Abstract
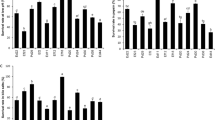
Fermented milk products are a major source of health-promoting microorganisms known as probiotics. To characterize the probiotic properties of lactic acid bacteria isolated from Ghanaian traditionally fermented milk, thirty (30) isolates comprising Enterococcus faecium (1), Lactobacillus fermentum (14), Lb. plantarum (2) and Pediococcus acidilactici (13) identified by 16S rRNA gene sequencing, were tested for survival at low pH (2.5) and bile salts (0.3% (w/v)), hydrophobicity, co-aggregation, auto-aggregation and antimicrobial activities against selected pathogens. Safety of potential probiotic bacteria was assessed by hemolytic activity on blood agar and susceptibility to nine different antibiotics. Majority (90%) of the strains showed survival rates above 80% at pH (2.5) and in bile salts (0.3% (w/v)). Hydrophobicity ranged from 5 to 61% while cell auto-aggregation ranged from 41 to 80% after 24 h. Co-aggregation with E. coli (3.7–43.9%) and S. Typhimurium (1.3–49.5%) were similar for the LAB strains at 24 h. Cell- free supernatants of all LAB strains inhibited E. coli while S. Typhimurium was not sensitive to cell-free supernatants of five Pd. acidilactici strains: OS24h20, OS18h3, OY9h19, OS9h8 and 24NL38. None of the LAB strains showed β-hemolysis but 38% of strains showed α-hemolysis. Susceptibilities to antibiotics were strain-specific; only four strains, two Lb. fermentum and two Pd. acidilactici were susceptible to all nine antibiotics tested. Based on high survival rates in bile salts, low pH and generally good hydrophobicity, auto-aggregation, co-aggregation and inhibitory activities, 15 out of 30 strains tested were considered qualified candidates for development of probiotic cultures for fermented milk products in sub-Saharan Africa.



Similar content being viewed by others
References
Abbasiliasi S et al (2017) In vitro assessment of Pediococcus acidilactici Kp10 for its potential use in the food industry. BMC Microbiol 17:121. https://doi.org/10.1186/s12866-017-1000-z
Adimpong DB, Nielsen DS, Sørensen KI, Derkx PM, Jespersen L (2012) Genotypic characterization and safety assessment of lactic acid bacteria from indigenous African fermented food products. BMC Microbiol 12:75. https://doi.org/10.1186/1471-2180-12-75
Akabanda F, Owusu-Kwarteng J, Tano-Debrah K, Glover RL, Nielsen DS, Jespersen L (2013) Taxonomic and molecular characterization of lactic acid bacteria and yeasts in nunu, a Ghanaian fermented milk product. Food Microbiol 34:277–283. https://doi.org/10.1016/j.fm.2012.09.025
Akabanda F, Owusu-Kwarteng J, Tano-Debrah K, Parkouda C, Jespersen L (2014) The use of lactic acid bacteria starter culture in the production of nunu, a spontaneously fermented milk product in Ghana. Int J Food Sci 2014:721067. https://doi.org/10.1155/2014/721067
Ammor MS, Flórez AB, Mayo B (2007) Antibiotic resistance in non-enterococcal lactic acid bacteria and bifidobacteria. Food Microbiol 24:559–570. https://doi.org/10.1016/j.fm.2006.11.001
Bao Y et al (2010) Screening of potential probiotic properties of Lactobacillus fermentum isolated from traditional dairy products. Food Control 21:695–701. https://doi.org/10.1016/j.foodcont.2009.10.010
Bayili GR, Johansen P, Nielsen DS, Sawadogo-Lingani H, Ouedraogo GA, Diawara B, Jespersen L (2019) Identification of the predominant microbiota during production of lait caillé, a spontaneously fermented milk product made in Burkina Faso. World J Microbiol Biotechnol 35:100. https://doi.org/10.1007/s11274-019-2672-3
Brandvold KR, Weaver JM, Whidbey C (2019) A continuous fluorescence assay for simple quantification of bile salt hydrolase activity in the gut microbiome. Sci Rep 9:1359. https://doi.org/10.1038/s41598-018-37656-7
Bujnakova D, Strakova E, Kmet V (2014) In vitro evaluation of the safety and probiotic properties of Lactobacilli isolated from chicken and calves. Anaerobe 29:118–127. https://doi.org/10.1016/j.anaerobe.2013.10.009
Drago L, Gismondo MR, Lombardi A, De Haën C, Gozzini L (1997) Inhibition of in vitro growth of enteropathogens by new Lactobacillus isolates of human intestinal origin. FEMS Microbiol Lett 153:455–463. https://doi.org/10.1111/j.1574-6968.1997.tb12610.x
Ejtahed H, Mohtadi-Nia J, Homayouni-Rad A, Niafar M, Asghari-Jafarabadi M, Mofid V, Akbarian-Moghari A (2011) Effect of probiotic yogurt containing Lactobacillus acidophilus and Bifidobacterium lactis on lipid profile in individuals with type 2 diabetes mellitus. J Dairy Sci 94:3288–3294. https://doi.org/10.3168/jds.2010-4128
Erginkaya Z, Turhan E, Tatlı D (2018) Determination of antibiotic resistance of lactic acid bacteria isolated from traditional Turkish fermented dairy products. Iran J Vet Res 19:53
FAO/WHO (2002) Joint FAO/WHO working group report on drafting guidelines for the evaluation of probiotics in food. London, Ontario, Canada
Franz CM, Huch M, Abriouel H, Holzapfel W, Gálvez A (2011) Enterococci as probiotics and their implications in food safety. Int J Food Microbiol 151:125–140. https://doi.org/10.1016/j.ijfoodmicro.2011.08.014
García-Cayuela T, Korany AM, Bustos I, de Cadiñanos LPG, Requena T, Peláez C, Martínez-Cuesta MC (2014) Adhesion abilities of dairy Lactobacillus plantarum strains showing an aggregation phenotype. Food Res Int 57:44–50. https://doi.org/10.1016/j.foodres.2014.01.010
Gevers D, Danielsen M, Huys G, Swings J (2003) Molecular characterization of tet (M) genes in Lactobacillus isolates from different types of fermented dry sausage. Appl Environ Microbiol 69:1270–1275
Gueimonde M, Sánchez B, GdLR-G C, Margolles A (2013) Antibiotic resistance in probiotic bacteria. Front Microbiol 4:202
Hammer Ø, Harper DA, Ryan PD (2001) PAST: Paleontological statistics software package for education and data analysis. Palaeontol Electron 4:1–9
Hati S, Mandal S, Prajapati J (2013) Novel starters for value added fermented dairy products. Curr Res Nutr Food Sci 1:83–91. https://doi.org/10.12944/CRNFSJ.1.1.09
Hill C et al (2014) Expert consensus document. The International Scientific Association for Probiotics and Prebiotics consensus statement on the scope and appropriate use of the term probiotic. Nat Rev Gastroenterol Hepatol 11:506–514. https://doi.org/10.1038/nrgastro.2014.66
Holzapfel W, Wood B (2014) Lactic acid bacteria: biodiversity and taxonomy. Wiley, Hoboken
Holzapfel WH (2002) Appropriate starter culture technologies for small-scale fermentation in developing countries. Int J Food Microbiol 75:197–212. https://doi.org/10.1016/S0168-1605(01)00707-3
Ilavenil S, Vijayakumar M, Kim DH, Valan Arasu M, Park HS, Ravikumar S, Choi KC (2016) Assessment of probiotic, antifungal and cholesterol lowering properties of Pediococcus pentosaceus KCC-23 isolated from Italian ryegrass. J Sci Food Agric 96:593–601. https://doi.org/10.1002/jsfa.7128
Jacobsen CN et al (1999) Screening of probiotic activities of forty-seven strains of Lactobacillus spp. by in vitro techniques and evaluation of the colonization ability of five selected strains in humans. Appl Environ Microbiol 65:4949–4956
Janković T, Frece J, Abram M, Gobin I (2012) Aggregation ability of potential probiotic Lactobacillus plantarum strains. Int J Sanit Eng Res 6:19–24
Kaushik JK, Kumar A, Duary RK, Mohanty AK, Grover S, Batish VK (2009) Functional and probiotic attributes of an indigenous isolate of Lactobacillus plantarum. PLoS ONE. https://doi.org/10.1371/journal.pone.0008099
Kirby LC, Klein RA, Coleman JP (1995) Continuous spectrophotometric assay of conjugated bile acid hydrolase. Lipids 30:863–867. https://doi.org/10.1007/bf02533963
Klose V, Bayer K, Kern C, Goelß F, Fibi S, Wegl G (2014) Antibiotic resistances of intestinal lactobacilli isolated from wild boars. Vet Microbiol 168:240–244. https://doi.org/10.1016/j.vetmic.2013.11.014
Kolenbrander PE (2000) Oral microbial communities: biofilms, interactions, and genetic systems. Annu Rev Microbiol 54:413–437. https://doi.org/10.1146/annurev.micro.54.1.413
Mathara JM et al (2008a) Functional characteristics of Lactobacillus spp. from traditional maasai fermented milk products in Kenya. Int J Food Microbiol 126:57–64. https://doi.org/10.1016/j.ijfoodmicro.2008.04.027
Mathara JM, Schillinger U, Kutima PM, Mbugua SK, Guigas C, Franz C, Holzapfel WH (2008b) Functional properties of Lactobacillus plantarum strains isolated from Maasai traditional fermented milk products in Kenya. Curr Microbiol 56:315–321. https://doi.org/10.1007/s00284-007-9084-6
Mathara JM, Schillinger U, Kutima PM, Mbugua SK, Holzapfel WH (2004) Isolation, identification and characterisation of the dominant microorganisms of kule naoto: the maasai traditional fermented milk in Kenya. Int J Food Microbiol 94:269–278. https://doi.org/10.1016/j.ijfoodmicro.2004.01.008
Nawaz M et al (2011) Characterization and transfer of antibiotic resistance in lactic acid bacteria from fermented food products. Curr Microbiol 62:1081–1089. https://doi.org/10.1007/s00284-010-9856-2
Noriega L, Gueimonde M, Sánchez B, Margolles A, de los Reyes-Gavilán CG (2004) Effect of the adaptation to high bile salts concentrations on glycosidic activity, survival at low pH and cross-resistance to bile salts in Bifidobacterium. Int J Food Microbiol 94:79–86. https://doi.org/10.1016/j.ijfoodmicro.2004.01.003
Obodai M, Dodd C (2006) Characterization of dominant microbiota of a Ghanaian fermented milk product, nyarmie, by culture-and nonculture-based methods. J Appl Microbiol 100:1355–1363. https://doi.org/10.1111/j.1365-2672.2006.02895.x
Ouoba LII, Lei V, Jensen LB (2008) Resistance of potential probiotic lactic acid bacteria and bifidobacteria of African and European origin to antimicrobials: determination and transferability of the resistance genes to other bacteria. Int J Food Microbiol 121:217–224. https://doi.org/10.1016/j.ijfoodmicro.2007.11.018
Owusu-Kwarteng J, Tano-Debrah K, Akabanda F, Jespersen L (2015) Technological properties and probiotic potential of Lactobacillus fermentum strains isolated from West African fermented millet dough. BMC Microbiol 15:261. https://doi.org/10.1186/s12866-015-0602-6
Ramakrishnan V, Goveas LC, Prakash M, Halami PM, Narayan B (2014) Optimization of conditions for probiotic curd formulation by Enterococcus faecium MTCC 5695 with probiotic properties using response surface methodology. J Food Sci Technol 51:3050–3060. https://doi.org/10.1007/s13197-012-0821-x
Ramos CL, Thorsen L, Schwan RF, Jespersen L (2013) Strain-specific probiotics properties of Lactobacillus fermentum, Lactobacillus plantarum and Lactobacillus brevis isolates from Brazilian food products. Food Microbiol 36:22–29. https://doi.org/10.1016/j.fm.2013.03.010
Roberts MC, Sutcliffe J, Courvalin P, Jensen LB, Rood J, Seppala H (1999) Nomenclature for macrolide and macrolide-lincosamide-streptogramin B resistance determinants. Antimicrob Agents Chemother 43:2823–2830. https://doi.org/10.1128/AAC.43.12.2823
Rojo-Bezares B, Sáenz Y, Poeta P, Zarazaga M, Ruiz-Larrea F, Torres C (2006) Assessment of antibiotic susceptibility within lactic acid bacteria strains isolated from wine. Int J Food Microbiol 111:234–240. https://doi.org/10.1016/j.ijfoodmicro.2006.06.007
Rychen G et al (2018) Guidance on the characterisation of microorganisms used as feed additives or as production organisms. EFSA J 16:1–24. https://doi.org/10.2903/j.efsa.2018.5206
Schillinger U, Guigas C, Holzapfel WH (2005) In vitro adherence and other properties of lactobacilli used in probiotic yoghurt-like products. Int Dairy J 15:1289–1297. https://doi.org/10.1016/j.idairyj.2004.12.008
Švec P, Franz CMAP (2014) The genus Enterococcus. In: Holzapfel W, Wood J (eds) Lactic acid bacteria: biodiversity and taxonomy. Wiley, Chichester, pp 175–211
Tokatlı M, Gülgör G, Bağder Elmacı S, Arslankoz İşleyen N, Özçelik F (2015) In vitro properties of potential probiotic indigenous lactic acid bacteria originating from traditional pickles. Biomed Res Int. https://doi.org/10.1155/2015/315819
Tsai Y-T, Cheng P-C, Pan T-M (2012) The immunomodulatory effects of lactic acid bacteria for improving immune functions and benefits. Appl Microbiol Biotechnol 96:853–862. https://doi.org/10.1007/s00253-012-4407-3
Vidhyasagar V, Jeevaratnam K (2013) Evaluation of Pediococcus pentosaceus strains isolated from Idly batter for probiotic properties in vitro. J Funct Foods 5:235–243. https://doi.org/10.1016/j.jff.2012.10.012
Wang Z, Zeng X, Mo Y, Smith K, Guo Y, Lin J (2012) Identification and characterization of a bile salt hydrolase from Lactobacillus salivarius for development of novel alternatives to antibiotic growth promoters. Appl Environ Microbiol 78:8795–8802. https://doi.org/10.1128/AEM.02519-12
Xu H, Jeong HS, Lee HY, Ahn J (2009) Assessment of cell surface properties and adhesion potential of selected probiotic strains. Lett Appl Microbiol 49:434–442. https://doi.org/10.1111/j.1472-765X.2009.02684.x
Zhang B, Wang Y, Tan Z, Li Z, Jiao Z, Huang Q (2016) Screening of probiotic activities of lactobacilli strains isolated from traditional Tibetan Qula, a raw yak milk cheese. Asian Australas J Anim Sci 29:1490–1499. https://doi.org/10.5713/ajas.15.0849
Zommiti M et al (2018) Evaluation of probiotic properties and safety of Enterococcus faecium isolated from artisanal Tunisian meat “Dried Ossban.” Front Microbiol 9:1685. https://doi.org/10.3389/fmicb.2018.01685
Acknowledgements
The authors would like to acknowledge Danida for providing funding for the project Preserving African Food Microorganisms for Green Growth (DFC No 13-04KU). We also appreciate the Navrongo Health Research Center for allowing their facilities to be used for aspects of the molecular work in this study.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Motey, G.A., Owusu-Kwarteng, J., Obiri-Danso, K. et al. In vitro properties of potential probiotic lactic acid bacteria originating from Ghanaian indigenous fermented milk products. World J Microbiol Biotechnol 37, 52 (2021). https://doi.org/10.1007/s11274-021-03013-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11274-021-03013-6