Abstract
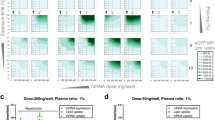
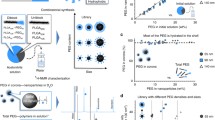
It is hypothesized that plasma proteins bind nanoparticles in vivo as they do in vitro, forming a protein corona. The resulting decorated nanoparticle surface could potentially alter nanoparticle pharmacokinetics, efficacy, and toxicity in vivo. A subset of the in vitro corona are high-density lipoproteins (HDLs). Since HDLs vary in patients based on diet, weight, and genetics, it is crucial to determine the affinity of HDLs for nanoparticles to generate a predictive model, which would provide information on the extent of HDL decoration on nanoparticles in the blood. Experiments that determined equilibrium affinities of HDLs for nanoparticles utilized isolated HDLs or HDL structural protein components such as ApoA-I. Thus, the effects of whole plasma on HDL-nanoparticle equilibrium binding are unclear. It is possible that competition from other plasma proteins for the nanoparticle surface could drastically change the affinity of HDLs for nanoparticles both in vitro and in vivo. Here, we determined effective equilibrium binding constants of Kdeff = 3.1 ± 0.7 μM, 1.2 ± 0.4 μM, and 2.0 ± 0.4 μM for polystyrene (PS), PS-COOH, and PS-NH2 nanospheres for ApoA-I, the main structural component of HDLs in whole mouse plasma. In comparison, binding constants were Kd = 400 nM, 900 nM, and 25 nM for PS, PS-COOH, and PS-NH2 nanospheres and HDLs isolated from mouse plasma. We utilized a binding model that is characterized by a nanoparticle with multiple identical and independent binding sites for HDLs. Our data show that HDL binding to nanoparticles could play a significant role in nanoparticle behavior in the vasculature of mammals.





Similar content being viewed by others
References
Bisswanger H (2008) Multiple equilibria. In: Enzyme kinetics: principles and methods, 2nd edn. Wiley-VCH Verlag GmbH & Co. KGaA, Weinheim, pp 13–20
Cedervall T, Lynch I, Foy M, Berggård T, Donnelly SC, Cagney G, Linse S, Dawson KA (2007) Detailed identification of plasma proteins adsorbed on copolymer nanoparticles. Angew Chem Int Ed 46:5754–5756
Cukalevski R, Lundqvist M, Oslakovic C, Dahlbäck B, Linse S, Cedervall T (2011) Structural changes in apolipoproteins bound to nanoparticles. Langmuir 27:14360–14369
Daniels KG, Suo Y, Oas TG (2015) Conformational kinetics reveals affinities of protein conformational states. Proc Natl Acad Sci USA 112:9352–9357
Davidson WS (2018) LDL Proteome Watch. http://homepages.uc.edu/~davidswm/LDLproteome.html. Accessed 17 July 2018
Dell’Orco D, Lundqvist M, Oslakovic C, Cedervall T, Linse S (2010) Modeling the time evolution of the nanoparticle-protein Corona in a body fluid. PLoS One 5:e10949
Deng ZJ, Mortimer G, Schiller T, Musumeci A, Martin D, Minchin RF (2009) Differential plasma protein binding to metal oxide nanoparticles. Nanotechnology 20:1–9
Dobrovolskaia MA, Patri AK, Zheng J, Clogston JD, Ayub N, Aggarwal P, Neun BW, Hall JB, McNeil SE (2009) Interaction of colloidal gold nanoparticles with human blood: effects on particle size and analysis of plasma protein binding profiles. Nanomedicine 5:106–117
Duan Y, Liu Y, Shen W, Zhong W (2017) Fluorescamine labeling for assessment of protein conformational change and binding affinity in protein−nanoparticle interaction. Anal Chem 89:12160–12167
Fagerberg L, Hallstrom BM, Oksvold P et al (2014) Analysis of the human tissue-specific expression by genome-wide integration of transcriptomics and antibody-based proteomics. Mol Cell Proteomics 13:397–406
Feingold KR, Grunfeld C (2019) Introduction to lipids and lipoproteins. In: Feingold KR, Anawalt B, Boyce A et al (eds) Endotext. MDText.com, Inc, South Dartmouth 2000-. https://www.ncbi.nlm.nih.gov/books/NBK305896/. Accessed 15 July 2019
Freire E, Mayorga OL, Straume M (1990) Isothermal titration calorimetry. Anal Chem 62:950A–959A
Göppert TM, Müller RH (2005) Protein adsorption patterns on poloxamer- and poloxamine-stabilized solid lipid nanoparticles (SLN). Eur J Pharm Biopharm 60:361–372
Gordon SM, Li H, Zhu X, Shah AS, Lu LJ, Davidson WS (2015) A comparison of the mouse and human lipoproteome: suitability of the mouse model for studies of human lipoproteins. J Proteome Res 14:2686–2695
Gossmann R, Fahrländer E, Hummel M, Mulac D, Brockmeyer J, Langer K (2015) Comparative examination of adsorption of serum proteins on HSA- and PLGA-based nanoparticles using SDS–PAGE and LC–MS. Eur J Pharm Biopharm 93:80–87
Hutchins PM, Ronsein GE, Monette JS, Pamir N, Wimberger J, He Y, Anantharamaiah GM, Kim DS, Ranchalis JE, Jarvik GP, Vaisar T, Heinecke JW (2014) Quantification of HDL particle concentration by calibrated ion mobility analysis. Clin Chem 60:1393–1401
Kono M, Okumura Y, Tanaka M, Nguyen D, Dhanasekaran P, Lund-Katz S, Phillips MC, Saito H (2008) Conformational flexibility of the N-terminal domain of apolipoprotein A-I bound to spherical lipid particles. Biochemistry 47:11340–11347
Lauffenburger DA, Linderman JJ (1996) Cell surface receptor binding models. In: Receptors: models for binding, trafficking, and signaling. Oxford University Press, Oxford, pp 19–26
Lundqvist M, Stigler J, Elia G, Lynch I, Cebervall T, Dawson KA (2008) Nanoparticle size and surface properties determine the protein corona with possible implications for biological impacts. Proc Natl Acad Sci U S A 105:14265–14270
Müller J, Prozeller D, Ghazaryan A, Kokkinopoulou M, Mailänder V, Morsbach S, Landfester K (2018) Beyond the protein corona—lipids matter for biological response of nanocarriers. Acta Biomater 71:420–431
Olbrich C, Gessner A, Schröder W, Kayser O, Müller RH (2004) Lipid–drug conjugate nanoparticles of the hydrophilic drug diminazene-cytotoxicity testing and mouse serum adsorption. J Control Release 96:425–435
Pamir N, Pan C, Plubell DL, Hutchins PM, Tang C, Wimberger J, Irwin A, Vallim TQA, Heinecke JW, Lusis AJ (2019) Genetic control of the mouse HDL proteome defines HDL traits, function, and heterogeneity. J Lipid Res 60:594–608
Pollard TD (2010) A Guide to Simple and Informative Binding Assays. Mol Biol Cell 21:4061–4067
Schneider CA, Rasband WS, Eliceiri KW (2012) NIH image to ImageJ: 25 years of image analysis. Nat Methods 9:671–675
Shah AS, Tan L, Long JL, Davidson WS (2013) Proteomic diversity of high-density lipoproteins: our emerging understanding of its importance in lipid transport and beyond. J Lipid Res 54:2575–2585
Shang J, Gao X (2014) Nanoparticle counting: towards accurate determination of the molar concentration. Chem Soc Rev 43:7267–7278
Shevchenko A, Tomas H, Havli J, Olsen JV, Mann M (2006) In-gel digestion for mass spectrometric characterization of proteins and proteomes. Nat Protoc 1:2856–2860
Tenzer S, Doctor D, Kuharev J, Musyanovych A et al (2013) Rapid formation of plasma protein corona critically affects nanoparticle pathophysiology. Nat Nanotechnol 8:772–781
Walkey CD, Chan WCW (2012) Understanding and controlling the interaction of nanomaterials with proteins in a physiological environment. Chem Soc Rev 41:2780–2799
Winzen S, Schoettler S, Baier G, Rosenauer C, Mailaender V, Landfester K, Mohr K (2015) Complementary analysis of the hard and soft protein corona: sample preparation critically effects corona composition. Nanoscale 7:2992–3001
Acknowledgements
We thank Dr. Eric Tague for mass spectrometry. We also thank Dr. Edward Wright and Liang Fang for their support with the isothermal titration calorimetry and gel filtration chromatography column, respectively.
Funding
This work was supported by the National Institute of General Medical Sciences R15GM116037.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(DOCX 24.4 mb)
Rights and permissions
About this article
Cite this article
Anozie, U.C., Quigley, K.J., Prescott, A. et al. Equilibrium binding of isolated and in-plasma high-density lipoproteins (HDLs) to polystyrene nanoparticles. J Nanopart Res 22, 223 (2020). https://doi.org/10.1007/s11051-020-04953-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11051-020-04953-0