Abstract
Proper chloroplast development and chlorophyll biosynthesis are essential for the photoautotrophic plants. The insertion of magnesium (Mg2+) into protoporphyrin IX (Proto), catalyzed by magnesium chelatase (Mg-chelatase), is the first committed step of chlorophyll biosynthesis. In dicot plants, a proposed model revealed that Mg-chelatase I subunit (CHLI) and Mg-chelatase D subunit (CHLD) can interact directly; however, their relation remains elusive in rice, a monocot model plant. In this study, we characterized a chlorophyll-deficiency mutant, etiolated leaf and lethal (ell), which displayed a yellow leaf in young seedlings and became lethal after three-leaf stage. Chlorophyll content in homozygous ell mutant was approximately 1 % of that in the wild type. Besides, chloroplast development in the mutant was entirely arrested and no thylakoid structure was observed. By map-based cloning, the ell locus was delimited to a 3.9-Mb interval in chromosome 3. A single-base mutation (G529C) in OsCHLI was identified, leading to an amino acid substitution (G177R) in a highly conserved region. Compared with the wild type, more Proto but less magnesium protoporphyrin IX (Mg-Proto) was measured in the ell mutant. Using protoplast transfection and callus transformation, we found that exogenous OsCHLI could consistently recover the lesion of chloroplast in the ell mutant. By subcellar localization analysis, OsCHLI was detected in the chloroplast. Despite the secondary structure of OsCHLI that was predicted to be altered in the mutant, the point mutation did not affect subcellular localization. Real-time PCR demonstrated that the ell mutation induced significantly transcriptional downregulation of the photosynthesis-associated nuclear and plastid genes. Additionally, yeast-two-hybrid experiments indicated that the single amino acid substitution blocked the intrinsic interaction between OsCHLI and OsCHLD. Moreover, OsCHLI showed physical interactions with some thioredoxins (TRXs), suggesting a similar regulatory mechanism of Mg-chelatase activity in both monocot and dicot plants.








Similar content being viewed by others
Abbreviations
- Proto:
-
Protoporphyrin IX
- Mg-chelatase:
-
Magnesium chelatase
- ell :
-
Etiolated leaf and lethal
- Mg-Proto:
-
Magnesium protoporphyrin IX
- TRX:
-
Thioredoxin
- EMS:
-
Ethyl methyl sulfonate
- CHLH:
-
Mg-chelatase H subunit
- CHLI:
-
Mg-chelatase I subunit
- CHLD:
-
Mg-chelatase D subunit
- Pchlide:
-
Protochlorophyllide
References
Arsova B, Hoja U, Wimmelbacher M, Greiner E, Üstün Ş, Melzer M, Börnke F (2010) Plastidial thioredoxin z interacts with two fructokinase-like proteins in a thiol-dependent manner: evidence for an essential role in chloroplast development in Arabidopsis and Nicotiana benthamiana. Plant Cell 22:1498–1515
Bollivar DW, Suzuki JY, Beatty JT, Dobrowolski JM, Bauer CE (1994) Directed mutational analysis of bacteriochlorophyll a biosynthesis in Rhodobacter capsulatus. J Mol Biol 237:622–640
Brindley AA, Raux E, Leech HK, Schubert HL, Warren MJ (2003) A story of chelatase evolution: identification and characterization of a small 13-15 kDa “ancestral” cobaltochelatase (CbiXS) in the archaea. J Biol Chem 278:22388–22395
Coomber SA, Chaudhri M, Connor A, Britton G, Hunter CN (1990) Localized transposon Tn5 mutagenesis of the photosynthetic gene cluster of Rhodobacter sphaeroides. Mol Microbiol 4:977–989
Czarnecki O, Grimm B (2012) Post-translational control of tetrapyrrole biosynthesis in plants, algae, and cyanobacteria. J Exp Bot 63:1675–1687
Dellaporta SL, Hicks JB (1983) A new plant DNA minipreparation: version II. Plant Mol Biol Report 1:19–21
Erzberger JP, Berger JM (2006) Evolutionary relationships and structural mechanisms of AAA+ proteins. Annu Rev Biophys Biomol Struct 35:93–114
Fodje MN, Hansson A, Hansson M, Olsen JG, Gough S, Willows RD, Al-Karadaghi S (2001) Interplay between an AAA module and an integrin I domain may regulate the function of magnesium chelatase. J Mol Biol 311:111–122
Gibson LC, Willows RD, Kannangara CG, von Wettstein D, Hunter CN (1995) Magnesium-protoporphyrin chelatase of Rhodobacter sphaeroides: reconstitution of activity by combining the products of the bchH, -I, and -D genes expressed in Escherichia coli. Proc Natl Acad Sci U S A 92:1941–1944
Hansson A, Kannangara CG, von Wettstein D, Hansson M (1999) Molecular basis for semidominance of missense mutations in the XANTHA-H (42-kDa) subunit of magnesium chelatase. Proc Natl Acad Sci 96:1744–1749
Hansson A, Willows RD, Roberts TH, Hansson M (2002) Three semidominant barley mutants with single amino acid substitutions in the smallest magnesium chelatase subunit form defective AAA+ hexamers. Proc Natl Acad Sci 99:13944–13949
Hiei Y, Ohta S, Komari T, Kumashiro T (1994) Efficient transformation of rice (Oryza sativa L.) mediated by Agrobacterium and sequence analysis of the boundaries of the T-DNA. Plant J 6:271–282
Hoober JK, Boyd CO, Paavola LG (1991) Origin of thylakoid membranes in Chlamydomonas reinhardtii y-1 at 38 °C. Plant Physiol 96:1321–1328
Huang YS, Li HM (2009) Arabidopsis CHLI2 can substitute for CHLI1. Plant Physiol 150:636–645
Ikegami A, Yoshimura N, Motohashi K, Takahashi S, Romano PG, Hisabori T, Masuda T (2007) The CHLI1 subunit of Arabidopsis thaliana magnesium chelatase is a target protein of the chloroplast thioredoxin. J Biol Chem 282:19282–19291
Iyer LM, Leipe DD, Koonin EV, Aravind L (2004) Evolutionary history and higher order classification of AAA+ ATPases. J Struct Biol 146:11–31
Jensen PE, Gibson LCD, Henningsen KW, Hunter CN (1996) Expression of the chlI, chlD, and chlH genes from the cyanobacterium Synechocystis PCC6803 in Escherichia coli and demonstration that the three cognate proteins are required for magnesium-protoporphyrin chelatase activity. J Biol Chem 271:16662–16667
Jensen P, Gibson L, Hunter C (1999) ATPase activity associated with the magnesium-protoporphyrin IX chelatase enzyme of Synechocystis PCC6803: evidence for ATP hydrolysis during Mg2+ insertion, and the MgATP-dependent interaction of the ChlI and ChlD subunits. Biochem J 339:127–134
Joyard J, Ferro M, Masselon C, Seigneurin-Berny D, Salvi D, Garin J, Rolland N (2009) Chloroplast proteomics and the compartmentation of plastidial isoprenoid biosynthetic pathways. Mol Plant 2:1154–1180
Kannangara CG, Gough SP, Bruyan P, Hoober JK, Kahn A, Von Wettstein D (1988) tRNAGlu as a cofactor in δ-aminolevulinate biosynthesis: steps that regulate chlorophyll synthesis. Trends Biochem Sci 13:139–143
Koncz CS, Mayerhofer R, Koncz-Kalman Z, Nawrath C, Reiss B, Redei GP, Schell J (1990) Isolation of a gene encoding a novel chloroplast protein by T-DNA tagging in Arabidopsis thaliana. EMBO J 9:1337–1346
Larkin RM, Alonso JM, Ecker JR, Chory J (2003) GUN4, a regulator of chlorophyll synthesis and intracellular signaling. Science 299:902–906
Lee HJ, Ball MD, Parham R, Rebeiz CA (1992) Chloroplast biogenesis 65. Enzymic conversion of protoporphyrin IX to Mg-protoporphyrin IX in a subplastidic membrane fraction of cucumber etiochloroplasts. Plant Physiol 99:1134–1140
Lichtenthaler HK (1987) Chlorophylls and carotenoids: pigments of photosynthetic biomembranes. Methods Enzymol 148:350–382
Lundqvist J, Elmlund H, Wulff RP, Berglund L, Elmlund D, Emanuelsson C, Al-Karadaghi S (2010) ATP-induced conformational dynamics in the AAA+ motor unit of magnesium chelatase. Structure 18:354–365
Luo T, Fan T, Liu Y, Rothbart M, Yu J, Zhou S, Luo M (2012) Thioredoxin redox regulates ATPase activity of magnesium chelatase CHLI subunit and modulates redox-mediated signaling in tetrapyrrole biosynthesis and homeostasis of reactive oxygen species in pea plants. Plant Physiol 159:118–130
Masuda T (2008) Recent overview of the Mg branch of the tetrapyrrole biosynthesis leading to chlorophylls. Photosynth Res 96:121–143
Matsumoto F, Obayashi T, Sasaki-Sekimoto Y, Ohta H, Takamiya K, Masuda T (2004) Gene expression profiling of the tetrapyrrole metabolic pathway in Arabidopsis with a mini-array system. Plant Physiol 135:2379–2391
Miernyk JA, Thelen JJ (2008) Biochemical approaches for discovering protein-protein interactions. Plant J 53:597–609
Mochizuki N, Brusslan JA, Larkin R, Nagatani A, Chory J (2001) Arabidopsis genomes uncoupled 5 (GUN5) mutant reveals the involvement of Mg-chelatase H subunit in plastid-to-nucleus signal transduction. Proc Natl Acad Sci U S A 98:2053–2058
Nakayama M, Masuda T, Sato N, Yamagata H, Bowler C, Ohta H, Takamiya KI (1995) Cloning, subcellular localization and expression of chl1, a subunit of magnesium chelatase in soybean. Biochem Biophys Res Commun 215:422–428
Nelson BK, Cai X, Nebenführ A (2007) A multicolored set of in vivo organelle markers for co-localization studies in Arabidopsis and other plants. Plant J 51:1126–1136
Pérez-Ruiza JM, Guineaa M, Puerto-Galána L, Cejudoa FJ (2014) NADPH thioredoxin reductase C is involved in redox regulation of the Mg-chelatase I subunit in Arabidopsis thaliana chloroplasts. Mol Plant 7:1252–1255
Petersen BL, Jensen PE, Gibson LC, Stummann BM, Hunter CN, Henningsen KW (1998) Reconstitution of an active magnesium chelatase enzyme complex from the bchI, -D, and -H gene products of the green sulfur bacterium Chlorobium vibrioforme expressed in Escherichia coli. J Bacteriol 180:699–704
Pogson BJ, Albrecht V (2011) Genetic dissection of chloroplast biogenesis and development: an overview. Plant Physiol 155:1545–1551
Reinhold T, Alawady A, Grimm B, Beran KC, Jahns P, Conrath U, Neuhaus HE (2007) Limitation of nocturnal import of ATP into Arabidopsis chloroplasts leads to photooxidative damage. Plant J 50:293–304
Richter AS, Peter E, Rothbart M, Schlicke H, Toivola J, Rintamäki E, Grimm B (2013) Posttranslational influence of NADPH-dependent thioredoxin reductase C on enzymes in tetrapyrrole synthesis. Plant Physiol 162:63–73
Rissler HM, Collakova E, DellaPenna D, Whelan J, Pogson BJ (2002) Chlorophyll biosynthesis. Expression of a second chl I gene of magnesium chelatase in Arabidopsis supports only limited chlorophyll synthesis. Plant Physiol 128:770–779
Sawers RJ, Viney J, Farmer PR, Bussey RR, Olsefski G, Anufrikova K, Brutnell TP (2006) The maize Oil yellow1 (Oy1) gene encodes the I subunit of magnesium chelatase. Plant Mol Biol 60:95–106
Solymosi K, Schoefs B (2010) Etioplast and etio-chloroplast formation under natural conditions: the dark side of chlorophyll biosynthesis in angiosperms. Photosynth Res 105:143–166
Tagwerker C, Flick K, Cui M, Guerrero C, Dou Y, Auer B, Baldi P, Huang L, Kaiser P (2006) A tandem affinity tag for two-step purification under fully denaturing conditions: application in ubiquitin profiling and protein complex identification combined with in vivo cross-linking. Mol Cell Proteomics 5:737–748
Tanaka R, Kobayashi K, Masuda T (2011) Tetrapyrrole metabolism in Arabidopsis thaliana. The Arabidopsis Book 9:e0145
Tripathy BC, Pattanayak GK (2012) Chlorophyll biosynthesis in higher plants. In: Eaton-Rye JJ, Tripathy BC, Sharkey TD (eds) Photosynthesis: plastid biology, energy conversion and carbon assimilation, advances in photosynthesis and respiration, vol 34. Springer, The Netherlands, pp 63–94
Tripathy BC, Rebeiz CA (1985) Chloroplast biogenesis: quantitative determination of monovinyl and divinyl Mg-protoporphyrins and protochlorophyll(ides) by spectrofluorometry. Anal Biochem 149:43–61
Voinnet O, Rivas S, Mestre P, Baulcombe D (2003) An enhanced transient expression system in plants based on suppression of gene silencing by the p19 protein of tomato bushy stunt virus. Plant J 33:949–956
Waadt R, Kudla J (2008) In planta visualization of protein interactions using bimolecular fluorescence complementation (BiFC). Cold Spring Harbor Protoc. port4995
Waadt R, Schmidt LK, Lohse M, Hashimoto K, Bock R, Kudla J (2008) Multicolor bimolecular fluorescence complementation reveals simultaneous formation of alternative CBL/CIPK complexes in planta. Plant J 56:505–516
Willows RD, Beale SI (1998) Heterologous expression of the Rhodobacter capsulatus BchI, -D, and -H genes that encode magnesium chelatase subunits and characterization of the reconstituted enzyme. J Biol Chem 273:34206–34213
Yoshida R, Kanno A, Sato T, Kameya T (1996) Cool-temperature-induced chlorosis in rice plants (I. Relationship between the induction and a disturbance of etioplast development). Plant Physiol 110:997–1005
Zhang H, Li J, Yoo JH, Yoo SC, Cho SH, Koh HJ, Paek NC (2006) Rice Chlorina-1 and Chlorina-9 encode ChlD and ChlI subunits of Mg-chelatase, a key enzyme for chlorophyll synthesis and chloroplast development. Plant Mol Biol 62:325–337
Zhang Y, Su J, Duan S, Ao Y, Dai J, Liu J, Wang H (2011) A highly efficient rice green tissue protoplast system for transient gene expression and studying light/chloroplast-related processes. Plant Methods 7:30
Acknowledgments
This project was financially supported by “the Fundamental Research Funds for the Central Universities (KYY201301)”, “the Key Science and Technology Specific Projects of Guizhou Province (Grant no. 2012-6005)”, the National Science and Technology supporting program (2013BAD01B02-16), the High Technology Program from NDRC ([2012]1961), and the Jiangsu Science and Technology Development Program (BE2012303, BE2013301).
Author information
Authors and Affiliations
Corresponding authors
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Supplementary Table S1
Summary of primers (DOCX 16 kb)
Supplementary Fig. S1
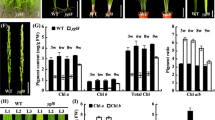
Abundance of Proto, Mg-Proto and Pchilide in wild type and ell mutant. Proto accumulated in ell mutant, but Mg-Proto and Pchilide were decreased compared with the wild type. WT, wild type; **p < 0.01 (GIF 15 kb)
Supplementary Fig. S2
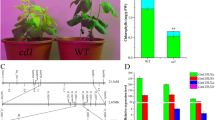
Expression analysis of photosynthesis-associated nuclear and plastid genes in the wild type and ell mutant. rpoA (the RNA polymerase A subunit), rpoB (the RNA polymerase B subunit), Lhc (light-harvesting complex protein), psbA (a core component of photosystem II), and rps7 (ribosomal protein S7). Relative amounts were recalculated using the level of Actin transcript as internal control; Nip, Nipponbare; *p < 0.05; **p < 0.01 (GIF 20 kb)
Supplementary Fig. S3
Negative controls of BiFC assay. YFP, fluorescence of yellow fluorescent protein; Chl, chlorophyll autofluorescence; ER-mCherry, fluorescence of ER marker, mCherry ER rk CD3-959; Merged, merged image of YFP, Chl and ER-mCherry; bars = 10 μm (GIF 442 kb)
Rights and permissions
About this article
Cite this article
Zhang, H., Liu, L., Cai, M. et al. A Point Mutation of Magnesium Chelatase OsCHLI Gene Dampens the Interaction Between CHLI and CHLD Subunits in Rice. Plant Mol Biol Rep 33, 1975–1987 (2015). https://doi.org/10.1007/s11105-015-0889-3
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11105-015-0889-3