Abstract
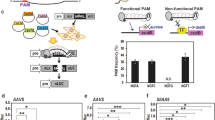
Genome editing is one of the most powerful tools for revealing gene function and improving crop plants. Recently, RNA-guided genome editing using the type II clustered regularly interspaced short palindromic repeats (CRISPR)-associated protein (Cas) system has been used as a powerful and efficient tool for genome editing in various organisms. Here, we report genome editing in tobacco (Nicotiana tabacum) mediated by the CRISPR/Cas9 system. Two genes, NtPDS and NtPDR6, were used for targeted mutagenesis. First, we examined the transient genome editing activity of this system in tobacco protoplasts, insertion and deletion (indel) mutations were observed with frequencies of 16.2–20.3 % after transfecting guide RNA (gRNA) and the nuclease Cas9 in tobacco protoplasts. The two genes were also mutated using multiplexing gRNA at a time. Additionally, targeted deletions and inversions of a 1.8-kb fragment between two target sites in the NtPDS locus were demonstrated, while indel mutations were also detected at both the sites. Second, we obtained transgenic tobacco plants with NtPDS and NtPDR6 mutations induced by Cas9/gRNA. The mutation percentage was 81.8 % for NtPDS gRNA4 and 87.5 % for NtPDR6 gRNA2. Obvious phenotypes were observed, etiolated leaves for the psd mutant and more branches for the pdr6 mutant, indicating that highly efficient biallelic mutations occurred in both transgenic lines. No significant off-target mutations were obtained. Our results show that the CRISPR/Cas9 system is a useful tool for targeted mutagenesis of the tobacco genome.





Similar content being viewed by others
References
Ansai S, Kinoshita M (2014) Targeted mutagenesis using CRISPR/Cas system in medaka. Biol Open: BIO20148177
Baltes NJ, Gil-Humanes J, Cermak T, Atkins PA, Voytas DF (2014) DNA replicons for plant genome engineering. Plant Cell Online 26:151–163
Beerli RR, Barbas CF (2002) Engineering polydactyl zinc-finger transcription factors. Nat Biotechnol 20:135–141
Bibikova M, Golic M, Golic KG, Carroll D (2002) Targeted chromosomal cleavage and mutagenesis in Drosophila using zinc-finger nucleases. Genetics 161:1169–1175
Bogdanove AJ, Voytas DF (2011) TAL effectors: customizable proteins for DNA targeting. Science 333:1843–1846
Cai CQ, Doyon Y, Ainley WM, Miller JC, DeKelver RC, Moehle EA, Rock JM, Lee Y-L, Garrison R, Schulenberg L (2009) Targeted transgene integration in plant cells using designed zinc finger nucleases. Plant Mol Biol 69:699–709
Cermak T, Doyle EL, Christian M, Wang L, Zhang Y, Schmidt C, Baller JA, Somia NV, Bogdanove AJ, Voytas DF (2011) Efficient design and assembly of custom TALEN and other TAL effector-based constructs for DNA targeting. Nucleic Acids Res 39:e82
Chen S, Songkumarn P, Liu J, Wang G-L (2009) A versatile zero background T-vector system for gene cloning and functional genomics. Plant Physiol 150:1111–1121
Chen K, Shan Q, Gao C (2014) An efficient TALEN mutagenesis system in rice. Methods 69:2–8
Christian M, Qi Y, Zhang Y, Voytas DF (2013) Targeted mutagenesis of Arabidopsis thaliana using engineered TAL effector nucleases. G3-Genes Genomes Genet 3:1697–1705. doi:10.1534/g3.113.007104
Cong L, Ran FA, Cox D, Lin S, Barretto R, Habib N, Hsu PD, Wu X, Jiang W, Marraffini LA (2013) Multiplex genome engineering using CRISPR/Cas systems. Science 339:819–823
Coutu C, Brandle J, Brown D, Brown K, Miki B, Simmonds J, Hegedus DD (2007) pORE: a modular binary vector series suited for both monocot and dicot plant transformation. Transgenic Res 16:771–781
Curtin SJ, Zhang F, Sander JD, Haun WJ, Starker C, Baltes NJ, Reyon D, Dahlborg EJ, Goodwin MJ, Coffman AP (2011) Targeted mutagenesis of duplicated genes in soybean with zinc-finger nucleases. Plant Physiol 156:466–473
De Pater S, Neuteboom LW, Pinas JE, Hooykaas PJ, Van Der Zaal BJ (2009) ZFN-induced mutagenesis and gene-targeting in Arabidopsis through Agrobacterium-mediated floral dip transformation. Plant Biotechnol J 7:821–835
Deng D, Yan C, Pan X, Mahfouz M, Wang J, Zhu J-K, Shi Y, Yan N (2012) Structural basis for sequence-specific recognition of DNA by TAL effectors. Science 335:720–723
D’Halluin K, Vanderstraeten C, Hulle J, Rosolowska J, Den Brande I, Pennewaert A, D’Hont K, Bossut M, Jantz D, Ruiter R (2013) Targeted molecular trait stacking in cotton through targeted double-strand break induction. Plant Biotechnol J 11:933–941
Doyon Y, McCammon JM, Miller JC, Faraji F, Ngo C, Katibah GE, Amora R, Hocking TD, Zhang L, Rebar EJ (2008) Heritable targeted gene disruption in zebrafish using designed zinc-finger nucleases. Nat Biotechnol 26:702–708
Feng Z, Zhang B, Ding W, Liu X, Yang D-L, Wei P, Cao F, Zhu S, Zhang F, Mao Y (2013) Efficient genome editing in plants using a CRISPR/Cas system. Cell Res 23:1229
Feng Z, Mao Y, Xu N, Zhang B, Wei P, Yang D-L, Wang Z, Zhang Z, Zheng R, Yang L (2014) Multigeneration analysis reveals the inheritance, specificity, and patterns of CRISPR/Cas-induced gene modifications in Arabidopsis. Proc Natl Acad Sci 111:4632–4637
Fu Y, Foden JA, Khayter C, Maeder ML, Reyon D, Joung JK, Sander JD (2013) High-frequency off-target mutagenesis induced by CRISPR-Cas nucleases in human cells. Nat Biotechnol 31:822–826
Gasiunas G, Barrangou R, Horvath P, Siksnys V (2012) Cas9-crRNA ribonucleoprotein complex mediates specific DNA cleavage for adaptive immunity in bacteria. Proc Natl Acad Sci 109:E2579–E2586
Gupta A, Hall VL, Kok FO, Shin M, McNulty JC, Lawson ND, Wolfe SA (2013) Targeted chromosomal deletions and inversions in zebrafish. Genome Res 23:1008–1017
Hruscha A, Krawitz P, Rechenberg A, Heinrich V, Hecht J, Haass C, Schmid B (2013) Efficient CRISPR/Cas9 genome editing with low off-target effects in zebrafish. Development 140:4982–4987
Hsu PD, Scott DA, Weinstein JA, Ran FA, Konermann S, Agarwala V, Li Y, Fine EJ, Wu X, Shalem O (2013) DNA targeting specificity of RNA-guided Cas9 nucleases. Nat Biotechnol 31:827–832
Jao L-E, Wente SR, Chen W (2013) Efficient multiplex biallelic zebrafish genome editing using a CRISPR nuclease system. Proc Natl Acad Sci 110:13904–13909
Jia H, Wang N (2014) Targeted genome editing of sweet orange using Cas9/sgRNA. PLoS One 9:e93806
Jiang W, Bikard D, Cox D, Zhang F, Marraffini LA (2013a) RNA-guided editing of bacterial genomes using CRISPR-Cas systems. Nat Biotechnol 31:233–239
Jiang W, Zhou H, Bi H, Fromm M, Yang B, Weeks DP (2013b) Demonstration of CRISPR/Cas9/sgRNA-mediated targeted gene modification in Arabidopsis, tobacco, sorghum and rice. Nucleic Acids Res 41. doi:10.1093/nar/gkt780
Jiang W, Yang B, Weeks DP (2014) Efficient CRISPR/Cas9-mediated gene editing in Arabidopsis thaliana and inheritance of modified genes in the T2 and T3 generations. PLoS One 9:e99225
Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E (2012) A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337:816–821
Kim Y-G, Cha J, Chandrasegaran S (1996) Hybrid restriction enzymes: zinc finger fusions to Fok I cleavage domain. Proc Natl Acad Sci 93:1156–1160
Lee HJ, Kim E, Kim J-S (2010) Targeted chromosomal deletions in human cells using zinc finger nucleases. Genome Res 20:81–89
Li T, Liu B, Spalding MH, Weeks DP, Yang B (2012) High-efficiency TALEN-based gene editing produces disease-resistant rice. Nat Biotechnol 30:390–392
Li J-F, Norville JE, Aach J, McCormack M, Zhang D, Bush J, Church GM, Sheen J (2013) Multiplex and homologous recombination-mediated genome editing in Arabidopsis and Nicotiana benthamiana using guide RNA and Cas9. Nat Biotechnol 31:688–691
Liang Z, Zhang K, Chen K, Gao C (2014) Targeted Mutagenesis in Zea mays using TALENs and the CRISPR/Cas System. J Genet Genomics 41:63–68
Lin Y, Cradick TJ, Brown MT, Deshmukh H, Ranjan P, Sarode N, Wile BM, Vertino PM, Stewart FJ, Bao G (2014) CRISPR/Cas9 systems have off-target activity with insertions or deletions between target DNA and guide RNA sequences. Nucleic Acids Res 42:7473–7485
Liu Y, Ma S, Wang X, Chang J, Gao J, Shi R, Zhang J, Lu W, Liu Y, Zhao P (2014) Highly efficient multiplex targeted mutagenesis and genomic structure variation in Bombyx mori cells using CRISPR/Cas9. Insect Biochem Mol Biol 49:35–42
Lloyd A, Plaisier CL, Carroll D, Drews GN (2005) Targeted mutagenesis using zinc-finger nucleases in Arabidopsis. Proc Natl Acad Sci USA 102:2232–2237
Ma S, Wang X, Liu Y, Gao J, Zhang S, Shi R, Chang J, Zhao P, Xia Q (2014) Multiplex genomic structure variation mediated by TALEN and ssODN. BMC Genom 15:41
Mahfouz MM, Li L, Shamimuzzaman M, Wibowo A, Fang X, Zhu J-K (2011) De novo-engineered transcription activator-like effector (TALE) hybrid nuclease with novel DNA binding specificity creates double-strand breaks. Proc Natl Acad Sci 108:2623–2628
Mali P, Yang L, Esvelt KM, Aach J, Guell M, DiCarlo JE, Norville JE, Church GM (2013) RNA-guided human genome engineering via Cas9. Science 339:823–826
Mani M, Kandavelou K, Dy FJ, Durai S, Chandrasegaran S (2005) Design, engineering, and characterization of zinc finger nucleases. Biochem Biophys Res Commun 335:447–457
Mao Y, Zhang H, Xu N, Zhang B, Gou F, Zhu J-K (2013) Application of the CRISPR-Cas system for efficient genome engineering in plants. Mol Plant 6:2008–2011
Marton I, Zuker A, Shklarman E, Zeevi V, Tovkach A, Roffe S, Ovadis M, Tzfira T, Vainstein A (2010) Nontransgenic genome modification in plant cells. Plant Physiol 154:1079–1087
Murashige T, Skoog F (1962) A revised medium for rapid growth and bio assays with tobacco tissue cultures. Physiol Plant 15:473–497
Mussolino C, Morbitzer R, Lütge F, Dannemann N, Lahaye T, Cathomen T (2011) A novel TALE nuclease scaffold enables high genome editing activity in combination with low toxicity. Nucleic Acids Res 39:9283–9293
Nekrasov V, Staskawicz B, Weigel D, Jones JD, Kamoun S (2013) Targeted mutagenesis in the model plant Nicotiana benthamiana using Cas9 RNA-guided endonuclease. Nat Biotechnol 31:691–693
Osakabe K, Osakabe Y, Toki S (2010) Site-directed mutagenesis in Arabidopsis using custom-designed zinc finger nucleases. Proc Natl Acad Sci 107:12034–12039
Pattanayak V, Ramirez CL, Joung JK, Liu DR (2011) Revealing off-target cleavage specificities of zinc-finger nucleases by in vitro selection. Nat Methods 8:765–770
Petolino JF, Worden A, Curlee K, Connell J, Moynahan TLS, Larsen C, Russell S (2010) Zinc finger nuclease-mediated transgene deletion. Plant Mol Biol 73:617–628
Puchta H, Fauser F (2014) Synthetic nucleases for genome engineering in plants: prospects for a bright future. Plant J 78:727–741
Schornack S, Moscou MJ, Ward ER, Horvath DM (2013) Engineering plant disease resistance based on TAL effectors. Annu Rev Phytopathol 51:383–406
Shan Q, Wang Y, Chen K, Liang Z, Li J, Zhang Y, Zhang K, Liu J, Voytas DF, Zheng X (2013a) Rapid and efficient gene modification in rice and Brachypodium using TALENs. Mol Plant 6:1365–1368
Shan Q, Wang Y, Li J, Zhang Y, Chen K, Liang Z, Zhang K, Liu J, Xi JJ, Qiu J-L (2013b) Targeted genome modification of crop plants using a CRISPR-Cas system. Nat Biotechnol 31:686–688
Shukla VK, Doyon Y, Miller JC, DeKelver RC, Moehle EA, Worden SE, Mitchell JC, Arnold NL, Gopalan S, Meng X (2009) Precise genome modification in the crop species Zea mays using zinc-finger nucleases. Nature 459:437–441
Sierro N, Battey JN, Ouadi S, Bakaher N, Bovet L, Willig A, Goepfert S, Peitsch MC, Ivanov NV (2014) The tobacco genome sequence and its comparison with those of tomato and potato. Nature Commun 5:3833
Sorek R, Lawrence CM, Wiedenheft B (2013) CRISPR-mediated adaptive immune systems in bacteria and archaea. Annu Rev Biochem 82:237–266
Sun Z, Li N, Huang G, Xu J, Pan Y, Wang Z, Tang Q, Song M, Wang X (2013) Site-specific gene targeting using transcription activator-like effector (TALE)-based nuclease in Brassica oleracea. J Integr Plant Biol 55:1092–1103
Tovkach A, Zeevi V, Tzfira T (2009) A toolbox and procedural notes for characterizing novel zinc finger nucleases for genome editing in plant cells. Plant J 57:747–757
Townsend JA, Wright DA, Winfrey RJ, Fu F, Maeder ML, Joung JK, Voytas DF (2009) High-frequency modification of plant genes using engineered zinc-finger nucleases. Nature 459:442–445
Upadhyay SK, Kumar J, Alok A, Tuli R (2013) RNA-guided genome editing for target gene mutations in wheat. G3-Genes Genomes Genet 3:2233–2238
Wang M, Wang G, Ji J, Wang J (2009) The effect of pds gene silencing on chloroplast pigment composition, thylakoid membrane structure and photosynthesis efficiency in tobacco plants. Plant Sci 177:222–226
Wendt T, Holm PB, Starker CG, Christian M, Voytas DF, Brinch-Pedersen H, Holme IB (2013) TAL effector nucleases induce mutations at a pre-selected location in the genome of primary barley transformants. Plant Mol Biol 83:279–285
Wright DA, Townsend JA, Winfrey RJ, Irwin PA, Rajagopal J, Lonosky PM, Hall BD, Jondle MD, Voytas DF (2005) High-frequency homologous recombination in plants mediated by zinc-finger nucleases. Plant J 44:693–705
Xiao A, Wang Z, Hu Y, Wu Y, Luo Z, Yang Z, Zu Y, Li W, Huang P, Tong X (2013) Chromosomal deletions and inversions mediated by TALENs and CRISPR/Cas in zebrafish. Nucleic Acids Res 41:e141
Xie K, Yang Y (2013) RNA-guided genome editing in plants using a CRISPR-Cas system. Mol Plant 6:1975–1983
Xie X, Wang G, Yang L, Cheng T, Gao J, Wu Y, Xia Q (2014) Cloning and characterization of a novel Nicotiana tabacum ABC transporter involved in shoot branching. Physiol Plant. doi:10.1111/ppl.12267
Xu R, Li H, Qin R, Wang L, Li L, Wei P, Yang J (2014) Gene targeting using the Agrobacterium tumefaciens-mediated CRISPR-Cas system in rice. Rice 7:5
Yoo S-D, Cho Y-H, Sheen J (2007) Arabidopsis mesophyll protoplasts: a versatile cell system for transient gene expression analysis. Nat Protoc 2:1565–1572
Zhang F, Maeder ML, Unger-Wallace E, Hoshaw JP, Reyon D, Christian M, Li X, Pierick CJ, Dobbs D, Peterson T (2010) High frequency targeted mutagenesis in Arabidopsis thaliana using zinc finger nucleases. Proc Natl Acad Sci 107:12028–12033
Zhang Y, Zhang F, Li X, Baller JA, Qi Y, Starker CG, Bogdanove AJ, Voytas DF (2013) Transcription activator-like effector nucleases enable efficient plant genome engineering. Plant Physiol 161:20–27
Zhang H, Zhang J, Wei P, Zhang B, Gou F, Feng Z, Mao Y, Yang L, Zhang H, Xu N (2014) The CRISPR/Cas9 system produces specific and homozygous targeted gene editing in rice in one generation. Plant Biotechnol J 12:797–807
Acknowledgments
This work was supported by grants from the following sources: the National Basic Research Program of China (No. 2012CB114600), the National Hi-Tech Research and Development Program of China (No. 2011AA100306), Fundamental Research Funds for the Central Universities (No. XDJK2013C043) and the Doctoral Fund of Southwest University (SWU112061).
Author information
Authors and Affiliations
Corresponding author
Additional information
Junping Gao and Genhong Wang have contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Gao, J., Wang, G., Ma, S. et al. CRISPR/Cas9-mediated targeted mutagenesis in Nicotiana tabacum . Plant Mol Biol 87, 99–110 (2015). https://doi.org/10.1007/s11103-014-0263-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11103-014-0263-0