Abstract
Aims
Soil legacy effects can have long-term impacts on soil microbial communities with implications for plant growth and community structure. These effects are well studied for invasive plants, particularly after removal of invasive species; however, we know less about the soil legacy effects post removal of native range expanding species.
Methods
We used a controlled greenhouse experiment with a range-expanding sagebrush species (Artemisia rothrockii (Asteraceae)) to determine how multiple metrics of sagebrush seedling performance (plant-soil feedback (PSF) ratio, height, leaf functional traits, and root:shoot biomass) were influenced by soil legacy effects in both the native and expansion range and over time since removal. We inoculated seedlings with field-collected soils from under sagebrush canopies and in herbaceous interspace, as well as in areas where sagebrush had been removed for 1 or 5 years. We then used ITS2 sequencing and extracellular enzyme assays to characterize the structure and function of soil microbial communities and to determine what microbial mechanisms drove seedling responses.
Results
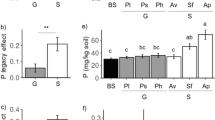
Conspecific sagebrush seedlings responded negatively to soil legacy effects of shrub removal, with a more negative PSF ratio, reduced height, and higher root:shoot ratios in shrub removal inoculum than in shrub and herbaceous soil inoculum. Seedlings in shrub removal inoculum also had enriched foliar isotope ratios, reflecting higher resource use efficiency. Soil communities of seedlings with shrub removal inoculum had increased fungal diversity, pathogen, and saprotroph richness, and altered fungal community composition. Legacy effects on soil fungal diversity and functional group richness were present in seedlings with 1-year shrub removal inoculum, while effects on fungal community composition were found in 1 and 5-year shrub removal inoculated seedlings. Despite changes in functional group richness, fungal diversity and community composition proved the strongest drivers of seedling performance overall.
Conclusions
This work provides novel insight into how soil legacy effects post removal of a native range expanding species may limit rather than promote the performance of conspecifics over short and long time periods, with important implications for management as global change continues to shift the geographic ranges of woody plants.






Similar content being viewed by others
Data availability
Fungal sequencing data can be found in the NCBI SRA database Accession # PRJNA924843 and all other data and analysis scripts can be found on Github https://github.com/cour10eygrace/Sagebrush_GH_PSFs.
References
Angert AL, Crozier LG, Rissler LJ et al (2011) Do species’ traits predict recent shifts at expanding range edges? Ecol Lett 14:677–689. https://doi.org/10.1111/j.1461-0248.2011.01620.x
Archer SR, Andersen EM, Predick KI et al (2017) Woody plant encroachment: causes and consequences. In: Briske DD (ed) Rangeland systems: processes, management and challenges. Springer International Publishing, Cham, pp 25–84
Austin AT, Vivanco L, González-Arzac A, Pérez LI (2014) There’s no place like home? An exploration of the mechanisms behind plant litter-decomposer affinity in terrestrial ecosystems. New Phytol 204:307–314. https://doi.org/10.1111/nph.12959
Bates D, Mächler M, Bolker B, Walker S (2014) Fitting linear mixed-effects models using lme4. J Stat Softw 67:. https://doi.org/10.18637/jss.v067.i01
Baxendale C, Orwin KH, Poly F et al (2014) Are plant-soil feedback responses explained by plant traits? J Physiol 204:408–423. https://doi.org/10.1111/nph.12915
Bennett JA, Franklin J, Ectomycorrhiza EM (2022) Plant - soil feedbacks persist following tree death, reducing survival and growth of Populus tremuloides seedlings. Plant Soil. https://doi.org/10.1007/s11104-022-05645-5
Berlow EL, D’Antonio CM, Swartz H (2003) Response of herbs to shrub removal across natural and experimental variation in soil moisture. Ecol Appl 13:1375–1387. https://doi.org/10.1890/02-5099
Bever JD (2003) Soil community feedback and the coexistence of competitors: conceptual frameworks and empirical tests. New Phytol 157:465–473. https://doi.org/10.1046/j.1469-8137.2003.00714.x
Bever JD, Westover KM, Antonovics J (1997) Incorporating the soil community into plant population dynamics: the utility of the feedback approach. J Ecol 85(5):561–573. https://doi.org/10.2307/2960528
Boddy L, Watkinson SC (1995) Wood decomposition, higher fungi, and their role in nutrient redistribution. Can J Bot 73:1377–1383. https://doi.org/10.1139/b95-400
Bonner FT, Karrfalt RP (2008) The woody plant seed manual. Agriculture Handbook 727. U.S. Department of Agriculture, Forest Service, Washington, DC, p 1223. https://www.fs.usda.gov/rm/pubs_series/wo/wo_ah727.pdf
Brandt AJ, De Kroon H, Reynolds HL, Burns JH (2013) Soil heterogeneity generated by plant-soil feedbacks has implications for species recruitment and coexistence. J Ecol 101:277–286. https://doi.org/10.1111/1365-2745.12042
Broadbent AAD, Bahn M, Pritchard WJ et al (2022) Shrub expansion modulates belowground impacts of changing snow conditions in alpine grasslands. Ecol Lett 25:52–64
Burnham KP, Anderson DR, Huyvaert KP (2011) AIC model selection and multimodel inference in behavioral ecology: Some background, observations, and comparisons. Behav Ecol Sociobiol 65:23–35. https://doi.org/10.1007/s00265-010-1029-6
Callaway RM, Cipollini D, Barto K et al (2008) Novel weapons: Invasive plant suppresses fungal mutualists in America but not in its native europe. Ecology 89:1043–1055. https://doi.org/10.1890/07-0370.1
Cantarel AAM, Pommier T, Desclos-Theveniau M et al (2015) Using plant traits to explain plant-microbe relationships involved in nitrogen acquisition. Ecology 96:788–799
Caporaso JG, Kuczynski J, Stombaugh J et al (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7:335
Caravaca F, Rodríguez-Caballero G, Campoy M et al (2020) The invasion of semiarid Mediterranean sites by Nicotiana glauca mediates temporary changes in mycorrhizal associations and a permanent decrease in rhizosphere activity. Plant Soil 450:217–229. https://doi.org/10.1007/s11104-020-04497-1
Chung YA, Rudgers JA (2016) Plant–soil feedbacks promote negative frequency dependence in the coexistence of two aridland grasses. Proc R Soc B Biol Sci 283:. https://doi.org/10.1098/rspb.2016.0608
Coats VC, Rumpho ME (2014) The rhizosphere microbiota of plant invaders: An overview of recent advances in the microbiomics of invasive plants. Front Microbiol 5:1–10. https://doi.org/10.3389/fmicb.2014.00368
Collins CG, Carey CJ, Aronson EL et al (2016) Direct and indirect effects of native range expansion on soil microbial community structure and function. J Ecol 104:1271–1283. https://doi.org/10.1111/1365-2745.12616
Collins CG, Stajich JE, Weber SE et al (2018) Shrub range expansion alters diversity and distribution of soil fungal communities across an alpine elevation gradient. Mol Ecol 27:2461–2476. https://doi.org/10.1111/mec.14694
Collins CG, Bohner TF, Diez JM (2019) Plant-soil feedbacks and facilitation influence the demography of herbaceous alpine species in response to woody plant range expansion. Front Ecol Evol 7:1–15. https://doi.org/10.3389/fevo.2019.00417
Collins CG, Spasojevic MJ, Alados CL et al (2020) Belowground impacts of alpine woody encroachment are determined by plant traits, local climate and soil conditions. Glob Chang Biol. https://doi.org/10.1111/gcb.15340
De Cáceres M, Legendre P (2009) Associations between species and groups of sites: indices and statistical inference. Ecology 90:3566–3574. https://doi.org/10.1890/08-1823.1
Duell EB, Zaiger K, Bever JD, Wilson GWT (2019) Climate affects plant-soil feedback of native and invasive grasses: negative feedbacks in stable but not in variable environments. Front Ecol Evol 7:1–10. https://doi.org/10.3389/fevo.2019.00419
Dufrêne M, Legendre P (1997) Species assemblages and indicator Specie: the need for a flexible asymetrical approach. Ecol Monogr 67:345–366. https://doi.org/10.1890/0012-9615(1997)067[0345:SAAIST]2.0.CO;2
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26:2460–2461. https://doi.org/10.1093/bioinformatics/btq461
Edgar RC (2013) UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat Methods 10:996
Eldridge DJ, Ding J (2021) Remove or retain: ecosystem effects of woody encroachment and removal are linked to plant structural and functional traits. New Phytol 229:2637–2646. https://doi.org/10.1111/nph.17045
Eldridge DJ, Beecham G, Grace JB (2015) Do shrubs reduce the adverse effects of grazing on soil properties? Ecohydrology. https://doi.org/10.1002/eco.1600
Elgersma KJ, Ehrenfeld JG, Yu S, Vor T (2011) Legacy effects overwhelm the short-term effects of exotic plant invasion and restoration on soil microbial community structure, enzyme activities, and nitrogen cycling. Oecologia 167:733–745
Eppinga MB, Rietkerk M, Dekker SC et al (2006) Accumulation of local pathogens: a new hypothesis to explain exotic plant invasions. Oikos 114:168–176. https://doi.org/10.1111/j.2006.0030-1299.14625.x
Esch CM, Kobe RK (2021) Short-lived legacies of Prunus serotina plant–soil feedbacks. Oecologia 196:529–538. https://doi.org/10.1007/s00442-021-04948-1
Friesen ML, Porter SS, Stark SC et al (2011) Microbially mediated plant functional traits. Annu Rev Ecol Evol Syst 42:23–46. https://doi.org/10.1146/annurev-ecolsys-102710-145039
GarcíaCriado M, Myers-Smith IH, Bjorkman AD et al (2020) Woody plant encroachment intensifies under climate change across tundra and savanna biomes. Glob Ecol Biogeogr 29:925–943. https://doi.org/10.1111/geb.13072
German DP, Chacon SS, Allison SD (2011) Substrate concentration and enzyme allocation can affect rates of microbial decomposition. Ecology 92:1471–1480. https://doi.org/10.1890/10-2028.1
Grove S, Parker IM, Haubensak KA (2015) Persistence of a soil legacy following removal of a nitrogen-fixing invader. Biol Invasions 17:2621–2631. https://doi.org/10.1007/s10530-015-0900-9
Gundale MJ, Kardol P (2021) Multi-dimensionality as a path forward in plant-soil feedback research. J Ecol 109:3446–3465. https://doi.org/10.1111/1365-2745.13679
Gundale MJ, Wardle DA, Kardol P, Nilsson MC (2019) Comparison of plant–soil feedback experimental approaches for testing soil biotic interactions among ecosystems. New Phytol 221:577–587. https://doi.org/10.1111/nph.15367
Hall CA Jr (ed) (1991) Natural History of the white-inyo range, Eastern California. University of California Press, Berkeley, p c1991. http://ark.cdlib.org/ark:/13030/ft3t1nb2pn/
Hannula SE, Heinen R, Huberty M et al (2021) Persistence of plant-mediated microbial soil legacy effects in soil and inside roots. Nat Commun 12:1–13. https://doi.org/10.1038/s41467-021-25971-z
Jančič S, Nguyen HDT, Frisvad JC et al (2015) A taxonomic revision of the Wallemia sebi species complex. PLoS ONE 10:1–25. https://doi.org/10.1371/journal.pone.0125933
Johnson NC, Graham JH, Smith FA (1997) Functioning of mycorrhizal associations along the mutualism-parasitism continuum. New Phytol 135:575–586. https://doi.org/10.1046/j.1469-8137.1997.00729.x
Kardol P, Veen GF (Ciska), Teste FP, Perring MP (2015) Peeking into the black box: a trait- based approach to predicting plant – soil feedback. New Phytol 206:1–4. https://doi.org/10.1111/nph.13283
Ke PJ, Miki T, Ding TS (2015) The soil microbial community predicts the importance of plant traits in plant-soil feedback. New Phytol 206:329–341. https://doi.org/10.1111/nph.13215
Koorem K, Snoek BL, Bloem J et al (2020) Community-level interactions between plants and soil biota during range expansion. J Ecol 108:1860–1873. https://doi.org/10.1111/1365-2745.13409
Kopp CW, Cleland EE (2014) Shifts in plant species elevational range limits and abundances observed over nearly five decades in a western North America mountain range. J Veg Sci 25:135–146. https://doi.org/10.1111/jvs.12072
Kopp CW, Cleland EE (2018) Plant community response to Artemisia rothrockii (sagebrush) encroachment and removal along an arid elevational gradient. J Veg Sci 29:859–866. https://doi.org/10.1111/jvs.12669
Kostenko O, Bezemer TM (2020) Abiotic and biotic soil legacy effects of plant diversity on plant performance. Front Ecol Evol 8:1–12. https://doi.org/10.3389/fevo.2020.00087
Kulmatiski A, Beard KH, Stevens JR, Cobbold SM (2008) Plant-soil feedbacks: a meta-analytical review. Ecol Lett 11:980–992. https://doi.org/10.1111/j.1461-0248.2008.01209.x
Kuťáková E, Herben T, Münzbergová Z (2018) Heterospecific plant–soil feedback and its relationship to plant traits, species relatedness, and co-occurrence in natural communities. Oecologia 187:679–688. https://doi.org/10.1007/s00442-018-4145-z
Lankau RA, Bauer JT, Anderson MR, Anderson RC (2014) Long-term legacies and partial recovery of mycorrhizal communities after invasive plant removal. Biol Invasions 16:1979–1990. https://doi.org/10.1007/s10530-014-0642-0
Lau JA, Lennon JT (2011) Evolutionary ecology of plant–microbe interactions: soil microbial structure alters selection on plant traits. New Phytol 192:215–224. https://doi.org/10.1111/j.1469-8137.2011.03790.x
Lee Taylor D, Sinsabaugh RL (2015) The soil fungi, 4th edn. Elsevier Inc, Burlington, MA & Oxford, UK
Li K, Veen GF, ten Hooven FC et al (2022) Soil legacy effects of plants and drought on aboveground insects in native and range-expanding plant communities. Ecol Lett:1–16. https://doi.org/10.1111/ele.14129
Lloret F, Casanovas C, Peñuelas J (1999) Seedling survival of Mediterranean shrubland species in relation to root:shoot ratio, seed size and water and nitrogen use. Funct Ecol 13:210–216. https://doi.org/10.1046/j.1365-2435.1999.00309.x
Lozano YM, Aguilar-Trigueros CA, Roy J, Rillig MC (2021) Drought induces shifts in soil fungal communities that can be linked to root traits across twenty-four plant species. New Phytol. https://doi.org/10.1111/nph.17707
MacLean SA, Beissinger SR (2017) Species’ traits as predictors of range shifts under contemporary climate change: A review and meta-analysis. Glob Chang Biol 23:4094–4105. https://doi.org/10.1111/gcb.13736
Manrubia M, Snoek LB, Weser C et al (2019) Belowground consequences of intracontinental range-expanding plants and related natives in novel environments. Front Microbiol 10:1–13. https://doi.org/10.3389/fmicb.2019.00505
Martinez Arbizu P (2020) pairwiseAdonis: Pairwise multilevel comparison using adonis. R package version 0.4
May JL, Oberbauer SF (2021) Simulated hurricane-induced changes in light and nutrient regimes change seedling performance in Everglades forest-dominant species. Ecol Evol 11:17762–17773. https://doi.org/10.1002/ece3.8273
Mayerhofer MS, Kernaghan G, Harper KA (2013) The effects of fungal root endophytes on plant growth: A meta-analysis. Mycorrhiza 23:119–128. https://doi.org/10.1007/s00572-012-0456-9
Mooney H, Andre G, Wright R (1962) Alpine and subalpine vegetation patterns in the white mountains of California. Am Midl Nat 68:257–273
Morrien E, Engelkes T, Macel M et al (2010) Climate change and invasion by intracontinental range-expanding exotic plants: the role of biotic interactions. Ann Bot 105:843–848. https://doi.org/10.1093/aob/mcq064
Muff S, Nilsen EB, O’Hara RB, Nater CR (2022) Rewriting results sections in the language of evidence. Trends Ecol Evol 37:203–210. https://doi.org/10.1016/j.tree.2021.10.009
Newsham KK (2011) A meta-analysis of plant responses to dark septate root endophytes. New Phytol 190:783–793. https://doi.org/10.1111/j.1469-8137.2010.03611.x
Nguyen NH, Song Z, Bates ST et al (2016) FUNGuild: An open annotation tool for parsing fungal community datasets by ecological guild. Fungal Ecol 20:241–248. https://doi.org/10.1016/j.funeco.2015.06.006
Ohm RA, Feau N, Henrissat B et al (2012) Diverse lifestyles and strategies of plant pathogenesis encoded in the genomes of eighteen dothideomycetes fungi. PLoS Pathog 8:e1003037. https://doi.org/10.1371/journal.ppat.1003037
Oksanen J, Blanchet F, Kindt R et al (2016) Vegan: community ecology package. R Packag 23–3 Available at: https://cran.r-project.org/web/packa
Olson Å, Aerts A, Asiegbu F et al (2012) Insight into trade-off between wood decay and parasitism from the genome of a fungal forest pathogen. New Phytol 194:1001–1013. https://doi.org/10.1111/j.1469-8137.2012.04128.x
Palmer JM, Jusino MA, Banik MT, Lindner DL (2018) Non-biological synthetic spike-in controls and the AMPtk software pipeline improve mycobiome data. bioRxiv. https://doi.org/10.1101/213470
Pérez-Harguindeguy N, Díaz S, Garnier E et al (2016) New handbook for standardised measurement of plant functional traits worldwide. Aust J Bot 64:715–716. https://doi.org/10.1071/BT12225_CO
Pernilla Brinkman E, Van der Putten WH, Bakker E-J, Verhoeven KJF (2010) Plant-soil feedback: experimental approaches, statistical analyses and ecological interpretations. J Ecol 98:1063–1073. https://doi.org/10.1111/j.1365-2745.2010.01695.x
Petipas RH, Geber MA, Lau JA (2021) Microbe-mediated adaptation in plants. Ecol Lett 24:1302–1317. https://doi.org/10.1111/ele.13755
Pinto-Figueroa EA, Seddon E, Yashiro E et al (2019) Archaeorhizomycetes spatial distribution in soils along wide elevational and environmental gradients reveal co-abundance patterns with other fungal saprobes and potential weathering capacities. Front Microbiol 10:1–13. https://doi.org/10.3389/fmicb.2019.00656
Purahong W, Pietsch KA, Bruelheide H et al (2019) Potential links between wood-inhabiting and soil fungal communities: Evidence from high-throughput sequencing. Microbiologyopen 8:1–14. https://doi.org/10.1002/mbo3.856
R Core Team (2020) R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. https://www.R-project.org/
Ramirez KS, Snoek LB, Koorem K et al (2019) Range-expansion effects on the belowground plant microbiome. Nat Ecol Evol 3:604–611. https://doi.org/10.1038/s41559-019-0828-z
Revilla TA., Veen GF (Ciska), Eppinga MB, Weissing FJ (2013) Plant-soil feedbacks and the coexistence of competing plants. Theor Ecol 6:99–113. https://doi.org/10.1007/s12080-012-0163-3
Rodríguez-Echeverría S, Lozano YM, Bardgett RD (2016) Influence of soil microbiota in nurse plant systems. Funct Ecol 30:30–40. https://doi.org/10.1111/1365-2435.12594
Rognes T, Flouri T, Nichols B et al (2016) VSEARCH: a versatile open source tool for metagenomics. PeerJ 4:e2584–e2584. https://doi.org/10.7717/peerj.2584
Rosling A, Cox F, Cruz-martinez K et al (2011) Archaeorhizomycetes : unearthing an ancient class of ubiquitous soil fungi. Science (80- ) 333:876–879
Rundel PW, Gibson AC, Sharifi MR (2005) Plant functional groups in alpine fellfield habitats of the White Mountains, California. Arctic Antarct Alp Res 37:358–365
Rundel P, Gibson A, Sharifi M (2008) The alpine flora of the White Mountains, California. Madroño 55:202–215
Saiya-Cork K, Sinsabaugh R, Zak D (2002) The effects of long term nitrogen deposition on extracellular enzyme activity in an Acer saccharum forest soil. Soil Biol Biochem 34:1309–1315. https://doi.org/10.1016/S0038-0717(02)00074-3
Scavo A, Abbate C, Mauromicale G (2019) Plant allelochemicals: agronomic, nutritional and ecological relevance in the soil system. Plant Soil 442:23–48. https://doi.org/10.1007/s11104-019-04190-y
Schadt CW, Martin AP, Lipson DA, Schmidt SK (2003) Seasonal dynamics of previously unknown fungal lineages in tundra soils. Science (80- ) 301:1359–1361. https://doi.org/10.1126/science.1086940
Schoch CL, Seifert KA, Huhndorf S et al (2012) Nuclear ribosomal internal transcribed spacer (ITS) region as a universal DNA barcode marker for Fungi. Proc Natl Acad Sci 109:6241–6246. https://doi.org/10.1073/pnas.1117018109
Semchenko M, Barry KE, de Vries FT et al (2022) Deciphering the role of specialist and generalist plant–microbial interactions as drivers of plant–soil feedback. New Phytol 234:1929–1944. https://doi.org/10.1111/nph.18118
Semchenko M, Leff JW, Lozano YM et al (2018) Fungal diversity regulates plant-soil feedbacks in temperate grassland. Sci Adv 4:. https://doi.org/10.1126/sciadv.aau4578
Smithers BV (2017) Soil preferences in germination and survival of limber pine in the Great Basin White Mountains. Forests 8:. https://doi.org/10.3390/f8110423
Spasojevic MJ, Weber S (2021) Variation in δ13C and δ15N within and among plant species in the alpine tundra. Arctic Antarct Alp Res 53:340–351. https://doi.org/10.1080/15230430.2021.2000567
Stinson KA, Campbell SA, Powell JR et al (2006) Invasive plant suppresses the growth of native tree seedlings by disrupting belowground mutualisms. PLoS Biol 4:727–731. https://doi.org/10.1371/journal.pbio.0040140
Suding KN, Stanley Harpole W, Fukami T et al (2013) Consequences of plant-soil feedbacks in invasion. J Ecol 101:298–308. https://doi.org/10.1111/1365-2745.12057
Tedersoo L, Sánchez-Ramírez S, Kõljalg U et al (2018) High-level classification of the Fungi and a tool for evolutionary ecological analyses. Fungal Divers 90:135–159. https://doi.org/10.1007/s13225-018-0401-0
Throop HL, Archer SR, McClaran MP (2020) Soil organic carbon in drylands: shrub encroachment and vegetation management effects dwarf those of livestock grazing. Ecol Appl 0:1–13. https://doi.org/10.1002/eap.2150
Tomiolo S, Ward D (2018) Species migrations and range shifts: A synthesis of causes and consequences. Perspect Plant Ecol Evol Syst 33:62–77. https://doi.org/10.1016/j.ppees.2018.06.001
Treseder KK, Lennon T (2015) Fungal traits that drive ecosystem dynamics on land. 79:243–262. https://doi.org/10.1128/MMBR.00001-15
Van de Voorde TFJ, van der Putten WH, Martijn Bezemer T (2011) Intra- and interspecific plant-soil interactions, soil legacies and priority effects during old-field succession. J Ecol 99:945–953. https://doi.org/10.1111/j.1365-2745.2011.01815.x
Van der Putten WH, Bardgett RD, Bever JD et al (2013) Plant-soil feedbacks: The past, the present and future challenges. J Ecol 101:265–276. https://doi.org/10.1111/1365-2745.12054
Van Nuland ME, Wooliver RC, Pfennigwerth AA et al (2016) Plant–soil feedbacks: connecting ecosystem ecology and evolution. Funct Ecol 30:1032–1042. https://doi.org/10.1111/1365-2435.12690
Walther GR, Roques A, Hulme PE et al (2009) Alien species in a warmer world: risks and opportunities. Trends Ecol Evol 24:686–693. https://doi.org/10.1016/j.tree.2009.06.008
Weber CF, King GM, Aho K (2015) Relative abundance of and composition within fungal orders differ between cheatgrass (Bromus tectorum) and sagebrush (Artemisia tridentate)-associated soils. PLoS ONE 10:1–22. https://doi.org/10.1371/journal.pone.0117026
Westerband AC, Funk JL, Barton KE (2021) Intraspecific trait variation in plants: a renewed focus on its role in ecological processes. Ann Bot 127:397–410. https://doi.org/10.1093/aob/mcab011
White TJ, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA Genes for phylogenetics. In: PCR - Protocols and Applications - A Laboratory Manual. pp 315–322
Wickham H (2009) Elegant graphics for data analysis, 2nd edn. Springer Publishing Company, Incorporated, New York, NY
Williams RJ, Hallgren SW, Wilson GWT, Palmer MW (2013) Juniperus virginiana encroachment into upland oak forests alters arbuscular mycorrhizal abundance and litter chemistry. Appl Soil Ecol 65:23–30. https://doi.org/10.1016/j.apsoil.2012.12.020
Xi N, Chu C, Bloor JMG (2018) Plant drought resistance is mediated by soil microbial community structure and soil-plant feedbacks in a savanna tree species. Environ Exp Bot 155:695–701. https://doi.org/10.1016/j.envexpbot.2018.08.013
Xi N, Adler PB, Chen D et al (2021) Relationships between plant–soil feedbacks and functional traits. J Ecol 109:3411–3423. https://doi.org/10.1111/1365-2745.13731
Yin BF, Zhang YM, Lou AR (2017) Impacts of the removal of shrubs on the physiological and biochemical characteristics of Syntrichia caninervis Mitt: In a temperate desert. Sci Rep 7:1–12. https://doi.org/10.1038/srep45268
Yuan J, Zhao J, Wen T et al (2018) Root exudates drive the soil-borne legacy of aboveground pathogen infection. Microbiome 6:156. https://doi.org/10.1186/s40168-018-0537-x
Zanne AE, Abarenkov K, Afkhami ME et al (2020) Fungal functional ecology: bringing a trait-based approach to plant-associated fungi. Biol Rev 95:409–433. https://doi.org/10.1111/brv.12570
Funding
Research funding was provided by the University of California Riverside Department of Botany and Plant Sciences, Department of Plant Pathology and Microbiology and a UCNRS White Mountain Research station mini-grant. This research project is partly supported by grants for development of new faculty staff, Ratchadaphiseksomphot Fund, Chulalongkorn University to Nuttapon Pombubpa. Courtney G. Collins was supported by a Biodiversity Research Centre Postdoctoral Fellowship at the University of British Columbia.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Field sampling, experimental design, data collection and analysis were performed by Courtney G. Collins. Nuttapon Pombubpa completed the bioinformatics for the DNA sequencing data. The first draft of the manuscript was written by Courtney G. Collins and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Responsible Editor: Jonathan Richard De Long.
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Collins, C.G., Spasojevic, M.J., Pombubpa, N. et al. Legacy effects post removal of a range-expanding shrub influence soil fungal communities and create negative plant-soil feedbacks for conspecific seedlings. Plant Soil 485, 143–165 (2023). https://doi.org/10.1007/s11104-023-05896-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11104-023-05896-w