Abstract
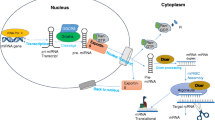
MicroRNAs (miRNAs) are increasingly being shown to play vital roles in development, apoptosis, and oncogenesis by interfering with gene expression at the post-transcriptional level. miRNAs, in principle, can contribute to the repertoire of host–pathogen interactions during infection by the Hepatitis B virus (HBV). Using a consensus-scoring approach, high-scoring miRNA-target pairs were selected, which were identified by four well-established target-prediction softwares. The miRNAs miR-7, miR196b, miR433, and miR511 target the polymerase or S gene of HBV, miR205 targets the X gene, and miR345 targets the preC gene. The minimum free-energy values for the bound complexes were the lowest, and the rules so far observed for miRNA-target pairing, namely, (1) pairing at a continuous stretch of 6–7 bases toward the 5′-end of the miRNA and (2) incomplete complementarity with the target sequence, were found to be valid. The target regions were highly conserved across the various clades of HBV. miRNA expression profiles from previously reported Solexa-sequencing based experiments showed that the four human miRNAs are expressed in the liver. This is the first report of human miRNAs that can target crucial HBV genes.



Similar content being viewed by others
References
D.P. Bartel, Cell 116, 281–297 (2004)
G.A. Calin, C.G. Liu, C. Sevignani, M. Ferracin, N. Felli, C.D. Dumitru, M. Shimizu, A. Cimmino, S. Zupo, M. Dono, M.L. Dell, A.H. Aquila, L. Rassenti, T.J. Kipps, F. Bullrich, M. Negrini, C.M. Croce, Proc. Natl. Acad. Sci. USA 101, 11755–11760 (2004)
S. Griffiths-Jones, Nucleic Acids Res. 32, D109–D111 (2004)
A. Enright, B. John, U. Gaul, T. Tuschl, C. Sander, D. Marks, Genome Biol. R1, 5 (2003)
M. Rehmsmeier, P. Steffen, M. Hochsmann, R. Giegerich, RNA 10, 1507–1517 (2004)
V. Rusinov, V. Baev, I.N. Minkov, M. Tabler, Nucleic Acids Res. 33, W696–W700 (2005)
M. Kiriakidou, P.T. Nelson, A. Kouranov, P. Fitziev, C. Bouyioukos, Z. Mourelatos, A. Hatzigeorgiou, Genes Dev. 18, 1165–1178 (2004)
T. Barrett, D.B. Troup, S.E. Wilhite, P.L.D. Rudnev, C.F. Evangelista, I.F. Kim, A. Soboleva, M. Tomashevsky, K.A. Marshall, K.H. Phillippy, P.M. Sherman, R.N. Muertter, R. Edgar. Nucleic Acids Res. 37(Database issue), D885–D890 (2009)
W.B. Jin, F.L. Wu, D. Kong, A.G. Guo, Computat Biol Chem. 31, 124–126 (2007)
C.L. Jopling, M. Yi, A.M. Lancaster, S.M. Lemon, P. Sarnow, Science 309, 1577–1581 (2005)
C.H. Lecellier, P. Dunoyer, K. Arar, J. Lehmann-Che, S. Eyquem, C. Himber, A. Saib, O. Voinnet, Science 308, 557–560 (2005)
Y. Tay, J. Zhang, A.M. Thomson, B. Lim, I. Rigoutsos, Nature 455, 1124–1129 (2008)
Acknowledgments
This work was supported by grant no. SJ08-ZT04 from Provincial Natural Science Foundation of Shaanxi and the grant No. 31000913 from the National Natural Science Foundation of China.
Author information
Authors and Affiliations
Corresponding author
Additional information
Fang-li Wu and Wei-Bo Jin contributed equally to this work.
Rights and permissions
About this article
Cite this article
Wu, Fl., Jin, WB., Li, Jh. et al. Targets for human encoded microRNAs in HBV genes. Virus Genes 42, 157–161 (2011). https://doi.org/10.1007/s11262-010-0555-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11262-010-0555-7