Key message
We genome-wide identified 28 JmjC domain-containing genes, further spatio-temporal expression profiling and genetic analysis defined them as epigenetic regulators in flowering initiation of Rosa chinensis.
Abstract
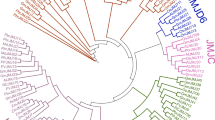
The JmjC domain-containing histone demethylases play critical roles in maintaining homeostasis of histone methylations, thus are vital for plant growth and development. Genome-wide identification of the JmjC domain-containing genes have been reported in several species, however, no systematic study has been performed in rose plants. In this paper, we identified 28 JmjC domain-containing genes from the newly published genome database of Rosa chinensis. The JmjC domain-containing proteins in R. chinensis were divided into seven groups, KDM3 was the largest group with 13 members, and JmjC domain-only A and KDM5B were the smallest clades both with only one member. Although all the JmjC domain proteins having a conserved JmjC domain, the gene and protein structure experienced differentiation and specification during the evolution, especially in KDM3 clade, one gene (RcJMJ40) was found carrying site deletions for cofactors Fe (II) and α-KG binding which were crucial for demethylase activities, three genes (RcJMJ41, RcJMJ43 and RcJMJ44) had no intron while two of them had tandem JmjC domains. Spatial expression pattern analysis of these JmjC domain-containing genes in different tissues showed most of them were highly expressed in reproductive tissues such as floral meristem and closed flowers than vegetative tissues, demonstrating their important functions in developmental switch from vegetative to reproductive growth of roses. Temporal expression profiling indicated majority of JmjC domain-containing genes from R. chinensis fluctuated along with floral bud differentiation and development, further proving their essential roles in flower organogenesis. VIGS induced silencing of RcJMJ12 led to delayed flowering time, and decreased the expression levels of flowering integrator such as RcFT, RcSOC1, RcFUL, RcLFY and RcAP1, therefore providing the genetic evidence of RcJMJ12 in flowering initiation. Collectively, spatio-temporal expression profiling and genetic analysis defined the JmjC domain-containing genes as important epigenetic regulators in flower development of R. chinensis.







Similar content being viewed by others
References
Allis DG, Fedor AM, Korter TM, Bjarnason JE, Brown ER (2007) Assignment of the lowest-lying THz absorption signatures in biotin and lactose monohydrate by solid-state density functional theory. Chem Phys Lett 440:203–209
Bendahmane M, Dubois A, Raymond O, Bris ML (2013) Genetics and genomics of flower initiation and development in roses. J Exp Bot 64:847–857
Binda E, Marcone GL, Berini F, Pollegioni L, Marinelli F (2013) Streptomyces spp. as efficient expression system for a D,D-peptidase/D,D-carboxypeptidase involved in glycopeptide antibiotic resistance. BMC Biotechnol 13: 24.
Chen Z, Zang J, Whetstine J, Hong X, Davrazou F, Kutateladze TG, Simpson M, Mao Q, Pan CH, Dai S, Haqman J, Hansen K, Shi Y, Zhang G (2006) Structural insights into histone demethylation by JMJD2 family members. Cell 125:691–702
Dubois A, Carrere S, Raymond O, Pouvreau B, Cottret L, Roccia A, Onesto JP, Sakr S, Atanassova R, Baudino S, Foucher F, Le Bris M, Gouzy J, Bendahmane M (2012) Transcriptome database resource and gene expression atlas for the rose. BMC Genomics 13:638
Fedorov VV, Voronin VV, Lapin EG (1992) On the search for neutron EDM using Laue diffraction by a crystal without a centre of symmetry. J Phys G 18:1133–1148
Feng S, Jacobsen SE (2011) Epigenetic modifications in plants: an evolutionary perspective. Curr Opin Plant Biol 14:179–186
Finn RD, Bateman A, Clements J, Coggill P, Eberhardt RY, Eddy SR, Heger A, Hetherington K, Holm L, Mistry J, Sonnhammer EL, Tate J, Punta M (2014) Pfam: the protein families database. Nucleic Acids Res 42:222–230
Gambino G, Perrone I, Gribaudo I (2008) A rapid and effective method for RNA extraction from different tissues of grapevine and other woody plants. Phytochem Anal 19:520–525
Gelato KA, Fischle W (2008) Role of histone modifications in defining chromatin structure and function. Biol Chem 389:353–363
Han Y, Li X, Lin C, Liu Y, Wang H, Ke D, Yuan H, Zhang L, Wang L (2016) Genome-wide analysis of soybean JmjC domain-containing proteins suggests evolutionary conservation following whole-genome duplication. Front Plant Sci 7:1800
Huang F, Chandrasekharan MB, Chun YC, Bhaskara S, Hiebert SW, Sun ZW (2010) The JmjN domain of Jhd2 is important for its protein stability, and the plant homeodomain (PHD) finger mediates its chromatin association independent of H3K4 methylation. J Biol Chem 285:24548–24561
Huang Y, Chen D, Liu C, Shen W, Ruan Y (2016) Evolution and conservation of JmjC domain proteins in the green lineage. Mol Genet Genomics 291:33–49
Jia B, Tang K, Chun BH, Jeon CO (2017) Large-scale examination of functional and sequence diversity of 2-oxoglutarate/Fe(II)-dependent oxygenases in Metazoa. Biochim Biophys Acta Gen Subj 1861:2922–2933
Klie M, Debener T (2011) Identification of superior reference genes for data normalisation of expression studies via quantitative PCR in hybrid roses (Rosa hybrida). BMC Res Notes. 4:518
Klose RJ, Kallin EM, Zhang Y (2006) JmjC-domain-containing proteins and histone demethylation. Nat Rev Genet 7:715–727
Kouzarides T (2007) Chromatin modifications and their function. Cell 128:693–705
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874
Letunic I, Doerks T, Bork P (2012) SMART 7: recent updates to the protein domain annotation resource. Nucleic Acids Res 40:302–305
Li T, Chen X, Zhong X, Zhao Y, Liu X, Zhou S, Cheng S, Zhou DX (2013) Jumonji C domain protein JMJ705-mediated removal of histone H3 lysine 27 trimethylation is involved in defense-related gene activation in rice. Plant Cell 25:4725–4736
Liu J, Fu X, Dong Y, Lu J, Ren M, Zhou N, Wang C (2018) MIKCC-type MADS-box genes in Rosa chinensis: the remarkable expansion of ABCDE model genes and their roles in floral organogenesis. Hortic Res 5:25
Lu F, Li G, Cui X, Liu C, Wang XJ, Cao X (2008) Comparative analysis of JmjC domain-containing proteins reveals the potential histone demethylases in Arabidopsis and rice. J Integr Plant Biol 50:886–896
Lu F, Cui X, Zhang S, Liu C, Cao X (2010) JMJ14 is an H3K4 demethylase regulating flowering time in Arabidopsis. Cell Res 20:387–390
Lu F, Cui X, Zhang S, Jenuwein T, Cao X (2011a) Arabidopsis REF6 is a histone H3 lysine 27 demethylase. Nat Genet 43:715–719
Lu SX, Knowles SM, Webb CJ, Celaya RB, Cha C, Siu JP, Tobin EM (2011b) The Jumonji C domain-containing protein JMJ30 regulates period length in the Arabidopsis circadian clock. Plant Physiol 155:906–915
Luo M, Hung FY, Yang S, Liu X, Wu K (2014) Histone lysine demethylases and their functions in plants. Plant Mol Biol Rep 32:558–565
Marchler-Bauer A, Lu S, Anderson JB, Chitsaz F, Derbyshire MK, DeWeese-Scott C, Fong JH, Geer LY, Geer RC, Gonzales NR, Gwadz M, Hurwitz DI, Jackson JD, Ke Z, Lanczycki CJ, Lu F, Marchler GH, Mullokandov M, Omelchenko MV, Robertson CL, Song JS, Thanki N, Yamashita RA, Zhang D, Zhang N, Zheng C, Bryant SH (2011) CDD: a conserved domain database for the functional annotation of proteins. Nucleic Acids Res 39:D225–D229
Mosammaparast N, Shi Y (2010) Reversal of histone methylation: biochemical and molecular mechanisms of histone demethylases. Annu Rev Biochem 79:155–179
Pontvianne F, Blevins T, Pikaard CS (2010) Arabidopsis histone lysine methyltransferases. Adv Bot Res 53:1–22
Quan Z, Oliver SG, Zhang N (2011) JmjN interacts with JmjC to ensure selective proteolysis of Gis1 by the proteasome. Microbiology 157:2694–2701
Raymond O, Gouzy J, Just J, Badouin H, Verdenaud M, Lemainque A, Vergne P, Moja S, Choisne N, Pont C, Carrère S, Caissard JC, Couloux A, Cottret L, Aury JM, Szécsi J, Latrasse D, Madoui MA, Francois L, Fu X, Yang SH, Dubois A, Piola F, Larrieu A, Perez M, Labadie K, Perrier L, Govetto B, Labrousse Y, Villand P, Bardoux C, Boltz V, Lopez-Roques C, Heitzler P, Vernoux T, Vandenbussche M, Quesneville H, Boualem A, Bendahmane A, Liu C, Le Bris M, Salse J, Baudino S, Benhamed M, Wincker P, Bendahmane M (2018) The Rosa genome provides new insights into the domestication of modern roses. Nat Genet 50:772–777
Rotili D, Mai A (2011) Targeting histone demethylases: a new avenue for the fight against cancer. Genes Cancer 2:663–679
Sun Q, Zhou DX (2008) Rice jmjC domain-containing gene JMJ706 encodes H3K9 demethylase required for floral organ development. Proc Natl Acad Sci USA 105:13679–13684
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Trewick SC, McLaughlin PJ, Allshire RC (2005) Methylation: lost in hydroxylation? EMBO Rep 6:315–320
Tsukada Y, Fang J, Erdjument-Bromage H, Warren ME, Borchers CH, Tempst P, Zhang Y (2006) Histone demethylation by a family of JmjC domain-containing proteins. Nature 439:811–816
van Leeuwen F, Gottschling DE (2002) Genome-wide histone modifications: gaining specificity by preventing promiscuity. Curr Opin Cell Biol 14:756–762
Wheeler TJ, Clements J, Eddy SR, Hubley R, Jones TA, Jurka J, Smit AF, Finn RD (2013) Dfam: a database of repetitive DNA based on profile hidden Markov models. Nucleic Acids Res 41:D70–82
Yang W, Jiang D, Jiang J, He Y (2010) A plant-specific histone H3 lysine 4 demethylase represses the floral transition in Arabidopsis. Plant J 62:663–673
Yokoo T, Saito H, Yoshitake Y, Xu Q, Asami T, Tsukiyama T, Teraishi M, Okumoto Y, Tanisaka T (2014) Se14, encoding a JmjC domain-containing protein, plays key roles in long-day suppression of rice flowering through the demethylation of H3K4me3 of RFT1. PLoS ONE 9:e96064
Zhang S, Feng M, Chen W, Zhou X, Lu J, Wang Y, Li Y, Jiang CZ, Gan SS, Ma N, Gao J (2019) In rose, transcription factor PTM balances growth and drought survival via PIP2;1 aquaporin. Nat Plants 5:290–299
Acknowledgements
This work was supported by the NSFC-XINJIANG joint foundation (U1803102) and NSFC (31972449) granted to Changquan Wang, and the China Postdoctoral Science Foundation (2016M600425) and the NSFC (31801890) granted to Jinyi Liu, and a Project Funded by the Priority Academic Program Development of Jiangsu Higher Education Institutions.
Author information
Authors and Affiliations
Contributions
Methodology, YD; formal analysis, JL; data curation, JL; validation, JA; writing—original draft, CW.
Corresponding author
Ethics declarations
Conflicts of interest
The authors declare no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
11103_2019_955_MOESM2_ESM.pdf
Fig. S2. The represented RT-PCR data of JmjC-domain containing genes in different tissues of Rosa chinensis. (PDF 513 kb)
11103_2019_955_MOESM3_ESM.pdf
Fig. S3. The represented RT-PCR data of JmjC-domain containing genes across different floral organ initiation stages of Rosa chinensis. (PDF 472 kb)
11103_2019_955_MOESM4_ESM.pdf
Fig. S4. Expression profile of JmjC-domain containing genes across different floral organ initiation stages from Rosa chinensis organized according to the expression pattern. (PDF 176 kb)
11103_2019_955_MOESM5_ESM.pdf
Fig. S5. The expression levels of RcJMJ11 and RcJMJ13 in RcJMJ12-silenced seedlings. Relative expression levels of RcJMJ11 (a) and RcJMJ13 (b) were determined by qRT-PCR in three independent RcJMJ12 silenced lines and mock treated seedlings of R. chinensis, RcGAPDH was used as an internal control. Three biological replicates were performed for each experiment. (PDF 68 kb)
Rights and permissions
About this article
Cite this article
Dong, Y., Lu, J., Liu, J. et al. Genome-wide identification and functional analysis of JmjC domain-containing genes in flower development of Rosa chinensis. Plant Mol Biol 102, 417–430 (2020). https://doi.org/10.1007/s11103-019-00955-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-019-00955-2