Abstract
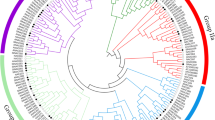
NAC (for NAM, ATAF1, 2, and CUC2) family genes have been found to play an important role in diversified developmental processes and environmental responses. A new NAC-type transcription factor SlNAC3 was primarily identified and isolated from the cDNA libraries of tomato cultivar Ailsa Craig. It contains three exons and two introns within genomic DNA sequence and encodes a polypeptide of 329 amino acids. A plant-specific and conserved NAC domain is located in the N-terminus of SlNAC3. The protein SlNAC3 is subcellularly localized in the nucleus of onion epidemical cells and it has a transcriptional activation domain in the C-terminal region which shows extremely divergent among NACs. Phylogenetic analysis showed that SlNAC3 belonged to the OsNAC3 subgroup of the NAC protein family. Tissue expression profile analysis revealed that SlNAC3 was expressed mainly in flower, fruit and root. The transcription expression of SlNAC3 was inhibited by salt, drought stress and ABA treatment. These data demonstrate that SlNAC3 might interact with environmental and endogenous stimuli and probably function when plants response to salt and drought stresses through ABA signaling pathways as a transcriptional activator.



Similar content being viewed by others
Abbreviations
- ABA:
-
Abscisic acid
- ABRE:
-
ABA-responsive element
- AC:
-
Ailsa Craig
- CaMV:
-
Cauliflower mosaic virus
- CUC:
-
Cup-shaped cotyledon
- DBD:
-
DNA-binding domain
- DRE/CRT:
-
Dehydration responsive element/C-repeat
- NAM:
-
No apical meristem
- NLS:
-
Nuclear localization signal
- SAM:
-
Shoot apical meristem
- UTR:
-
Untranslated region
- X-Gal:
-
5-Bromo-4-chloro-3-indolyl β-d-galactopyranoside
References
Zhu JK (2002) Salt and drought stress signal transduction in plants. Annu Rev Plant Biol 53:247–273. doi:10.1146/annurev.arplant.53.091401.143329
Ma H, Zhou H, Zhang H, Zhao J (2010) Cloning and expression analysis of an AP2/ERF gene and its responses to phytohormones and abiotic stresses in rice. Rice Sci 17:1–9. doi:10.1016/S1672-6308(08)60098-0
Hirota A, Kato T, Fukaki H, Aida M, Tasaka M (2007) The auxin-regulated AP2/EREBP gene PUCHI is required for morphogenesis in the early lateral root primordium of Arabidopsis. Plant Cell 19:2156–2168. doi:10.1105/tpc.107.050674
Ulm R, Baumann A, Oravecz A, Mate Z, Adam E, Oakeley EJ, Schafer E, Nagy F (2004) Genome-wide analysis of gene expression reveals function of the bZIP transcription factor HY5 in the UV-B response of Arabidopsis. Proc Natl Acad Sci USA 101:1397–1402. doi:10.1073/pnas.0308044100
Souer E, van Houwelingen A, Kloos D, Mol J, Koes R (1996) The no apical meristem gene of Petunia is required for pattern formation in embryos and flowers and is expressed at meristem and primordia boundaries. Cell 85:159–170. doi:10.1016/S0092-8674(00)81093-4
Aida M, Ishida T, Fukaki H, Fujisawa H, Tasaka M (1997) Genes involved in organ separation in Arabidopsis: an analysis of the cup-shaped cotyledon mutant. Plant Cell 9:841–857. doi:10.1105/tpc.9.6.841
Wang X, Basnayake BM, Zhang H, Li G, Li W, Virk N, Mengiste T, Song F (2009) The Arabidopsis ATAF1, a NAC transcription factor, is a negative regulator of defense responses against necrotrophic fungal and bacterial pathogens. Mol Plant Microbe Interact 22:1227–1238. doi:10.1094/MPMI-22-10-1227
Delessert C, Kazan K, Wilson IW, Van Der Straeten D, Manners J, Dennis ES, Dolferus R (2005) The transcription factor ATAF2 represses the expression of pathogenesis-related genes in Arabidopsis. Plant J 43:745–757. doi:10.1111/j.1365-313X.2005.02488.x
Hibara K, Takada S, Tasaka M (2003) CUC1 gene activates the expression of SAM-related genes to induce adventitious shoot formation. Plant J 36:687–696. doi:10.1046/j.1365-313X.2003.01911.x
Ernst HA, Olsen AN, Larsen S, Lo Leggio L (2004) Structure of the conserved domain of ANAC, a member of the NAC family of transcription factors. EMBO Rep 5:297–303. doi:10.1038/sj.embor.740009
He XJ, Mu RL, Cao WH, Zhang ZG, Zhang JS, Chen SY (2005) AtNAC2, a transcription factor downstream of ethylene and auxin signaling pathways, is involved in salt stress response and lateral root development. Plant J 44:903–916. doi:10.1111/j.1365-313X.2005.02575.x
Xie Q, Frugis G, Colgan D, Chua NH (2000) Arabidopsis NAC1 transduces auxin signal downstream of TIR1 to promote lateral root development. Genes Dev 14:3024–3036. doi:10.1101/gad.852200
Duval M, Hsieh TF, Kim SY, Thomas TL (2002) Molecular characterization of AtNAM: a member of the Arabidopsis NAC domain superfamily. Plant Mol Biol 50:237–248. doi:10.1023/A:1016028530943
Kikuchi K, Ueguchi-Tanaka M, Yoshida KT, Nagato Y, Matsusoka M, Hirano HY (2000) Molecular analysis of the NAC gene family in rice. Mol Gen Genet 262:1047–1051. doi:10.1007/PL00008647
Olsen AN, Ernst HA, Leggio LL, Skriver K (2005) NAC transcription factors: structurally distinct, functionally diverse. Trends Plant Sci 10:79–87. doi:10.1016/j.tplants.2004.12.010
Sablowski RW, Meyerowitz EM (1998) A homolog of NO APICAL MERISTEM is an immediate target of the floral homeotic genes APETALA3/PISTILLATA. Cell 92:93–103. doi:10.1016/S0092-8674(00)80902-2
Ren T, Qu F, Morris TJ (2000) HRT gene function requires interaction between a NAC protein and viral capsid protein to confer resistance to turnip crinkle virus. Plant Cell 12:1917–1926. doi:10.1105/tpc.12.10.1917
Kim SG, Kim SY, Park CM (2007) A membrane-associated NAC transcription factor regulates salt-responsive flowering via FLOWRING LOCUS T in Arabidopsis. Planta 226:647–654. doi:10.1007/s00425-007-0513-3
Hu H, Dai M, Yao J, Xiao B, Li X, Zhang Q, Xiong L (2006) Overexpressing a NAM, ATAF, and CUC (NAC) transcription factor enhances drought resistance and salt tolerance in rice. Proc Natl Acad Sci USA 103:12987–12992. doi:10.1073/pnas.0604882103
Wang YJ, Zhang ZG, He XJ, Zhou HL, Wen YX, Dai JX, Zhang JS, Chen SY (2003) A rice transcription factor OsbHLH1 is involved in cold stress response. Theor Appl Genet 107:1402–1409. doi:10.1007/s00122-003-1378-x
Selth LA, Dogra SC, Rasheed MS, Healy H, Randles JW, Rezaian MA (2005) A NAC domain protein interacts with tomato leaf curl virus replication accessory protein and enhances viral replication. Plant Cell 17:311–325. doi:10.1105/tpc.104.027235
Uppalapati SR, Ishiga Y, Wangdi T, Urbanczyk-Wochniak E, Ishiga T, Mysore KS, Bender CL (2008) Pathogenicity of Pseudomonas syringae pv. tomato on tomato seedlings: phenotypic and gene expression analyses of the virulence function of coronatine. Mol Plant Microbe Interact 21:383–395. doi:10.1094/MPMI-21-4-0383
Lu PL, Chen NZ, An R, Su Z, Qi BS, Ren F, Chen J, Wang XC (2007) A novel drought-inducible gene, ATAF1, encodes a NAC family protein that negatively regulates the expression of stress-responsive genes in Arabidopsis. Plant Mol Biol 63:289–305. doi:10.1007/s11103-006-9089-8
Skriver K, Olsen FL, Rogers JC, Mundy J (1991) cis-Acting DNA elements responsive to gibberellin and its antagonist abscisic acid. Proc Natl Acad Sci USA 88:7266–7270. doi:10.1073/pnas.88.16.7266
Xiong L, Ishitani M, Lee H, Zhu J (2001) The Arabidopsis LOS5/ABA3 locus encodes a molybdenum cofactor sulfurase and modulates cold stress- and osmotic stress-responsive gene expression. Plant Cell 13:2063–2083. doi:10.1105/tpc.13.9.2063
Riechmann JL, Heard J, Martin G, Reuber L, Jiang C, Keddie J, Adam L, Pineda O, Ratcliffe OJ, Samaha RR, Creelman R, Pilgrim M, Broun P, Zhang JZ, Ghandehari D, Sherman BK, Yu G (2000) Arabidopsis transcription factors: genome-wide comparative analysis among eukaryotes. Science 290:2105–2110. doi:10.1126/science.290.5499.2105
Fang Y, You J, Xie K, Xie W, Xiong L (2008) Systematic sequence analysis and identification of tissue-specific or stress-responsive genes of NAC transcription factor family in rice. Mol Genet Genomics 280:547–563. doi:10.1007/s00438-008-0386-6
Yang RC, Deng CT, Ouyang B, Ye ZB (2010) Molecular analysis of two salt-responsive NAC-family genes and their expression analysis in tomato. Mol Biol Rep. doi:10.1007/s11033-010-0177-0
Xie Q, Sanz-Burgos AP, Guo H, García JA, Gutiérrez C (1999) GRAB proteins, novel members of the NAC domain family, isolated by their interaction with a geminivirus protein. Plant Mol Biol 39:647–656. doi:10.1023/A:1006138221874
Lin RM, Zhao WS, Meng XB, Wang M, Peng YL (2007) Rice gene OsNAC19 encodes a novel NAC-domain transcription factor and responds to infection by Magnaporthe grisea. Plant Sci 172:120–130. doi:10.1016/j.plantsci.2006.07.019
Peng H, Cheng HY, Chen C, Yua XW, Yang JN, Gao WR, Shi QC, Zhang H, Li JG, Ma H (2009) A NAC transcription facto gene of Chickpea (Cicer arietinum), CarNAC3, is involved in drought stress response and various developmental processes. J Plant Physiol 166:1934–1945. doi:10.1016/j.jplph.2009.05.013
Ooka H, Satoh K, Doi K, Nagata T, Otomo Y, Murakami K, Matsubara K, Osato N, Kawai J, Carninci P, Hayashizaki Y, Suzuki K, Kojima K, Takahara Y, Yamamoto K, Kikuchi S (2003) Comprehensive analysis of NAC family genes in Oryza sativa and Arabidopsis thaliana. DNA Res 10:239–247. doi:10.1093/dnares/10.6.239
Cantu D, Blanco-Ulate B, Yang L, Labavitch JM, Bennett AB, Powell AL (2009) Ripening-regulated susceptibility of tomato fruit to Botrytis cinerea requires NOR but not RIN or ethylene. Plant Physiol 150:1434–1449. doi:10.1016/S0092-8674(00)81093-4
Nuruzzaman M, Manimekalai R, Sharoni AM, Satoh K, Kondoh H, Ooka H, Kikuchi S (2010) Genome-wide analysis of NAC transcription factor family in rice. Gene 465:30–44. doi:10.1016/j.gene.2010.06.008
Shinozaki K, Yamaguchi-Shinozaki K (2007) Gene networks involved in drought stress response and tolerance. J Exp Bot 58:221–227. doi:10.1093/jxb/erl164
Fan J, Gao X, Yang YW, Deng W, Li ZG (2007) Molecular cloning and characterization of a NAC-like gene in “navel” orange fruit response to postharvest stresses. Plant Mol Biol Rep 25:145–153. doi:10.1007/s11105-007-0016-1
Peng H, Cheng HY, Yu XW, Shi QH, Zhang H, Li JG, Ma H (2009) Characterization of a chickpea (Cicer arietinum L.) NAC family gene, CarNAC5, which is both developmentally- and stress-regulated. Plant Physiol Biochem 47:1037–1045. doi:10.1016/j.plaphy.2009.09.002
Xia N, Zhang G, Liu XY, Deng L, Cai GL, Zhang Y, Wang XJ, Zhao J, Huang LL, Kang ZS (2010) Characterization of a novel wheat NAC transcription factor gene involved in a defense response against stripe rust pathogen infection and abiotic stresses. Mol Biol Rep 37:3703–3712. doi:10.1007/s11033-010-0023-4
Shinozaki K, Yamaguchi-Shinozaki K (1997) Gene expression and signal transduction in water-stress response. Plant Physiol 115:327–334. doi:10.1104/pp.115.2.327
Shinozaki K, Yamaguchi-Shinozaki K (2000) Molecular responses to dehydration and low temperature, differences and cross-talk between two stress signaling pathways. Curr Opin Plant Biol 3:217–223. doi:10.1016/S1369-5266(00)80068-0
Finkelstein RR, Gampala SSL, Rock CD (2002) Abscisic acid signaling in seeds and seedlings. Plant Cell 14(Suppl):S15–S45. doi:10.1105/tpc.010441
Shinozaki K, Yamaguchi-Shinozaki K, Seki M (2003) Regulatory network of gene expression in the drought and cold stress responses. Curr Opin Plant Biol 5:410–417. doi:10.1016/S1369-5266(03)00092-X
Busk PK, Pagès M (1998) Regulation of abscisic acid-induced transcription. Plant Mol Biol 37:425–435. doi:10.1023/A:1006058700720
Narusaka Y, Nakashima K, Shinwari ZK, Sakuma Y, Furihata T, Abe H, Narusaka M, Shinozaki K, Yamaguchi-Shinozaki K (2003) Interaction between two cis-acting elements, ABRE and DRE, in ABA-dependent expression of Arabidopsis RD29, a gene in response to dehydration and high-salinity stresses. Plant J 34:137–148. doi:10.1046/j.1365-313X.2003.01708.x
Kant P, Kant S, Gordon M, Shaked R, Barak S (2007) Stress responsive suppressor1 and stress responsive suppressor2, two DEAD-box RNA helicases that attenuate Arabidopsis responses to multiple abiotic stresses. Plant Physiol 145:814–830. doi:10.1104/pp.107.099895
Abe H, Yamaguchi-Shinozaki K, Urao T, Iwasaki T, Hosokawa D, Shinozaki K (1997) Role of Arabidopsis MYC and MYB homologs in drought- and abscisic acid-regulated gene expression. Plant Cell 9:1859–1868. doi:10.1105/tpc.9.10.1859
Abe H, Urao T, Ito T, Seki M, Shinozaki K, Yamaguchi-Shinozaki K (2003) Arabidopsis AtMYC2 (bHLH) and AtMYB2 (MYB) function as transcriptional activators in abscisic acid signaling. Plant Cell 15:63–78. doi:10.1105/tpc.006130
Uno Y, Furihata T, Abe H, Yoshida R, Shinozaki K, Yamaguchi-Shinozaki K (2000) Arabidopsis basic leucine zipper transcription factors involved in an abscisic acid-dependent signal transduction pathway under drought and high-salinity conditions. Proc Natl Acad Sci USA 97:11632–11637. doi:10.1073/pnas.190309197
Yamaquchi-Shinozaki K, Shinozaki K (1992) The plant hormone abscisic acid mediates the drought-induced expression but not the seed-specific expression of rd22, a gene responsive to dehydration stress in Arabidopsis thaliana. Mol Gen Genet 238:17–25. doi:10.1007/BF00279525
Plesch G, Ehrhardt T, Mueller-Roeber B (2001) Involvement of TAAAG elements suggests a role for Dof transcription factors in guard cell-specific gene expression. Plant J 28:455–464. doi:10.1046/j.1365-313X.2001.01166.x
Acknowledgments
This work was supported by the grants of the Ministry of Science and Technology of China (973 Project, 2009CB119000), the National Science Foundation of China (NSFC Grants No. 30800755, 30871712, and 30921002).
Author information
Authors and Affiliations
Corresponding author
Additional information
Qinqin Han and Junhong Zhang contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Han, Q., Zhang, J., Li, H. et al. Identification and expression pattern of one stress-responsive NAC gene from Solanum lycopersicum . Mol Biol Rep 39, 1713–1720 (2012). https://doi.org/10.1007/s11033-011-0911-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11033-011-0911-2