Abstract
Purpose
The solvent effect on skin permeability is important for assessing the effectiveness and toxicological risk of new dermatological formulations in pharmaceuticals and cosmetics development. The solvent effect occurs by diverse mechanisms, which could be elucidated by efficient and reliable prediction models. However, such prediction models have been hampered by the small variety of permeants and mixture components archived in databases and by low predictive performance. Here, we propose a solution to both problems.
Methods
We first compiled a novel large database of 412 samples from 261 structurally diverse permeants and 31 solvents reported in the literature. The data were carefully screened to ensure their collection under consistent experimental conditions. To construct a high-performance predictive model, we then applied support vector regression (SVR) and random forest (RF) with greedy stepwise descriptor selection to our database. The models were internally and externally validated.
Results
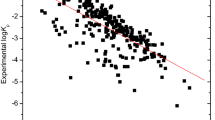
The SVR achieved higher performance statistics than RF. The (externally validated) determination coefficient, root mean square error, and mean absolute error of SVR were 0.899, 0.351, and 0.268, respectively. Moreover, because all descriptors are fully computational, our method can predict as-yet unsynthesized compounds.
Conclusion
Our high-performance prediction model offers an attractive alternative to permeability experiments for pharmaceutical and cosmetic candidate screening and optimizing skin-permeable topical formulations.







Similar content being viewed by others
Abbreviations
- ALOGP:
-
Ghose–Crippen octanol–water partition coefficient
- ANN:
-
Artificial neural network
- C d :
-
Chemical concentration in dose formulation
- J ss :
-
Steady state flux of the solute
- k p :
-
Permeability coefficient
- log P:
-
Octanol–water partition coefficient
- MAE:
-
Mean absolute error
- MW:
-
Molecular weight
- PCA:
-
Principal component analysis
- QSPR:
-
Quantitative structure–property relationship
- r 2 :
-
Determination coefficient
- RF:
-
Random forest
- RMSE:
-
Root mean square error
- SVR:
-
Support vector regression
References
Prausnitz MR, Mitragotri S, Langer R. Current status and future potential of transdermal drug delivery. Nat Rev Drug Discov. 2004;3(2):115–24.
Bartek MJ, LaBudde JA, Maibach HI. Skin permeability in vivo: comparison in rat, rabbit, pig and man. J Invest Dermatol. 1972;58(3):114–23.
Franz TJ. Percutaneous absorption on the relevance of in vitro data. J Invest Dermatol. 1975;64(3):190–5.
Zhang Q, Grice JE, Li P, Jepps OG, Wang GJ, Roberts MS. Skin solubility determines maximum transepidermal flux for similar size molecules. Pharm Res. 2009;26(8):1974–85.
Takeuchi H, Ishida M, Furuya A, Todo H, Urano H, Sugibayashi K. Influence of skin thickness on the in vitro permeabilities of drugs through Sprague–Dawley rat or Yucatan micropig skin. Biol Pharm Bull. 2012;35(2):192–202.
Karadzovska D, Riviere JE. Assessing vehicle effects on skin absorption using artificial membrane assays. Eur J Pharm Sci. 2013;50(5):569–76.
Chauhan P, Shakya M. Role of physicochemical properties in the estimation of skin permeability: in vitro data assessment by partial least-squares regression. SAR QSAR Environ Res. 2010;21(5–6):481–94.
Abraham MH, Martins F. Human skin permeation and partition: general linear free-energy relationship analyses. J Pharm Sci. 2004;93(6):1508–23.
Buchwald P, Bodor N. A simple, predictive, structure-based skin permeability model. J Pharm Pharmacol. 2001;53(8):1087–98.
Cronin MT, Dearden JC, Moss GP, Murray-Dickson G. Investigation of the mechanism of flux across human skin in vitro by quantitative structure-permeability relationships. Eur J Pharm Sci. 1999;7(4):325–30.
Potts RO, Guy RH. Predicting skin permeability. Pharm Res. 1992;9(5):663–9.
Baba H, Takahara JI, Mamitsuka H. In silico predictions of human skin permeability using nonlinear quantitative structure–property relationship models. Pharm Res. 2015. doi:10.1007/s11095-015-1629-y.
Khajeh A, Modarress H. Linear and nonlinear quantitative structure–property relationship modelling of skin permeability. SAR QSAR Environ Res. 2014;25(1):35–50.
Neely BJ, Madihally SV, Robinson Jr RL, Gasem KA. Nonlinear quantitative structure–property relationship modeling of skin permeation coefficient. J Pharm Sci. 2009;98(11):4069–84.
Baert B, Deconinck E, Van Gele M, Slodicka M, Stoppie P, Bodé S, et al. Transdermal penetration behaviour of drugs: CART-clustering, QSPR and selection of model compounds. Bioorg Med Chem. 2007;15(22):6943–55.
Katritzky AR, Dobchev DA, Fara DC, Hür E, Tämm K, Kurunczi L, et al. Skin permeation rate as a function of chemical structure. J Med Chem. 2006;49(11):3305–14.
Neumann D, Kohlbacher O, Merkwirth C, Lengauer T. A fully computational model for predicting percutaneous drug absorption. J Chem Inf Model. 2006;46(1):424–9.
Lim CW, Fujiwara S, Yamashita F, Hashida M. Prediction of human skin permeability using a combination of molecular orbital calculations and artificial neural network. Biol Pharm Bull. 2002;25(3):361–6.
Smith EW, Maibach HI, editors. Percutaneous penetration enhancers, second ed. New York: CRC Press; 2005.
Barry BW. Breaching the skin’s barrier to drugs. Nat Biotechnol. 2004;22(2):165–7.
Lane ME. Skin penetration enhancers. Int J Pharm. 2013;447(1–2):12–21.
Watkinson RM, Guy RH, Hadgraft J, Lane ME. Optimisation of cosolvent concentration for topical drug delivery - II: influence of propylene glycol on ibuprofen permeation. Skin Pharmacol Physiol. 2009;22(4):225–30.
Roy SD, Manoukian E. Transdermal delivery of ketorolac tromethamine: permeation enhancement, device design, and pharmacokinetics in healthy humans. J Pharm Sci. 1995;84(10):1190–6.
Sathyan G, Ritschel WA, Hussain AS. Transdermal delivery of tacrine: I: Identification of a suitable delivery vehicle. Int J Pharm. 1995;114(1):75–83.
Goldberg-Cettina M, Liu P, Nightingale J, Kurihara-Bergstrom T. Enhanced transdermal delivery of estradiol in vitro using binary vehicles of isopropyl myristate and short-chain alkanols. Int J Pharm. 1995;114(2):237–45.
Pardo A, Shiri Y, Cohen S. Percutaneous absorption of physostigmine: optimization of delivery from a binary solvent by thermodynamic control. J Pharm Sci. 1990;79(7):573–8.
Riviere JE, Brooks JD. Prediction of dermal absorption from complex chemical mixtures: incorporation of vehicle effects and interactions into a QSPR framework. SAR QSAR Environ Res. 2007;18(1–2):31–44.
Riviere JE, Brooks JD. Predicting skin permeability from complex chemical mixtures: dependency of quantitative structure permeation relationships on biology of skin model used. Toxicol Sci. 2011;119(1):224–32.
Ghafourian T, Samaras EG, Brooks JD, Riviere JE. Modelling the effect of mixture components on permeation through skin. Int J Pharm. 2010;398(1–2):28–32.
Ghafourian T, Samaras EG, Brooks JD, Riviere JE. Validated models for predicting skin penetration from different vehicles. Eur J Pharm Sci. 2010;41(5):612–6.
Netzeva TI, Worth A, Aldenberg T, Benigni R, Cronin MT, Gramatica P, et al. Current status of methods for defining the applicability domain of (quantitative) structure-activity relationships. The report and recommendations of ECVAM Workshop 52. Altern Lab Anim. 2005;33(2):155–73.
Moss GP, Sun Y, Prapopoulou M, Davey N, Adams R, Pugh WJ, et al. The application of Gaussian processes in the prediction of percutaneous absorption. J Pharm Pharmacol. 2009;61(9):1147–53.
Patel J. Science of the science, drug discovery and artificial neural networks. Curr Drug Discov Technol. 2013;10(1):2–7.
Smola AJ, Schölkopf B. A tutorial on support vector regression. Stat Comput. 2004;14(3):199–222.
Breiman L. Random forests. Mach Learn. 2001;45(1):5–32.
Monte-Moreno E. Non-invasive estimate of blood glucose and blood pressure from a photoplethysmograph by means of machine learning techniques. Artif Intell Med. 2011;53(2):127–38.
Yap CW, Li ZR, Chen YZ. Quantitative structure-pharmacokinetic relationships for drug clearance by using statistical learning methods. J Mol Graph Model. 2006;24(5):383–95.
El-Sebakhy EA. Forecasting PVT properties of crude oil systems based on support vector machines modeling scheme. J Petrol Sci Eng. 2009;64(1–4):25–34.
Bharathidason S, Venkataeswaran CJ. Improving classification accuracy based on random forest model with uncorrelated high performing trees. Int J Comput Appl. 2014;101(13):26–30.
Blank IH, McAuliffe DJ. Penetration of benzene through human skin. J Investig Dermatol. 1985;85(6):522–6.
Burkert U, Norman LA. Molecular mechanics, ACS monograph 177. Washington, DC: American Chemical Society; 1982.
Stewart JJ. Optimization of parameters for semiempirical methods VI: more modifications to the NDDO approximations and re-optimization of parameters. J Mol Model. 2013;19(1):1–32.
Law V, Knox C, Djoumbou Y, Jewison T, Guo AC, Liu Y, et al. DrugBank 4.0: shedding new light on drug metabolism. Nucleic Acids Res. 2014;42:D1091–7.
We have obtained structures of 6475 compounds as approved drugs in DrugBank: http://www.drugbank.ca/downloads#structures/.
Meyer D, Dimitriadou E, Hornik K, Weingessel A, Leisch F. e1071: Misc Functions of the Department of Statistics (e1071), TU Wien. R package version 1.6-2. 2014. Available from http://CRAN.R-project.org/package=e1071/.
Svetnik V, Liaw A, Tong C, Culberson JC, Sheridan RP, Feuston BP. Random forest: a classification and regression tool for compound classification and QSAR modeling. J Chem Inf Comput Sci. 2003;43(6):1947–58.
Liaw A, Wiener M. Classification and regression by random Forest. R News. 2002;2:18–22.
Shao J. Linear model selection by cross-validation. J Am Stat Assoc. 1993;88(442):486–94.
Zhang P. Model selection via multifold cross validation. Ann Stat. 1993;21(1):299–313.
Burman P. A Comparative study of ordinary cross-validation, v-fold cross-validation and the repeated learning-testing methods. Biometrika. 1989;76(3):503–14.
Golbraikh A, Tropsha A. Beware of q2! J Mol Graph Model. 2002;20(4):269–76.
Roy PP, Roy K. On some aspects of variable selection for partial least squares regression models. QSAR Comb Sci. 2008;27(3):302–13.
Consonni V, Ballabio D, Todeschini R. Comments on the definition of the Q2 parameter for QSAR validation. J Chem Inf Model. 2009;49(7):1669–78.
Ojha PK, Mitra I, Das RN, Roy K. Further exploring rm2 metrics for validation of QSPR models. Chemom Intell Lab Syst. 2011;107(1):194–205.
Chirico N, Gramatica P. Real external predictivity of QSAR models: how to evaluate it? Comparison of different validation criteria and proposal of using the concordance correlation coefficient. J Chem Inf Model. 2011;51(9):2320–35.
Chirico N, Gramatica P. Real external predictivity of QSAR models.Part 2.New intercomparable thresholds for different validation criteria and the need for scatter plot inspection. J Chem Inf Model. 2012;52(8):2044–58.
Roy K, Mitra I, Kar S, Ojha PK, Das RN, Kabir H. Comparative studies on some metrics for external validation of QSPR models. J Chem Inf Model. 2012;52(2):396–408.
Funar-Timofei S, Iliescu S, Suzuki T. Correlations of limiting oxygen index with structural polyphosphoester features by QSPR approaches. Struct Chem. 2014;25(6):1847–63.
Fornberg B, Sloan DM. A review of pseudospectral methods for solving partial differential equations. Acta Numer. 1994;3:203–67.
Bos JD, Meinardi MM. The 500 Dalton rule for the skin penetration of chemical compounds and drugs. Exp Dermatol. 2000;9(3):165–9.
Yano T, Nakagawa A, Tsuji M, Noda K. Skin permeability of various non-steroidal anti-inflammatory drugs in man. Life Sci. 1986;39(12):1043–50.
Kasting GB, Smith RL, Cooper ER. Effect of lipid solubility and molecular size on percutaneous absorption. In: Shroot B, Schaefer H, editors. Skin pharmacokinetics. Basel: Kargar; 1987. p. 138–53.
Boonen J, Veryser L, Taevernier L, Roche N, Peremans K, Burvenich C, et al. Risk evaluation of impurities in topical excipients: the acetol case. J Pharm Anal. 2014;4(5):303–15.
Inselberg A. Parallel coordinates: a tool for visualizing multidimensional geometry. New York: Springer; 2009.
Shahlaei M. Descriptor selection methods in quantitative structure-activity relationship studies: a review study. Chem Rev. 2013;113(10):8093–103.
Geinoz S, Guy RH, Testa B, Carrupt PA. Quantitative structure-permeation relationships (QSPeRs) to predict skin permeation: A critical evaluation. Pharm Res. 2004;21(1):83–92.
Michaels AS, Chandrasekaran SK, Shaw JE. Drug permeation through human skin: Theory and invitro experimental measurement. AIChE J. 1975;21(5):985–96.
Verhaar HJM, van Leeuwen CJ, Hermens JLM. Classifying environmental pollutants 1: Structure-activity relationships for prediction of aquatic toxicity. Chemosphere. 1992;25(4):471–91.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Baba, H., Takahara, Ji., Yamashita, F. et al. Modeling and Prediction of Solvent Effect on Human Skin Permeability using Support Vector Regression and Random Forest. Pharm Res 32, 3604–3617 (2015). https://doi.org/10.1007/s11095-015-1720-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11095-015-1720-4