Abstract
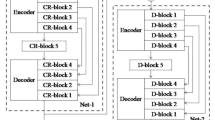
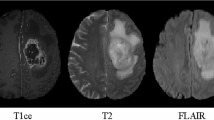
Accurate tumor delineation in medical images is of great importance in guiding radiotherapy. In nasopharyngeal carcinoma (NPC), due to its high variability, low contrast and discontinuous boundaries in magnetic resonance images (MRI), the margin of the tumor is especially difficult to be identified, making the radiotherapy planning a more challenging problem. The objective of this paper is to develop an automatic segmentation method of NPC in MRI for radiosurgery applications. To this end, we present to segment NPC using a deep convolutional neural network. Specifically, to obtain spatial consistency as well as accurate feature details for segmentation, multiple convolution kernel sizes are employed. The network contains a large number of trainable parameters which capture the relationship between the MRI intensity images and the corresponding label maps. When trained on subjects with pre-labeled MRI, the network can estimate the label class of each voxel for the testing subject which is only given the intensity image. To demonstrate the segmentation performance, we carry on our method on the T1-weighted images of 15 NPC patients, and compare the segmentation results against the radiologist’s reference outline. Experimental results show that the proposed method outperforms the traditional hand-crafted features based segmentation methods. The presented method in this paper could be useful for NPC diagnosis and helpful for guiding radiotherapy.







Similar content being viewed by others
References
Stewart BWKP, Wild CP (2014) World cancer report
Chang ET, Adami HO (2006) The enigmatic epidemiology of nasopharyngeal carcinoma. Cancer Epidemiol Biomark Prev 15(10):1765–1777
Parkin D, Bray F, Ferlay J, Pisani P (2005) Global cancer statistics. CA Cancer J Clin 55:74–108
Bertolino A, Frye M, Callicott JH, Mattay VS, Rakow R, Shelton-Repella J et al (2003) Neuronal pathology in the hippocampal area of patients with bipolar disorder: a study with proton magnetic resonance spectroscopic imaging. Biol Psychiatry 53(10):1177–1194
Lodi R, Tonon PC, Manners D, Capellari S, Strammiello R, Rinaldi R et al (2009) Magnetic resonance diagnostic markers in clinically sporadic prion disease: a combined brain magnetic resonance imaging and spectroscopy study. Brain 132(Pt10):2669–79
Satoh T, Omi M, Nabeshima M, Onoda K, Date I (2009) Severity analysis of neurovascular contact in patients with trigeminal neuralgia: assessment with the inner view of the 3D MR cisternogram and angiogram fusion imaging. Am J Neuroradiol 30(3):603–7
Wang Y, Zhang P, An L, Ma G, Kang J, Wu X et al (2016) Predicting standard-dose pet image from low-dose pet and multimodal MR images using mapping-based sparse representation. Phys Med Biol 61(2):791–812
Huang KW, Zhao ZY, Gong Q, Zha J, Chen L, Yang R (2015) Nasopharyngeal carcinoma segmentation via HMRF-EM with maximum entropy. Eng Med Biol Soc 2015:2968
Gao Y, Zhang H, Zhao X, Yan S (2017) Event classification in microblogs via social tracking. ACM Trans Intell Syst Technol (TIST) 8(3):35
Zu C, Wang Z, Zhang D, Liang P, Shi Y, Shen D, Wu G (2017) Robust multi-atlas label propagation by deep sparse representation. Pattern Recogn 63:511–517
Gao Y, Wang M, Tao D, Ji R, Dai Q (2012) 3-D object retrieval and recognition with hypergraph analysis. IEEE Trans Image Process 21(9):4290–4303
Chen L, Chen L, Cui L, Cui L, Huang R, Huang R, Ren Z (2016) Bio-inspired neural network with application to license plate recognition: hysteretic ELM approach. Assembly Autom 36(2):172–178
Shi J, Zheng X, Li Y, Zhang Q, Ying S (2017) Multimodal neuroimaging feature learning with multimodal stacked deep polynomial networks for diagnosis of Alzheimer’s disease. IEEE J Biomed Health Inform 22(1):173–183
Shi J, Wu J, Li Y, Zhang Q, Ying S (2017) Histopathological image classification with color pattern random binary hashing based PCANet and matrix-form classifier. IEEE J Biomed Health Inform 21(5):1327–1337
Ying S, Wen Z, Shi J, Peng Y, Peng J, Qiao H (2017) Manifold preserving: an intrinsic approach for semisupervised distance metric learning. IEEE Trans Neural Netw Learn Syst. https://doi.org/10.1109/TNNLS.2017.2691005
Ramirez-Quintana JA, Chacon-Murguia MI (2015) An adaptive unsupervised neural network based on perceptual mechanism for dynamic object detection in videos with real scenarios. Neural Process Lett 42(3):665–689
Brendel M, Roska T, Bártfai G (2002) Gradient computation of continuous-time cellular neural/nonlinear networks with linear templates via the CNN universal machine. Neural Process Lett 16(2):111–120
Pereira S, Pinto A, Alves V, Silva CA (2016) Brain tumor segmentation using convolutional neural networks in MRI images. IEEE Trans Med Imaging 35(5):1240–1251
Zhang Y, Zhao D, Sun J, Zou G, Li W (2016) Adaptive convolutional neural network and its application in face recognition. Neural Process Lett 43(2):389–399
Zhang W, Li R, Deng H, Wang L, Lin W, Ji S et al (2015) Deep convolutional neural networks for multi-modality isointense infant brain image segmentation. Neuroimage 108:214–224
Brebisson AD, Montana G (2015) Deep neural networks for anatomical brain segmentation. In: CVPR bioimage computing workshop, vol 35, pp 20–28
Zikic D, Ioannou Y, Brown M, Criminisi A (2014) Segmentation of brain tumor tissues with convolutional neural networks. In: MICCAI workshop on multimodal brain tumor segmentation challenge, pp 36–39
Rao V, Sharifi M, Jaiswal A (2015) Brain tumor segmentation with deep learning. In: MICCAI multimodal brain tumor segmentation challenge, pp 56–59
Fitton I, Cornelissen SAP, Duppen JC, Steenbakkers RJHM, Peeters STH, Hoebers FJP et al (2011) Semi- automatic delineation using weighted CT-MRI registered images for radiotherapy of nasopharyngeal cancer. Med Phys 38(8):4662–4666
Huang KW, Zhao ZY, Gong Q, Zha J, Chen L, Yang R (2015) Nasopharyngeal carcinoma segmentation via HMRF-EM with maximum entropy. In: Engineering in Medicine and Biology Society, vol 2015, pp 2968
Lee FK, Yeung DK, King AD, Leung SF, Ahuja A (2005) Segmentation of nasopharyngeal carcinoma (NPC) lesions in MR images. Int J Radiat Oncol Biol Phys 61(2):608–620
Ritthipravat P, Tatanun C, Bhongmakapat T, Tuntiyatorn L (2008) Automatic segmentation of nasopharyngeal carcinoma from CT images. Int Conf Biomed Eng Inform 2:18–22
Chanapai W, Bhongmakapat T, Tuntiyatorn L, Ritthipravat P (2012) Nasopharyngeal carcinoma segmentation using a region growing technique. Int J Comput Assist Radiol Surg 7(3):413–422
Hong R, Ye S (2014) Segmentation of nasopharyngeal MR medical image base on improved region growing. J Fuzhou Univ (Nat Sci Edn) 42(5):683–688
Zhang J, Ma KK, Meng HE, Chong V (2004) Tumor segmentation from magnetic resonance imaging by learning via one-class support vector machine. In: International workshop on advanced image technology, pp 207–211
Zhou J, Chan KL, Xu P, Chong VFH (2006) Nasopharyngeal carcinoma lesion segmentation from MR images by support vector machine. In: IEEE international symposium on biomedical imaging: nano to macro, pp 1364–1367
Zhou J, Chong V, Lim TK, Houng J (2002) MRI tumor segmentation for nasopharyngeal carcinoma using knowledge-based fuzzy clustering. Int J Inf Technol 8(2):36–45
Huang W, Chan KL, Zhou J (2013) Region-based nasopharyngeal carcinoma lesion segmentation from MRI using clustering-and classification-based methods with learning. J Digit Imaging 26(3):472–482
Tustison NJ, Avants BB, Cook PA, Zheng Y, Egan A, Yushkevich PA et al (2010) N4ITK: improved N3 bias correction. IEEE Trans Med Imaging 29(6):1310–20
Acknowledgements
This work was supported in part by NSFC61701324, Science and Technology Department of Sichuan Province 2016JZ0014 and Open Fund Project of Fujian Provincial Key Laboratory of Information Processing and Intelligent Control (Minjiang University) (No. MJUKF201715).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Wang, Y., Zu, C., Hu, G. et al. Automatic Tumor Segmentation with Deep Convolutional Neural Networks for Radiotherapy Applications. Neural Process Lett 48, 1323–1334 (2018). https://doi.org/10.1007/s11063-017-9759-3
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11063-017-9759-3