Abstract
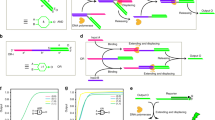
A functional machine is not only an assembly of parts, but also an assembly of processes. The processing of each part must obey laws that respect to the property of this part. For example, building any kind of computer entails selecting appropriate components and assembling their properties to function in computation. Here, we describe computation using a DNA strand as the basic unit and we have used this unit to achieve the function of multiplication. We exploit the phenomenon of DNA hybridization, in which each strand can represent two individual units that can pair to form a single unit. We represent the numbers we multiply in binary, with different lengths representing each digit present in the number. In principle, all combinations of the numbers will be present in solution. Following hybridization, there is present a collection of duplex molecules that are tailed by single-stranded ends. These intermediates are converted to fully duplex molecules by filling in the ends with DNA polymerase. The lengths that are present represent the digits that are present, and they may be separated by denaturing PAGE. The results give a series of bands for each power of two. The number of bands in the size domain for a particular power of two is converted to binary and the sum of all present bands is then added together. Experimentally, the result of this process always yields the correct answer.
Similar content being viewed by others
References
Adleman LM (1994) Molecular computation of solutions to combinatorial problems. Science 266: 1021–1024
Braich RS, Chelyapov N, Johnson C, Rothemund PWK, and Adleman L (2002) Solution of a 20-Variable 3-SAT problem on a DNA computer. Science 296: 499–502
Egholm M, Buchardt O, Christensen L, Behrens C, Freier SM, Driver DA, Berg RH, Kim SK, Norden B, Nielsen PE (1993) PNA hybridizes to complementary oligonucleotides obeying the Watson-Crick hydrogen-bonding rules. Nature 365: 566–568
Faulhammer D, Cukras AR, Lipton RJ, Landweber LF (2000) Molecular computation: RNA solutions to chess problems. Proceedings of the National Academy of Sciences (USA) 97: 1385–1389
Frank R, Heikens W, Heisterberg-Moutsis G, Blöcker H (1983) A new general approach for the simultaneous chemical synthesis of large numbers of oligonucleotides: segmental solid supports. Nucleic Acids Research 11: 4366–4377
Guarnieri F, Fliss M, Bancroft C (1996) Making DNA add. Science 273: 220–223
Horn T, Urdea MS (1989) Forks and combs and DNA – the synthesis of branched oligodeoxyribonucleotides. Nucleic Acids Research 17: 6959–6967
Lipton RJ (1995) DNA solution of hard computational problems. Science 268: 542–545
Liu Q, Wang L, Frutos AG, Condon AE, Corn RM, Smith LM (2000) DNA computing on surfaces. Nature 403: 175–179
Sakamoto K, Gouzu H, Komiya K, Kiga D, Yokoyama S, Yokomori T, Hagiya M (2000) Molecular computation by DNA hairpin formation. Science 288: 1223–1226
Seeman NC (1990) De Novo design of sequences for nucleic acid structural engineering. Journal of Biomolecular Structure & Dynamics 8: 573–581
Acknowledgements
This research has been supported by grants GM-29554 from NIGMS, grants, EIA-0086015, CCF-0432009, CCF-0523290, and CTS-0548774 from the NSF, 48681-EL from ARO and NTI-001 from Nanoscience Technologies, Inc. to N.C.S.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wu, G., Seeman, N.C. Multiplying with DNA. Nat Comput 5, 427–441 (2006). https://doi.org/10.1007/s11047-006-9006-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11047-006-9006-8