Abstract
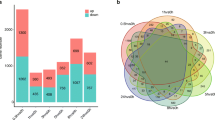
To identify genes that are differentially expressed in tobacco in response to environmental changes and to decipher the mechanisms by which aromatic carotenoids are formed in tobacco, an Agilent Tobacco Gene Expression microarray was adapted for transcriptome comparison of tobacco leaves derived from three cultivated regions of China, Kaiyang (KY), Weining (WN) and Tianzhu (TZ). A total of 1,005 genes were differentially expressed between leaves derived from KY and TZ, 733 between KY and WN, and 517 between TZ and WN. Genes that were upregulated in leaves from WN and TZ tended to be involved in secondary metabolism pathways, and included several carotenoid pathway genes, e.g., NtPYS, NtPDS, and NtLCYE, whereas those that were down-regulated tended to be involved in the response to temperature and light. The expression of 10 differentially expressed genes (DEGs) was evaluated by real-time quantitative polymerase chain reaction (qRT-PCR) and found to be consistent with the microarray data. Gene Ontology and MapMan analyses indicate that the genes that were differentially expressed among the three cultivated regions were associated with the light reaction of photosystem II, response to stimuli, and secondary metabolism. High-performance liquid chromatography (HPLC) analysis showed that leaves derived from KY had the lowest levels of lutein, β-carotene, and neoxanthin, whereas the total carotenoid content in leaves from TZ was greatest, a finding that could well be explained by the expression patterns of DEGs in the carotenoid pathway. These results may help elucidate the molecular mechanisms underlying environmental adaptation and accumulation of aroma compounds in tobacco.






Similar content being viewed by others
Abbreviations
- a.s.l.:
-
Above sea level
- DEG:
-
Differentially expressed gene
References
Cramer GR, Urano K, Delrot S, Pezzotti M, Shinozaki K (2011) Effects of abiotic stress on plants: a systems biology perspective. BMC Plant Biol 11:163
Wang Y, Frei M (2011) Stressed food-The impact of abiotic environmental stresses on crop quality. Agric Ecosyst Environ 141:271–286
Wahlberg I (2002) Carotenoid-derived aroma compounds in tobacco. In: Winterhalter P, Rouseff RL (eds) Carotenoid-derived aroma compounds, vol 802. American Chemical Society, Washington, pp 131–144
Liu P, Wang X, Wang J, Yan Q, Shi G, Yang T (2010) Changes of aroma constituents of different flue-cured tobacco genotypes in different ecoregions. Acta Agric Zhejiangensis 22:239–242 (in Chinese with English Abstract)
Shi JL, Yang HQ, Song CM, Deng JH, Pang T (2011) Comparison on the main volatile aroma components of flue-cured tobacco from different ecological regions in Yunnan Province. J Yunnan Agric Univ 26:790–795 (in Chinese with English Abstract)
Wang N, Li Z, Wang D, Xu Z, Zhou H, Zhu X (2009) Preliminary study on principal aroma and flavor constituents of flue-cured tobacco in China. Chin Tobacco Sci 30(3):1–6 (in Chinese)
Edwards KD, Bombarely A, Story GW, Allen F, Mueller LA, Coates SA, Jones L (2010) TobEA: an atlas of tobacco gene expression from seed to senescence. BMC Genomics 11:142. doi:10.1186/1471-2164-11-142
Lu K, Chai Y, Zhang K, Wang R, Chen L, Lei B, Lu J, Xu X, Li J (2008) Cloning and characterization of phosphorus starvation inducible Brassica napus PURPLE ACID PHOSPHATASE12 gene family, and imprinting of a recently evolved MITE-minisatellite twin structure. Theor Appl Genet 117:963–975
Guo Y, Guo H, Zhang L, Xie H, Zhao X, Wang F, Li Z, Wang Y, Ma S, Tao J (2005) Genomic analysis of anti-hepatitis B virus (HBV) activity by small interfering RNA and lamivudine in stable HBV-producing cells. J Virol 79:14392–14403
Yu L, Guo N, Meng R, Liu B, Tang X, Jin J, Cui Y, Deng X (2010) Allicin-induced global gene expression profile of Saccharomyces cerevisiae. Appl Microbiol Biotechnol 88:219–229. doi:10.1007/s00253-010-2709-x
Yang YH, Dudoit S, Luu P, Lin DM, Peng V, Ngai J, Speed TP (2002) Normalization for cDNA microarray data: a robust composite method addressing single and multiple slide systematic variation. Nucleic Acids Res 30:e15
Patterson TA, Lobenhofer EK, Fulmer-Smentek SB, Collins PJ, Chu TM, Bao W, Fang H, Kawasaki ES, Hager J, Tikhonova IR (2006) Performance comparison of one-color and two-color platforms within the MicroArray Quality Control (MAQC) project. Nat Biotechnol 24:1140–1150
Du Z, Zhou X, Ling Y, Zhang Z, Su Z (2010) agriGO: a GO analysis toolkit for the agricultural community. Nucleic Acids Res 38:W64–W70. doi:10.1093/nar/gkq310
Ye J, Fang L, Zheng H, Zhang Y, Chen J, Zhang Z, Wang J, Li S, Li R, Bolund L (2006) WEGO: a web tool for plotting GO annotations. Nucleic Acids Res 34:W293–W297
Thimm O, Blasing O, Gibon Y, Nagel A, Meyer S, Kruger P, Selbig J, Muller LA, Rhee SY, Stitt M (2004) MAPMAN: a user-driven tool to display genomics data sets onto diagrams of metabolic pathways and other biological processes. Plant J 37:914–939
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods 25:402–408. doi:10.1006/meth.2001.1262
Drummond AJ, Ashton B, Cheung M, Heled J, Kearse M, Moir R, Stones-Havas S, Thierer T, Wilson A (2009) Geneious v4.8. http://www.geneious.com/
Edgar R, Domrachev M, Lash AE (2002) Gene Expression Omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res 30:207–210
Benjamini Y, Yekutieli D (2001) The control of the false discovery rate in multiple testing under dependency. Ann Stat 29:1165–1188
Dijk JP, Leifert C, Barros E, Kok EJ (2010) Gene expression profiling for food safety assessment: examples in potato and maize. Regul Toxicol Pharmacol 58:S21–S25
Mueller L, Solow T, Taylor N, Skwarecki B, Buels R, Binns J, Lin C, Wright M, Ahrens R, Wang Y, Herbst E, Keyder E, Menda N, Zamir D, Tanksley S (2005) The SOL Genomics Network: a comparative resource for Solanaceae biology and beyond. Plant Physiol 138:1310–1317
Wang X, Wang H, Wang J, Sun R, Wu J, Liu S, Bai Y, Mun JH, Bancroft I, Cheng F (2011) The genome of the mesopolyploid crop species Brassica rapa. Nat Genet 43:1035–1039
Brandt S, Kloska S, Altmann T, Kehr J (2002) Using array hybridization to monitor gene expression at the single cell level. J Exp Bot 53:2315–2323
Kawamura Y, Uemura M (2003) Mass spectrometric approach for identifying putative plasma membrane proteins of Arabidopsis leaves associated with cold acclimation. Plant J 36:141–154
Hu H, Boisson-Dernier A, Israelsson-Nordström M, Böhmer M, Xue S, Ries A, Godoski J, Kuhn JM, Schroeder JI (2009) Carbonic anhydrases are upstream regulators of CO2-controlled stomatal movements in guard cells. Nat Cell Biol 12:87–93
Sanchez R, Flores A, Cejudo FJ (2006) Arabidopsis phosphoenolpyruvate carboxylase genes encode immunologically unrelated polypeptides and are differentially expressed in response to drought and salt stress. Planta 223:901–909
Zhou BY, Ding ZS, Zhao M (2011) Alleviation of drought stress inhibition on photosynthesis by overexpression of PEPC in rice. Acta Agron Sin 37:112–118
Wang W, Vinocur B, Shoseyov O, Altman A (2004) Role of plant heat-shock proteins and molecular chaperones in the abiotic stress response. Trends Plant Sci 9:244–252
Sun W, Bernard C, Van De Cotte B, Van Montagu M, Verbruggen N (2001) At-HSP17.6A, encoding a small heat-shock protein in Arabidopsis, can enhance osmotolerance upon overexpression. Plant J 27:407–415
Nishizawa A, Yabuta Y, Yoshida E, Maruta T, Yoshimura K, Shigeoka S (2006) Arabidopsis heat shock transcription factor A2 as a key regulator in response to several types of environmental stress. Plant J 48:535–547
Verpoorte R (2000) Secondary metabolies. In: Verpoorte R, Alfermann AW (eds) Metabolic engineering of plant secondary metabolism. Kluwer, Dordrecht, pp 1–29
Goossens A, Häkkinen ST, Laakso I, Seppänen-Laakso T, Biondi S, De Sutter V, Lammertyn F, Nuutila AM, Söderlund H, Zabeau M (2003) A functional genomics approach toward the understanding of secondary metabolism in plant cells. Proc Nat Acad Sci USA 100:8595–8600
Nugroho LH, Verpoorte R (2002) Secondary metabolism in tobacco. Plant Cell, Tissue Organ Cult 68:105–125
Ruiz-Sola MÁ, Rodríguez-Concepción M (2012) Carotenoid biosynthesis in Arabidopsis: a colorful pathway. The Arabidopsis Book 10:e0158. doi:10.1199/tab.0158
Bezman Y, Bilkis I, Winterhalter P, Fleischmann P, Rouseff RL, Baldermann S, Naim M (2005) Thermal oxidation of 9′-cis-neoxanthin in a model system containing peroxyacetic acid leads to the potent odorant β-damascenone. J Agric Food Chem 53:9199–9206
Schwartz SH, Qin X, Zeevaart JAD (2003) Elucidation of the indirect pathway of abscisic acid biosynthesis by mutants, genes, and enzymes. Plant Physiol 131:1591–1601
Davison P, Hunter C, Horton P (2002) Overexpression of beta-carotene hydroxylase enhances stress tolerance in Arabidopsis. Nature 418:203–206
Auldridge ME, McCarty DR, Klee HJ (2006) Plant carotenoid cleavage oxygenases and their apocarotenoid products. Curr Opin Plant Biol 9:315–321
Acknowledgments
This work was supported by grants from the Key Special Program of China National Tobacco Corporation (TS-02-20110014), the Program of Guizhou Provincial Tobacco Company (2007-04 and 2010-02), the Foundation of Science and Technology of Guizhou Province of China (J[2010]2088), the Fundamental Research Funds for the Central Universities (XDJK2009B009 and XDJK2012A009), the Specialized Research Fund for the Doctoral Program of Higher Education of China (20100182120016), the “111” Project (B12006) and the Natural Science Foundation of Chongqing of China (cstc2011jjA80026).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Bo Lei, Xue-Hua Zhao, Kai Zhang and Fu-Zhang Ding equally contributed to this study.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Lei, B., Zhao, XH., Zhang, K. et al. Comparative transcriptome analysis of tobacco (Nicotiana tabacum) leaves to identify aroma compound-related genes expressed in different cultivated regions. Mol Biol Rep 40, 345–357 (2013). https://doi.org/10.1007/s11033-012-2067-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-012-2067-0