV GPAs include an additional crosslinked F-O-G ring between aromatic groups at AA1 and AA3 residues compared to Type I/II GPAs. The difference between Type III and IV GPAs is that there is an acyl chain substitute in the structural scaffold of Type IV, but there isn’t in Type III. Type V GPAs contain a typical tryptophan (Trp) moiety linked to the central residue Hpg to form a conserved Trp-Hpg-(m)Tyr motif and are not generally glycosylated. The core scaffold of GPAs, known as the aglycone, requires the essential crosslinked rings, including A-B, C-O-D, and D-O-E, which are necessary for GPAs to present their antibacterial activity (Fig. 1; Table 1).
Schematic representation of GPAs grouped into five (I − V) structural subtypes. The core structural scaffold of GPAs (I–V) is marked in red. The typical chemical structure of different type GPA is presented using red core scaffold plus specific tailoring moieties marked in the enclosing ring. Type I: red scaffold plus black circle; Type II: red scaffold plus blue ellipse; Type III: red scaffold plus pink dashed ellipse; Type IV: red scaffold plus pink dashed ellipse and green dashed ellipse; Type V: red scaffold plus cyan ellipse
Diversity and conservation of GPA BGCs
Generally, GPA BGCs are modularly composed of the genes required for the amino acid precursor supply, peptide assembly, and scaffold crosslinking by non-ribosomal peptide synthetases (NRPSs) and cytochrome P450s, tailoring enzymes (halogenation, sulfation, glycosylation, methylation, and acylation), transport, regulation, and resistance. Recently, Wright’s group revealed that the abilities of glycopeptide biosynthesis and resistance in Actinobacteria emerged approximately 150–400 million years ago by phylogenetic analysis based on the sources of 71 GPA BGCs. Especially noted in this research, the precursors of GPA biosynthesis are much older than other components (Waglechner et al. 2019).
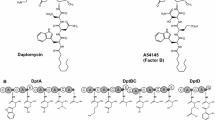
NRPSs are a class of ribosome-independent large modular enzymes involved in the biosynthesis of many different peptide-derived natural products (Fig. 2). A typical linear NRPS structure consists of multiple modules, and each module on the assembly line is responsible for transferring and inserting an amino acid monomer into the final peptide. Typically, these modules include adenylation (A)-domain (responsible for amino acid selection and activation), condensation (C)-domain (responsible for peptide bond formation), thioesterase (TE)-domain (responsible for cleaving the mature peptide from the NRPS), epimerization (E)-domain (responsible for the altering the stereochemistry of peptide residues), methyltransferase (MT)-domain (responsible for methylation of amino acid), and P450 recruitment (X)-domain (responsible for interaction with P450 oxygenases). Except above module, peptidyl carrier proteins (PCPs) act as the attachment point for all biosynthetic intermediates during peptide assembly (Izore and Cryle 2018). In NRPS modules, the specific combination of A domain, C domain, and E domain determines the corresponding peptide sequence, which can be as a vital machinery to guide the development of new GPAs.
A domains, requiring ATP activation, are composed of the larger N-terminal subdomain and the smaller C-terminal subdomain and can recognize and activate more than 500 different substrates for peptide synthesis (Walsh et al. 2013). Previous studies had demonstrated that redesigning the A domain could extend a range of non-natural monomers and produce different novel peptides (Weist et al. 2004). C domains are responsible for peptide bond formation and the stereochemical selectivity of the substrates with the L- or D-configuration, indicating a tight relationship between the C domains and E domains (Chen et al. 2016). Generally, A domains mainly select and activate L-configured monomers, requiring E domains to epimerize the substrate from the L to the D form. TE domains in NRPS modular are typically located at the end of the NRPS assembly line, which is responsible for the cleavage of thioester products and for the peptide cyclization (Owen et al. 2016). The conserved X domains without catalytical activity are located in the final module of all GPA NRPS lines. They are responsible for recruiting P450s to the NRPS-bound peptide to perform the crosslink of linear peptide (Haslinger et al. 2015). Additionally, a previous study revealed that the P450 recruitment abided by a continuous association/dissociation model between the P450 enzymes and the NRPS, namely the P450 scanning (Peschke et al. 2016).
Ingenious biosynthetic cascades of GPAs
Preparation of amino acids and assembly of linear polypeptide
Within the biosynthesis of GPAs, the amino acids as monomers include non-proteinogenic amino acids and common amino acids. In general, non-proteinogenic amino acids are derived from the classic 20 proteinogenic amino acids, but there are some special cases, which are produced de novo according to the corresponding modules in BGCs of GPAs.
The common amino acid precursors in the GPA family mainly include Asn, Leu, Ala, Phe, Glu, Trp, and Tyr with different epitopic configurations. The other non-proteinogenic amino acids are hydroxyphenylglycine (Hpg), 3,5-dihydroxyphenylglycine (Dpg), and β-hydroxytyrosine (Bht) (Fig. 2). The three amino acids share common aromatic amino acid biosynthetic pathways containing the crucial enzymes 3-deoxy-D-arabinoheptulosonate-7-phosphate synthase (DAHPS), chorismate mutase (CM), and prephenate dehydrogenase (PDH) (Waglechner et al. 2019).
During the NRPS-mediated biosynthesis of GPAs, A domains, C domains, and Type-II TE domains can act collaboratively to ensure the incorporation of the correct amino acid to the polypeptide line, even if the A domain displays weak specificities for substrate selection. Furthermore, the amino acid modification in trans can be selectively controlled during non-ribosomal peptide biosynthesis through the exchange of selective C-domains (Kaniusaite et al. 2019). In addition, the C domain has a broad tolerance to changes to the amino acids. It can select both L- and D-configured peptides and significantly prefer linear peptides over crosslinked ones. Although the C domain is regarded as a stereochemical gatekeeper during GPA synthesis, its selectivity depends on the presence or absence of the adjacent E domain. Furthermore, the peptide bond formation as an essential role for the C domain is tightly related to the rate of monomer activation exerted by the neighboring A domain (Schoppet et al. 2019).
Crosslinking of GPAs guided by P450s
Oxidative crosslinking of aromatic side chains during the biosynthesis of GPAs is a vital modification depending on the catalytic action of cytochrome P450 enzymes (P450s). P450s are a superfamily of monooxygenases given their diverse oxidative reactions and abundant catalytic substrates by a cysteine thiolate-ligated heme iron moiety which is crucial for their ability to generate extremely powerful oxidization. During the biosynthesis of GPAs, different P450s generally catalyze the respective oxidative crosslinking of aromatic side chains. Those known as “Oxy enzymes” play a crucial role in the biosynthesis of GPAs (Haslinger and Cryle 2016).
During the scaffold crosslinking of GPAs, the P450s are recruited to the PCP-bound peptide substrate by binding the same site on the X domain surface. Therefore, these cytochrome oxygenases should compete the binding site of X domain. The related research had verified that the P450s enzymes performed the reactions as the scanning model depending on the experimental evidence of the continuous association/dissociation of P450s with the peptide substrates (Peschke et al. 2016). In addition, the P450 proteins have many Tyr residues to maintain the typical heme orientation and avoid potential oxidative damage based on crystallographic experiments (Greule et al. 2022).
In the first step of substrate selection in the crosslinking oxidation, different P450s present distinct binding abilities to the stereochemical substrate. For example, OxyBtei has a higher tolerance to the seventh amino acid’s stereochemical change than OxyAtei. During the complicated crosslinking process, there exist one or more intermediates of the P450-substrate complex that perform a continuous cycle of combination and dissociation with the NRPS. In vivo and in vitro, previous studies presented that the conserved P450s conducted the oxygenation in a specific order. Generally, the P450s OxyA, OxyB, OxyC, and OxyE act sequentially on the polypeptide precursor in OxyB-(OxyE)-OxyA-OxyC to produce the GPAs (Forneris and Seyedsayamdost 2018; Peschke et al. 2016; Tailhades et al. 2020; Zhao et al. 2020). In another case of teicoplanin biosynthesis, the TE domain located in the last module of Tcp12 is responsible for acting as a logic gate to ensure the formation of mature linear peptide during NRPS biosynthesis. Furthermore, once the Oxy cascade is completed, the finished aglycone is rapidly hydrolyzed to be released (Peschke et al. 2018).
Here, we summarized the classification and protein structure of P450s involved in the oxidational biosynthesis of GPAs (Fig. 3 and Table 2). In general, the D-O-E ring is catalyzed by the OxyA and its homologous enzymes; the OxyB enzymes oxidize the C-O-D ring; the A-(O)-B ring is formed by the OxyC and its homologous counterparts; the F-O-G ring is catalyzed by the OxyE and its homologous enzymes. In particular, oxygenase OxyAkis conducts the formation of a D-E ring in the biosynthesis of kistamicin belonging to type V GPAs. As well, the P450 OxyCkis oxidizes the A-O-B ring (Fig. 3). These obvious differences in P450s indicate that the Type V GPAs are atypical and structurally diverse. In another special case, the enzyme OxyD involved in vancomycin biosynthesis performs hydroxylation (Fig. 3). Additionally, the catalytic activity of the OxyCkis enzyme is flexible and can install two crosslinks of A-O-B ring and C-O-D ring in GPA kistamicin (Greule et al. 2019). Furthermore, the Oxy enzymes have broad tolerance for modification to the N-terminal peptide of GPAs and result in generating diverse derivatives. For instance, a method of click chemistry was used to diversify the cyclic peptide based on the wide tolerance of the enzyme OxyB/A for substituted amino acids at position 3 of GPA (Tailhades et al. 2020).
P450 oxygenases involved in the crosslinking scaffold formation during GPAs biosynthesis. The crystallographic structures of some P450s loaded in the Protein Data Bank (PDB) database are showed. NA means no available of crystallographic information from PDB. kis kistamicin, Oxy vancomycin, Sta A47934, tei teicoplanin, Cep chloroeremomycin, bal balhimycin, Dbv A40926
Glycosylation of GPAs
Glycosylation is one of the essential modifications which are performed by glycosyltransferases (GTs) found in different BGCs of GPAs. Generally, glycosylation and acylation are necessary for the antimicrobial activities of some GPAs, such as teicoplanin and A40926, belonging to Type IV GPAs. Furthermore, it should be noted that all types of GPAs are modified by glycosylation except for Type V and Type III GPA A47934 (Fig. 4) (Pootoolal et al. 2002; Xu et al. 2022).
The monosaccharide moieties and their corresponding glycosyltransferases of GPAs. Solid ellipses contain the different sugars; dashed ellipses encircle glycosyltransferases. B Balhimycin, V Vancomycin, Dbv A40926, K Keratinimicin A, C Chloroeremomycin, Orf Ristocetin A, Pek Pekiskomycin, Tei Teicoplanin, Auk UK-68,597
As for sugar donors, their resources are from primary metabolism or non-conventional metabolism. In detail, the primary metabolite sugar, namely conventional sugars, are D-glucose, D-mannose, D-arabinose, N-acetylglucosamine (GlcNAc), and L-rhamnose. For example, Type IV GPAs, including teicoplanin and A40926, are decorated with sugars deriving from primary metabolites such as α-D-mannose and GlcNAc. Non-conventional sugar residues include L-vancosamine, L-epivancosamine, L-4-oxovancosamine, L-ristosamine, and L-actinosamine. For instance, Type I GPAs vancomycin, balhimycin, and chloroeremonycin have sugar moiety of non-conventional monosaccharides such as L-vancosamine, L-4-oxovancosamine, and L-epivancosamine. These non-conventional monosaccharides are biosynthesized by enzymes from the BGCs of GPAs. Furthermore, all GTs involved in the glycosylation belong to two families GT1 and GT39, according to the Carbohydrate-Active enZYmes Databse (CAZy, http://www.cazy.org) (Drula et al. 2022). The GT1 family is responsible for transferring non-conventional aminosugars and conventional D-glucose, D-arabinose, GlcNAc, and L-rhamnose, and requires the dTDP- or UDP-activated sugar substrates. These enzymic proteins characterize a unique two-domain scaffold, including C domains as the recognition sites for the donor NDP-activated sugars and N domains as the acceptor binding sites for polypeptide aglycones (Zhang et al. 2020). As for the second GT family, the installation of conventional sugar D-mannose is the only case performed by the GT39 family, which requires D-mannosyl-1-phosphoundecaprenol as a donor substrate.
Generally, the GT1 enzymes prefer the glycosylation sites at aa-4 (Hpg) and aa-6 (Bht) of the aglycone or add additional monosaccharides to already existing mono/di/trisaccharides at aa-4. The other family GT39 is responsible for attaching mannose at aa-7 (Dpg).
Halogenation of GPAs
Halogenation is also an important tailoring modification for the stability and activity of natural GPAs, in which chlorination is the most common halogenation modification. For example, dechlorinated vancomycin showed a decrease in antibacterial activity (Wadzinski et al. 2016). Similarly, the halogenation of A47934 is not only necessary for antimicrobial activity but also for avoiding the induction of resistance mechanisms in some pathogens (Yim et al. 2018). In contrast, the antimicrobial activity of dechlorinated A40926 is similar to that of A40926 (Beltrametti et al. 2003).
So far, the known GPA BGCs contain up to two halogenases, and can lead to six chlorinations at most in the known GPA structures. Generally, one halogenase gene is responsible for one to two chlorinations. If there exist two halogenases in the BGCs, three or four chlorinations will be installed in the GPA structure. As an exception, StaI encoding only one halogenase in the sta cluster of A47934 presents three halogenations at different moieties of amino acids on the A47934 scaffold (Yim et al. 2018). In another specific case, atypical GPA complestatin contains six chlorine atoms derived from three different amino acid monomers of 1,3-dichloro-Hpg, 3,5-dichloro-Hpg, and 3,5-dichloro-hydroxybenzoylformate catalyzed by nonheme metal-free halogenase ComH (Chiu et al. 2001).
During the halogenation, halogenase recognizes the amino acid substrate loaded to the corresponding PCP domain, which is necessary and contributes some role in the procedure. Additionally, the halogenation of GPAs can improve oxidative processing by assisting the P450s to maintain higher activity. This reveals that GPA chlorination occurs before the oxidative crosslinking cascade (Peschke et al. 2017).
Acylation of GPAs
Compared with Types I-III GPAs, Type IV GPAs include an emblematic modification of N-lipidated glucosamine on the residue aa4. This particular feature inspired the development of novel semisynthetic GPA analogues such as oritavancin and telavancin. Therefore, these GPAs featured a long aliphatic acyl side chain and are also named lipoglycopeptide antibiotics. This type of antibiotic is powered by the N-acyltransferases (NAT), which introduce the N-acylation on glucosamine at the central residue of pseudoglycone. The X-ray crystallographic analysis of Orf11 protein (N-acyltransferase from tei cluster) demonstrates that it can dock the bulky or lengthy acyl moiety and thus allows diversity in novel derivatives. Based on the above characteristics of NAT, many teicoplanin analogues with different acyl modifications are developed and show excellent antibacterial activities (Lyu et al. 2014).
New development of GPAs analogues
Over the past 50 years, GPAs have been regarded as the last resort to cure serious Gram-positive infections. However, drug resistance has always been a thorny problem accompanying the clinical application of GPAs. The emergence of glycopeptide resistance has resulted in developing the synthetic and semisynthetic glycopeptide analogues to target acquired resistance. Recently, many methodologies, including chemistry, biochemistry, enzymology, microbiology, and the mixing method based on the above methods, have been applied to develop new GPAs with higher activity and more broad antibacterial spectra. Here, we summed up the progress of some GPA analogues as described below.
Vancomycin analogues
Although more novel semisynthetic GPAs such as telavancin, dalbavancin, and oritavancin have been used in clinic, vancomycin, a representation of the first-generation GPA, has been a mainstream antibiotic against serious Gram-positive infections. Over the past few years, many analogues of vancomycin have been developed via different modification strategies, including C-terminal modification (membrane targeting approaches) (Blaskovich et al. 2018), core scaffold modifications (Okano et al. 2017), aryl ring functionalization (Guan et al. 2018), hydroxyl modifications (Han and Miller 2013), N-terminal modifications(Chang et al. 2013), glycopeptide dimers (Silverman et al. 2017) and conjugates (Varisco et al. 2014). For example, a series of compounds named vancapticins 7 is developed by modifying the C-terminus of vancomycin with a modular assembly containing a bis-amine linker moiety, a cationic peptide, and a hydrophobic cap. These analogues are designed to target bacterial membranes with the aid of hydrophobic caps and cationic peptides. The results of antibacterial tests show that the combined modifications present an over 100-fold increase in potency against MRSA (Blaskovich et al. 2018). Similarly, the other studies reported that the modification of vancomycin C-terminus including the zinc-binding dipicolyl moiety, the benzoxaborole group, the quaternary ammonium propylamine, and positively-charged amino acids (such as lysine and arginine) contribute to the increase of antibacterial activity and the extend of antimicrobial spectra (Antonoplis et al. 2018; Antonoplis et al. 2019).
Teicoplanin analogues
Teicoplanin, which belongs to the second-generation and Type IV GPA, has a typical long-chain N-acyl moiety, resulting in teicoplanin being a mixture of five related derivatives and cannot form non-covalent dimers because of the apolar tag. In order to develop new teicoplanin analogues, teicoplanin generally is treated to remove the one or two β-D-GlcNAc substituents and further form new analogues by lipidation, N-terminal guanidalization, and dimerization. These new teicoplanin analogues show high activity against VanA teicoplanin-resistant enterococci (Bereczki et al. 2022a; Szucs et al. 2019, 2020). In addition, the latest research reported that teicoplanin derivatives bearing hydrophobic or super-basic side chain showed activity against severe acute respiratory syndrome coronavirus (SARS-CoV-2) in human Calu-3 cells and HCoV-229E in HEL cells (Bereczki et al. 2022b).
A40926 analogues
A40926 is the precursor of the third-generation GPA dalbavancin and is biosynthesized by actinobacteria ATCC 39,727. A previous study reported that a series of alkylated derivatives were synthesized on the deacyl A40926 scaffold. However, the derivatives harboring only the lipophilic mono- or dialkylation of the amino groups showed poor antibacterial activity. Interestingly, further modification of the two carboxylic acids contributed to the increase in antibiotic activity (Maffioli et al. 2005). In addition, a research had reported that a double mutant lacking dbv8 and dbv23 was constructed to produce A40926 intermediates in order to obtain the single A40926 component by chemical synthesis method in vitro (Alt et al. 2019). Recently, a novel GPA A50926 was discovered from Nonomuraea coxensis DSM45129 and shared a similar structure with A40926, lacking the carboxyl group on the N-acylglucosamine moiety (Yushchuk et al. 2021). The semisynthetic antibiotic dalbavancin as the derivative of A40926 is proven to has a high affinity to human angiotensin-concerting enzyme 2 (ACE2), which is the docking site of the SARS-COV-2 spike protein, indicating that dalbavancin is a promising anti-COVID-19 drug candidate (Wang et al. 2021).
Discovery of novel natural polypeptide
The NRPS-encoded polypeptides, including the known GPAs, have always been an appealing source of novel antibiotics. The genome sequencing technology dramatically improves the discovery of NRPS BGCs encoding potential natural lipopeptides from bacteria, especially actinomycetes. Based on the bioinformatic analysis, a new lipopeptide cilagicin containing ten amino acid residues encoded by the cil cluster has been discovered recently. This novel antibiotic apparently resists antibiotic-resistant pathogens (Wang et al. 2022a). Furthermore, macolacin, an analogue of colistin, is chemically synthesized depending on bioinformatic analysis of identified BGC from sequenced bacterial genomes. The colistin congener is effective against pathogens that express the mcr-1 (phosphoethanolamine transferase) resistance gene (Wang et al. 2022b). As the above strategy, a similar route for discovering and verifying new GPAs is developed and described as the GPA Heterologous expression (GPAHex) platform, which is a synthetic biology tool to exploit the cryptic non-expressed GPA under laboratory conditions. With the aid of GPAHex platform, the novel Type V GPAs corbomycin and GP6738 are biosynthesized successfully and effectively (Xu et al. 2020). Furthermore, a series of new Type V GPAs rimomycin (A/B/C) and misaugamycin (A/B) are discovered by combining phylogeny-guided genome mining and the synthetic biology platform GPAHex. These new Type V GPAs have novel chemical scaffolds that expand this type of GPAs’ diversity. In detail, rimomycin shares a similar chemical structure with kistamicin, but presents an N-methyl-tyrosine group at aa6 residue and two Hpg moieties at aa1 and aa3 residues, and while misaugamycin contains an unprecedented N-C ring between aa2 and aa4 and sole N-terminal acylation (Xu et al. 2022).
Perspectives
Few compound classes play dramatically essential roles in human health protection as antibiotics. As typical representatives, GPAs are a group of complicated molecules used to treat serious Gram-positive bacterial infections, which mainly are obtained by the sole pathway of actinomycetes fermentation directly or indirectly. GPA researches continue to generate new discoveries and new entity molecules for antibiotic development. For instance, the reassembly of the NRPS line is performed to generate novel potential GPAs based on synthetic biology. A recent study presented that a total of thirteen tailoring enzymes from seven GPA BGCs in different combinations were introduced into the model actinomycete S. coelicolor, which had priorly loaded a heterogenous minimal scaffold of teicoplanin, which tried to explore the criteria for GPA modification in vivo and to expand GPA chemical diversity (Yim et al. 2016). For another example, a team from Princeton University developed a universal high-throughput platform to screen the secondary metabolome based on an imaging mass spectrometric technology. They found a new glycopeptide substance with potent inhibitory activity against a virus (Xu et al. 2019).
In a word, we believe that with the development and interdisciplinary integration of synthetic biology, NGS, and AI, more GPAs with new chemical structures and action mechanisms will constantly be emerging.
References
Acar J, Casewell M, Freeman J, Friis C, Goossens H (2000) Avoparcin and virginiamycin as animal growth promoters: a plea for science in decision-making. Clin Microbiol Infect 6:477–482. https://doi.org/10.1046/j.1469-0691.2000.00128.x
Acharya Y, Bhattacharyya S, Dhanda G, Haldar J (2022) Emerging roles of glycopeptide antibiotics: moving beyond Gram-positive bacteria. ACS Infect Dis 8:1–28. https://doi.org/10.1021/acsinfecdis.1c00367
Alt S, Bernasconi A, Sosio M, Brunati C, Donadio S, Maffioli SI (2019) Toward single-peak dalbavancin analogs through biology and chemistry. Acs Chem Biol 14:356–360. https://doi.org/10.1021/acschembio.9b00050
Antonoplis A, Zang X, Huttner MA, Chong KKL, Lee YB, Co JY, Amieva MR, Kline KA, Wender PA, Cegelski L (2018) A dual-function antibiotic-transporter conjugate exhibits superior activity in sterilizing MRSA biofilms and killing persister cells. J Am Chem Soc 140:16140–16151. https://doi.org/10.1021/jacs.8b08711
Antonoplis A, Zang X, Wegner T, Wender PA, Cegelski L (2019) Vancomycin-arginine conjugate inhibits growth of carbapenem-resistant E. coli and targets cell-wall synthesis. Acs Chem Biol 14:2065–2070. https://doi.org/10.1021/acschembio.9b00565
Beltrametti F, Lazzarini A, Brunati C, Marazzi A, Jovetic S, Selva E, Marinelli F (2003) Production and characterization of monochlorinated and dechlorinated A40926 derivatives. J Antibiot (Tokyo) 56:773–782. https://doi.org/10.7164/antibiotics.56.773
Bereczki I, Szűcs Z, Batta G, Nagy TM, Ostorházi E, Kövér KE, Borbás A, Herczegh P (2022a) The first dimeric derivatives of the glycopeptide antibiotic teicoplanin. pharmaceuticals (Basel) 15(1):77–92. https://doi.org/10.3390/ph15010077
Bereczki I, Vimberg V, Lőrincz E, Papp H, Nagy L, Kéki S, Batta G, Mitrović A et al (2022b) Semisynthetic teicoplanin derivatives with dual antimicrobial activity against SARS-CoV-2 and multiresistant bacteria. Sci Rep 12:16001–16015. https://doi.org/10.1038/s41598-022-20182-y
Blaskovich MAT, Hansford KA, Gong Y, Butler MS, Muldoon C, Huang JX, Ramu S, Silva AB et al (2018) Protein-inspired antibiotics active against vancomycin- and daptomycin-resistant bacteria. Nat Commun 9:22–38. https://doi.org/10.1038/s41467-017-02123-w
Butler MS, Hansford KA, Blaskovich MA, Halai R, Cooper MA (2014) Glycopeptide antibiotics: back to the future. J Antibiot (Tokyo) 67:631–644. https://doi.org/10.1038/ja.2014.111
Chang J, Zhang SJ, Jiang YW, Xu L, Yu JM, Zhou WJ, Sun X (2013) Design, synthesis, and antibacterial activity of demethylvancomycin analogues against drug-resistant bacteria. ChemMedChem 8:976–984. https://doi.org/10.1002/cmdc.201300011
Chen WH, Li K, Guntaka NS, Bruner SD (2016) Interdomain and intermodule organization in epimerization domain containing nonribosomal peptide synthetases. Acs Chem Biol 11:2293–2303. https://doi.org/10.1021/acschembio.6b00332
Chiu HT, Hubbard BK, Shah AN, Eide J, Fredenburg RA, Walsh CT, Khosla C (2001) Molecular cloning and sequence analysis of the complestatin biosynthetic gene cluster. Proc Natl Acad Sci U S A 98:8548–8553. https://doi.org/10.1073/pnas.151246498
Cryle MJ, Meinhart A, Schlichting I (2010) Structural characterization of OxyD, a cytochrome P450 involved in beta-hydroxytyrosine formation in vancomycin biosynthesis. J Biol Chem 285:24562–24574. https://doi.org/10.1074/jbc.M110.131904
Cryle MJ, Staaden J, Schlichting I (2011) Structural characterization of CYP165D3, a cytochrome P450 involved in phenolic coupling in teicoplanin biosynthesis. Arch Biochem Biophys 507:163–173. https://doi.org/10.1016/j.abb.2010.10.017
Culp EJ, Waglechner N, Wang W, Fiebig-Comyn AA, Hsu YP, Koteva K, Sychantha D, Coombes BK et al (2020) Evolution-guided discovery of antibiotics that inhibit peptidoglycan remodelling. Nature 578:582–587. https://doi.org/10.1038/s41586-020-1990-9
Drula E, Garron ML, Dogan S, Lombard V, Henrissat B, Terrapon N (2022) The carbohydrate-active enzyme database: functions and literature. Nucleic Acids Res 50:D571–D577. https://doi.org/10.1093/nar/gkab1045
Forneris CC, Seyedsayamdost MR (2018) In vitro reconstitution of OxyC activity enables total chemoenzymatic syntheses of vancomycin aglycone variants. Angew Chem Int Ed Engl 57:8048–8052. https://doi.org/10.1002/anie.201802856
Frasch HJ, Kalan L, Kilian R, Martin T, Wright GD, Stegmann E (2015) Alternative pathway to a glycopeptide-resistant cell wall in the balhimycin producer Amycolatopsis balhimycina. ACS Infect Dis 1:243–252. https://doi.org/10.1021/acsinfecdis.5b00011
Greule A, Izoré T, Iftime D, Tailhades J, Schoppet M, Zhao Y, Peschke M, Ahmed I et al (2019) Kistamicin biosynthesis reveals the biosynthetic requirements for production of highly crosslinked glycopeptide antibiotics. Nat Commun 10(1):2613–2627. https://doi.org/10.1038/s41467-019-10384-w
Greule A, Izoré T, Machell D, Hansen MH, Schoppet M, De Voss JJ, Charkoudian LK, Schittenhelm RB et al (2022) The cytochrome P450 OxyA from the kistamicin biosynthesis cyclization cascade is highly sensitive to oxidative damage. Front Chem 10:868240. https://doi.org/10.3389/fchem.2022.868240
Guan DL, Chen F, Xiong L, Tang F, Faridoon, Qiu Y, Zhang N, Gong L et al (2018) Extra sugar on vancomycin: new analogues for combating multidrug-resistant Staphylococcus aureus and vancomycin-resistant enterococci. J Med Chem 61:286–304. https://doi.org/10.1021/acs.jmedchem.7b01345
Han S, Miller SJ (2013) Asymmetric catalysis at a distance: catalytic, site-selective phosphorylation of teicoplanin. J Am Chem Soc 135:12414–12421. https://doi.org/10.1021/ja406067v
Haslinger K, Maximowitsch E, Brieke C, Koch A, Cryle MJ (2014) Cytochrome P450 OxyBtei catalyzes the first phenolic coupling step in teicoplanin biosynthesis. ChemBioChem 15(18):2719–2728. https://doi.org/10.1002/cbic.201402441
Haslinger K, Peschke M, Brieke C, Maximowitsch E, Cryle MJ (2015) X-domain of peptide synthetases recruits oxygenases crucial for glycopeptide biosynthesis. Nature 521:105–109. https://doi.org/10.1038/nature14141
Haslinger K, Cryle MJ (2016) Structure of OxyA tei: completing our picture of the glycopeptide antibiotic producing cytochrome P450 cascade. FEBS Lett 590:571–581. https://doi.org/10.1002/1873-3468.12081
Heald SL, Mueller L, Jeffs PW (1987) Actinoidins a and A2: structure determination using 2D NMR methods. J Antibiot (Tokyo) 40:630–645. https://doi.org/10.7164/antibiotics.40.630
Hu Q, Peng H, Rao X (2016) Molecular events for promotion of vancomycin resistance in Vancomycin Intermediate Staphylococcus aureus. Front Microbiol 7:1601–1617. https://doi.org/10.3389/fmicb.2016.01601
Izore T, Cryle MJ (2018) The many faces and important roles of protein-protein interactions during non-ribosomal peptide synthesis. Nat Prod Rep 35:1120–1139. https://doi.org/10.1039/c8np00038g
Jeong H, Sim YM, Kim HJ, Lee DW, Lim SK, Lee SJ (2013) Genome sequence of the vancomycin-producing Amycolatopsis orientalis subsp. orientalis strain KCTC 9412T. Genome Announc 1(3):e00408–413. https://doi.org/10.1128/genomeA.00408-13
Kaniusaite M, Tailhades J, Marschall EA, Goode RJA, Schittenhelm RB, Cryle MJ (2019) A proof-reading mechanism for non-proteinogenic amino acid incorporation into glycopeptide antibiotics. Chem Sci 10:9466–9482. https://doi.org/10.1039/c9sc03678d
Kwun MJ, Hong HJ (2014) Draft genome sequence of Amycolatopsis lurida NRRL 2430, producer of the glycopeptide family antibiotic ristocetin. Genome Announc 2(5):e01050–e01014. https://doi.org/10.1128/genomeA.01050-14
Lyu SY, Liu YC, Chang CY, Huang CJ, Chiu YH, Huang CM, Hsu NS, Lin KH et al (2014) Multiple complexes of long aliphatic N-acyltransferases lead to synthesis of 2,6-diacylated/2-acyl-substituted glycopeptide antibiotics, effectively killing vancomycin-resistant enterococcus. J Am Chem Soc 136(31):10989–10995. https://doi.org/10.1021/ja504125v
Maffioli SI, Ciabatti R, Romano G, Marzorati E, Preobrazhenskaya M, Pavlov A (2005) Synthesis and antibacterial activity of alkyl derivatives of the glycopeptide antibiotic A40926 and their amides. Bioorg Med Chem Lett 15:3801–3805. https://doi.org/10.1016/j.bmcl.2005.05.076
Mollo A, von Krusenstiern AN, Bulos JA, Ulrich V, Akerfeldt KS, Cryle MJ, Charkoudian LK (2017) P450 monooxygenase ComJ catalyses side chain phenolic cross-coupling during complestatin biosynthesis. Rsc Adv 7:35376–35384. https://doi.org/10.1039/c7ra06518c
Nazari B, Forneris CC, Gibson MI, Moon K, Schramma KR, Seyedsayamdost MR (2017) Nonomuraea sp. ATCC 55076 harbours the largest actinomycete chromosome to date and the kistamicin biosynthetic gene cluster. Medchemcomm 8:780–788. https://doi.org/10.1039/c6md00637j
Okano A, Isley NA, Boger DL (2017) Peripheral modifications of [Psi[CH2NH]tpg(4)]vancomycin with added synergistic mechanisms of action provide durable and potent antibiotics. Proc Natl Acad Sci USA 114:E5052–E5061. https://doi.org/10.1073/pnas.1704125114
Owen JG, Calcott MJ, Robins KJ, Ackerley DF (2016) Generating functional recombinant NRPS enzymes in the laboratory setting via peptidyl carrier protein engineering cell. Chem Biol 23:1395–1406. https://doi.org/10.1016/j.chembiol.2016.09.014
Peschke M, Haslinger K, Brieke C, Reinstein J, Cryle MJ (2016) Regulation of the P450 oxygenation cascade involved in glycopeptide antibiotic biosynthesis. J Am Chem Soc 138:6746–6753. https://doi.org/10.1021/jacs.6b00307
Peschke M, Brieke C, Goode RJ, Schittenhelm RB, Cryle MJ (2017) Chlorinated glycopeptide antibiotic peptide precursors improve cytochrome P450-catalyzed cyclization cascade efficiency. Biochemistry 56:1239–1247. https://doi.org/10.1021/acs.biochem.6b01102
Peschke M, Brieke C, Heimes M, Cryle MJ (2018) The thioesterase domain in glycopeptide antibiotic biosynthesis is selective for cross-linked aglycones. Acs Chem Biol 13:110–120. https://doi.org/10.1021/acschembio.7b00943
Pootoolal J, Thomas MG, Marshall CG, Neu JM, Hubbard BK, Walsh CT, Wright GD (2002) Assembling the glycopeptide antibiotic scaffold: the biosynthesis of A47934 from Streptomyces toyocaensis NRRL15009. Proc Natl Acad Sci USA 99:8962–8967. https://doi.org/10.1073/pnas.102285099
Pylypenko O, Vitali F, Zerbe K, Robinson JA, Schlichting I (2003) Crystal structure of OxyC, a cytochrome P450 implicated in an oxidative C-C coupling reaction during vancomycin biosynthesis. J Biol Chem 278:46727–46733. https://doi.org/10.1074/jbc.M306486200
Schaenzer AJ, Wright GD (2020) Antibiotic resistance by enzymatic modification of antibiotic targets trends. Mol Med 26:768–782. https://doi.org/10.1016/j.molmed.2020.05.001
Schoppet M, Peschke M, Kirchberg A, Wiebach V, Sussmuth RD, Stegmann E, Cryle MJ (2019) The biosynthetic implications of late-stage condensation domain selectivity during glycopeptide antibiotic biosynthesis. Chem Sci 10:118–133. https://doi.org/10.1039/c8sc03530j
Shawky RM, Puk O, Wietzorrek A, Pelzer S, Takano E, Wohlleben W, Stegmann E (2007) The border sequence of the balhimycin biosynthesis gene cluster from Amycolatopsis balhimycina contains bbr, encoding a StrR-like pathway-specific regulator. J Mol Microbiol Biotechnol 13:76–88. https://doi.org/10.1159/000103599
Silverman SM, Moses JE, Sharpless KB (2017) Reengineering antibiotics to combat bacterial resistance: click chemistry [1,2,3]-triazole vancomycin dimers with potent activity against MRSA and VRE. Chemistry 23:79–83. https://doi.org/10.1002/chem.201604765
Singh M, Chang J, Coffman L, Kim SJ (2017) Hidden mode of action of glycopeptide antibiotics: inhibition of wall teichoic acid biosynthesis. J Phys Chem B 121:3925–3932. https://doi.org/10.1021/acs.jpcb.7b00324
Sosio M, Stinchi S, Beltrametti F, Lazzarini A, Donadio S (2003) The gene cluster for the biosynthesis of the glycopeptide antibiotic A40926 by Nonomuraea species. Chem Biol 10:541–549. https://doi.org/10.1016/s1074-5521(03)00120-0
Sosio M, Kloosterman H, Bianchi A, de Vreugd P, Dijkhuizen L, Donadio S (2004) Organization of the teicoplanin gene cluster in Actinoplanes teichomyceticus. Microbiol (Reading) 150:95–102. https://doi.org/10.1099/mic.0.26507-0
Stokes JM, French S, Ovchinnikova OG, Bouwman C, Whitfield C, Brown ED (2016) Cold stress makes Escherichia coli susceptible to glycopeptide antibiotics by altering outer membrane integrity. Cell Chem Biol 23:267–277. https://doi.org/10.1016/j.chembiol.2015.12.011
Szucs Z, Ostorhazi E, Kicsak M, Nagy L, Borbas A, Herczegh P (2019) New semisynthetic teicoplanin derivatives have comparable in vitro activity to that of oritavancin against clinical isolates of VRE. J Antibiot (Tokyo) 72:524–534. https://doi.org/10.1038/s41429-019-0164-1
Szucs Z, Bereczki I, Rőth E, Milánkovits M, Ostorházi E, Batta G, Nagy L, Dombrádi Z et al (2020) N-Terminal guanidine derivatives of teicoplanin antibiotics strongly active against glycopeptide resistant Enterococcus faecium. J Antibiot (Tokyo) 73:603–614. https://doi.org/10.1038/s41429-020-0313-6
Tailhades J, Zhao Y, Ho YTC, Greule A, Ahmed I, Schoppet M, Kulkarni K, Goode RJA et al (2020) A chemoenzymatic approach to the synthesis of glycopeptide antibiotic analogues. Angew Chem Int Ed Engl 59:10899–10903. https://doi.org/10.1002/anie.202003726
Thaker MN, Wang W, Spanogiannopoulos P, Waglechner N, King AM, Medina R, Wright GD (2013) Identifying producers of antibacterial compounds by screening for antibiotic resistance. Nat Biotechnol 31:922–927. https://doi.org/10.1038/nbt.2685
Ulrich V, Peschke M, Brieke C, Cryle MJ (2016a) More than just recruitment: the X-domain influences catalysis of the first phenolic coupling reaction in A47934 biosynthesis by cytochrome P450 StaH. Mol Biosyst 12(10):2992–3004. https://doi.org/10.1039/c6mb00373g
Ulrich V, Brieke C, Cryle MJ (2016b) Biochemical and structural characterisation of the second oxidative crosslinking step during the biosynthesis of the glycopeptide antibiotic A47934. Beilstein J Org Chem 12:2849–2864. https://doi.org/10.3762/bjoc.12.284
van Wageningen AM, Kirkpatrick PN, Williams DH, Harris BR, Kershaw JK, Lennard NJ, Jones M, Jones SJ, Solenberg PJ (1998) Sequencing and analysis of genes involved in the biosynthesis of a vancomycin group antibiotic. Chem Biol 5:155–162. https://doi.org/10.1016/s1074-5521(98)90060-6
Varisco M, Khanna N, Brunetto PS, Fromm KM (2014) New antimicrobial and biocompatible implant coating with synergic silver-vancomycin conjugate action. ChemMedChem 9:1221–1230. https://doi.org/10.1002/cmdc.201400072
Vimberg V, Gazak R, Szűcs Z, Borbás A, Herczegh P, Cavanagh JP, Zieglerova L, Závora J et al (2019) Fluorescence assay to predict activity of the glycopeptide antibiotics. J Antibiot (Tokyo) 72:114–117. https://doi.org/10.1038/s41429-018-0120-5
Wadzinski TJ, Gea KD, Miller SJ (2016) A stepwise dechlorination/cross-coupling strategy to diversify the vancomycin ‘in-chloride’. Bioorg Med Chem Lett 26:1025–1028. https://doi.org/10.1016/j.bmcl.2015.12.027
Waglechner N, McArthur AG, Wright GD (2019) Phylogenetic reconciliation reveals the natural history of glycopeptide antibiotic biosynthesis and resistance. Nat Microbiol 4:1862–1871. https://doi.org/10.1038/s41564-019-0531-5
Walsh CT, O’Brien RV, Khosla C (2013) Nonproteinogenic amino acid building blocks for non-ribosomal peptide and hybrid polyketide scaffolds. Angew Chem Int Ed Engl 52:7098–7124. https://doi.org/10.1002/anie.201208344
Wang G, Yang ML, Duan ZL, Liu FL, Jin L, Long CB, Zhang M, Tang XP et al (2021) Dalbavancin binds ACE2 to block its interaction with SARS-CoV-2 spike protein and is effective in inhibiting SARS-CoV-2 infection in animal models. Cell Res 31:17–24. https://doi.org/10.1038/s41422-020-00450-0
Wang Z, Koirala B, Hernandez Y, Zimmerman M, Brady SF (2022a) Bioinformatic prospecting and synthesis of a bifunctional lipopeptide antibiotic that evades resistance. Science 376:991–996. https://doi.org/10.1126/science.abn4213
Wang Z, Koirala B, Hernandez Y, Zimmerman M, Park S, Perlin DS, Brady SF (2022b) A naturally inspired antibiotic to target multidrug-resistant pathogens. Nature 601:606–611. https://doi.org/10.1038/s41586-021-04264-x
Weist S, Kittel C, Bischoff D, Bister B, Pfeifer V, Nicholson GJ, Wohlleben W, Süssmuth RD (2004) Mutasynthesis of glycopeptide antibiotics: variations of vancomycin’s AB-ring amino acid 3,5-dihydroxyphenylglycine. J Am Chem Soc 126:5942–5943. https://doi.org/10.1021/ja0499389
Xu F, Wu Y, Zhang C, Davis KM, Moon K, Bushin LB, Seyedsayamdost MR (2019) A genetics-free method for high-throughput discovery of cryptic microbial metabolites. Nat Chem Biol 15:161–168. https://doi.org/10.1038/s41589-018-0193-2
Xu M, Wang W, Waglechner N, Culp EJ, Guitor AK, Wright GD (2020) GPAHex-A synthetic biology platform for type IV-V glycopeptide antibiotic production and discovery. Nat Commun 11:5232–5243. https://doi.org/10.1038/s41467-020-19138-5
Xu M, Wang W, Waglechner N, Culp EJ, Guitor AK, Wright GD (2022) Phylogeny-informed synthetic biology reveals unprecedented structural novelty in type V glycopeptide antibiotics. ACS Cent Sci 8:615–626. https://doi.org/10.1021/acscentsci.1c01389
Yarlagadda V, Manjunath GB, Sarkar P, Akkapeddi P, Paramanandham K, Shome BR, Ravikumar R, Haldar J (2016) Glycopeptide antibiotic to overcome the intrinsic resistance of Gram-negative bacteria. ACS Infect Dis 2:132–139. https://doi.org/10.1021/acsinfecdis.5b00114
Yim G, Kalan L, Koteva K, Thaker MN, Waglechner N, Tang I, Wright GD (2014) Harnessing the synthetic capabilities of glycopeptide antibiotic tailoring enzymes: characterization of the UK-68,597 biosynthetic cluster. ChemBioChem 15:2613–2623. https://doi.org/10.1002/cbic.201402179
Yim G, Wang W, Thaker MN, Tan S, Wright GD (2016) How to make a glycopeptide: a synthetic biology approach to expand antibiotic chemical diversity. ACS Infect Dis 2:642–650. https://doi.org/10.1021/acsinfecdis.6b00105
Yim G, Wang W, Pawlowski AC, Wright GD (2018) Trichlorination of a teicoplanin-type glycopeptide antibiotic by the halogenase StaI evades resistance. Antimicrob Agents Chemother 62(12):e01540–e01518. https://doi.org/10.1128/AAC.01540-18
Yushchuk O, Vior NM, Andreo-Vidal A, Berini F, Rückert C, Busche T, Binda E, Kalinowski J et al (2021) Genomic-led discovery of a novel glycopeptide antibiotic by Nonomuraea coxensis DSM 45129. Acs Chem Biol 16:915–928. https://doi.org/10.1021/acschembio.1c00170
Zerbe K, Pylypenko O, Vitali F, Zhang W, Rouset S, Heck M, Vrijbloed JW, Bischoff D et al (2002) Crystal structure of OxyB, a cytochrome P450 implicated in an oxidative phenol coupling reaction during vancomycin biosynthesis. J Biol Chem 277:47476–47485. https://doi.org/10.1074/jbc.M206342200
Zhang P, Zhang Z, Zhang L, Wang J, Wu C (2020) Glycosyltransferase GT1 family: phylogenetic distribution, substrates coverage, and representative structural features. Comput Struct Biotechnol J 18:1383–1390. https://doi.org/10.1016/j.csbj.2020.06.003
Zhao Y, Goode RJA, Schittenhelm RB, Tailhades J, Cryle MJ (2020) Exploring the tetracyclization of teicoplanin precursor peptides through chemoenzymatic synthesis. J Org Chem 85:1537–1547. https://doi.org/10.1021/acs.joc.9b02640
Zhong J, Lu Z, Dai J, He W (2017) Identification of two regulatory genes involved in carbomycin biosynthesis in Streptomyces thermotolerans. Arch Microbiol 199:1023–1033. https://doi.org/10.1007/s00203-017-1376-z
Zhou N et al (2016) Glycopeptide antibiotics potently inhibit cathepsin L in the late endosome/lysosome and block the entry of Ebola Virus, Middle East respiratory syndrome coronavirus (MERS-CoV), and severe Acute Respiratory Syndrome Coronavirus (SARS-CoV). J Biol Chem 291:9218–9232. https://doi.org/10.1074/jbc.M116.716100
Author information
Authors and Affiliations
Contributions
All the authors contributed to the Writing-Reviewing and Editing of the paper. XZ, FH, and HD share the ideas for the article. LT and SS conducted the literature search and the information summarization. LT and SS wrote the original draft.
Corresponding author
Ethics declarations
Conflict of interest
All authors seriously declare that they have no any conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Tian, L., Shi, S., Zhang, X. et al. Newest perspectives of glycopeptide antibiotics: biosynthetic cascades, novel derivatives, and new appealing antimicrobial applications. World J Microbiol Biotechnol 39, 67 (2023). https://doi.org/10.1007/s11274-022-03512-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-022-03512-0