Abstract
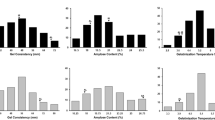
Improvement of eating and cooking quality (ECQ) traits is a major breeding target in indica rice due to market demand. In this study, amylose content, gel consistency, gelatinization temperature and 15 parameters from the viscosity profile were analyzed for quantitative trait loci (QTLs) with main effect, epistatic effects and their environmental interactions with ECQ, using 237 recombinant inbred lines (RILs) derived from a cross between indica cultivars Huahui 3 and Zhongguoxiangdao. Sixty-six QTLs were found across 2 years, of which 17 loci were detected in both years. No QTL was detected at the Wx locus, possibly because of allelism between the parents. A QTL cluster in the interval ALK6–RM549 on chromosome 6 had the largest effect on alkali spreading value, pasting time, time needed from initial viscosity increase to peak viscosity, pasting temperature, temperature needed from initial viscosity increase to peak viscosity, and pasting viscosity, and a minor effect on six traits (parameters): gel consistency, amylose content, peak viscosity, cool paste viscosity, breakdown viscosity and setback viscosity. Moreover, the Alk gene was responsible for two traits studied for the first time, namely peak temperature and pasting viscosity, controlled by QTLs qBAtime6 and qVA6, respectively. QTL clusters on other chromosomes showed similar characteristics to the Alk locus, although the variations explained were relatively minor. The OsBEIIb gene locus is linked to the interval RM301–RM424, which defined an important QTL cluster on chromosome 2. Moreover, a study using the same RIL population showed that a QTL for protein content and Rapid Visco Analyzer parameters was tightly linked to the same gene and directly affected ECQ. These findings not only demonstrate the complexity of ECQ, but also have important implications for map-based cloning of the underlying genetic factors and for breeding for rice quality.

Similar content being viewed by others
Abbreviations
- AC:
-
Amylose content
- Add:
-
Additive effect
- Alk :
-
Alkali gene locus
- ASV:
-
Alkali spreading value
- Atemp:
-
Pasting temperature
- Atime:
-
Pasting time
- BAtime:
-
Time needed from initial viscosity increase to PKV
- BAtemp:
-
Temperature needed from initial viscosity increase to PKV
- BE:
-
Branching enzyme
- BDV:
-
Breakdown viscosity
- BPC:
-
Brown rice protein content
- Btemp:
-
Peak temperature
- Btime:
-
Peak time
- CPV:
-
Cool paste viscosity
- CS:
-
Consistency viscosity
- ECQ:
-
Eating and cooking quality
- FV:
-
Final viscosity at 40 °C
- GC:
-
Gel consistency
- GT:
-
Gelatinization temperature
- HH3:
-
HuaHui 3
- HPV:
-
Hot paste viscosity
- InDel:
-
Insertion/deletion
- PKV:
-
Peak viscosity
- QTL:
-
Quantitative trait locus
- RIL:
-
Recombinant inbred lines
- RVA:
-
Rapid Visco Analyzer
- SB:
-
Setback viscosity
- SD:
-
Standard deviation
- SS:
-
Starch synthase
- SSR:
-
Simple sequence repeat
- V95:
-
Viscosity at 95 °C
- VA:
-
Pasting viscosity
- R 2 %:
-
Percentage phenotypic variation explained
- Wx :
-
Waxy gene locus
- ZGXD:
-
Zhongguoxiandao
References
Aluko G, Martinez C, Tohme J, Castano C, Bergman C, Oard JH (2004) QTL mapping of grain quality traits from the interspecific cross Oryza sativa & #x00D7; Oryza glaberrima. Theor Appl Genet 109:630–639
American Association of Cereal Chemists (2000) Approved methods for the AACC, 10th edn. Method 61-01 (amylograph method for milled rice) and Method 61-02 (determination of the pasting properties of rice with rapid visco analyzer). American Association of Cereal Chemists, St. Paul
Ayres NM, McClung AM, Larkin PD, Bligh HFJ, Jones CA, Park WD (1997) Microsatellites and a single-nucleotide polymorphism differentiate apparent amylose classes in an extended pedigree of US rice germplasm. Theor Appl Genet 94:773–781
Bao JS, Xia YW (1999) Genetic control of the paste viscosity characteristics in indica rice (Oryza sativa L.). Theor Appl Genet 98:1120–1124
Bao JS, Zheng XW, Xia YW, He P, Shu QY, Lu X, Chen Y, Zhu LH (2000) QTL mapping for the paste viscosity characteristics in rice (Oryza sativa L.). Theor Appl Genet 100:280–284
Bao JS, Wu YR, Hu B, Wu P, Cui HR, Shu QY (2002) QTL for rice grain quality based on a DH population derived from parents with similar apparent amylose content. Euphytica 128:317–324
Bo J (1996) Rice quality screening with the Rapid Visco Analyser. In: Walker CE, Hazelton JL (eds) Applications of the rapid visco analyser. Newport Scientific, Sydney, pp 19–24
Cagampang GB, Perez CM, Juliano BO (1973) A gel consistency test for eating quality in rice. J Sci Food Agric 24:1589–1594
Fan CC, Yu XQ, Xing YZ, Xu CG, Luo LJ, Zhang QF (2005) The main effects, epistatic effects and environmental interactions of QTLs on the cooking and eating quality of rice in a doubled-haploid line population. Theor Appl Genet 110:1445–1452
Gao ZY, Zeng DL, Cui X, Zhou YH, Yan MX, Huang DN, Li JY, Qian Q (2003) Map-based cloning of the ALK gene, which controls the gelatinization temperature of rice. Sci China (Ser C) 46:661–668
Gao ZY, Zeng DL, Cheng FM, Tian ZX, Guo LB, Su Y, Yan MX, Jiang H, Dong GJ, Huang YC, Han B, Li JY, Qian Q (2011) ALK, the key gene for gelatinization temperature, is a modifier gene for gel consistency in rice. J Integr Plant Biol 53:756–765
Harushima Y, Yano M, Shomura A, Sato M, Shimano T, Kuboki Y, Yamamoto T, Lin SY, Antonio BA, Parco A, Kajiya H, Huang N, Yamamoto K, Nagamura Y, Kurata N, Khush GS, Sasaki T (1998) A high-density rice genetic linkage map with 2,275 markers using a single F2 population. Genetics 148:479–494
He P, Li SG, Qian Q, Ma YQ, Li JZ, Wang WM, Chen Y, Zhu LH (1999) Genetic analysis of grain quality. Theor Appl Genet 98:502–508
Jantaboon J, Siangliw M, Im-mark S, Jamboonsri W, Vanavichit A, Toojinda T (2011) Ideotype breeding for submergence tolerance and cooking quality by marker-assisted selection in rice. Field Crops Res 123:206–213
Jiang GH, Xu CG, Tu JM, Li XH, He YQ, Zhang QF (2004) Pyramiding of insect- and disease-resistance genes into an elite indica, cytoplasm male sterile restorer line of rice, ‘Minghui 63’. Plant Breed 123:112–116
Juliano BO (1985) Rice chemistry and technology, 2nd edn. American Association of Cereal Chemists, St. Paul
Kharabian-Masouleh A, Waters DLE, Reinke RF, Ward R, Henry RJ (2012) SNP in starch biosynthesis genes associated with nutritional and functional properties of rice. Sci Rep 2(557):1–9
Kudo M (1968) Genetical and thremmatological studies of characters, physiological or ecological, in the hybrids between ecological rice groups. Bull Natl Inst Agric Sci Ser D 19:1–84
Lanceras JC, Huang ZL, Naivikul O, Vanavichit A, Ruanjaichon V, Tragoonrung S (2000) Mapping of genes for cooking and eating qualities in Thai jasmine rice (KDML105). DNA Res 7:93–101
Lincoln S, Daly M, Lander E (1992) Constructing genetics maps with MAPMAKER/EXP 3.0. Whitehead Institute Technical Report, Whitehead Institute, Cambridge
Little RR, Hilder GB, Dawson EH (1958) Differential effect of dilute alkali on 25 varieties of milled white rice. Cereal Chem 35:111–126
Liu XL, Wan XY, Ma XD, Wan JM (2011) Dissecting the genetic basis for the effect of rice chalkiness, amylose content, protein content, and rapid viscosity analyzer profile characteristics on the eating quality of cooked rice using the chromosome segment substitution line population across eight environments. Genome 54:64–80
Nassirou TY, Bo Y, Guang JG, Qing LZ, Cai GX, Jin HX, Yu QH (2013) QTL analysis of eating quality and cooking process of rice using a new RIL population derived from a cross between Minghui 63 and Khao Dawk Mali105. AJCS 7(13):2036–2047
Ni DH, Zhang SL, Chen S, Xu Y, Li L, Li H, Wang ZY, Cai XL, Li ZF, Yang JB (2011) Improving cooking and eating quality of Xieyou57, an elite indica hybrid rice, by marker-assisted selection of the Wx locus. Euphytica 179:355–362
Sano Y (1984) Differential regulation of waxy gene expression in rice endosperm. Theor Appl Genet 68:467–473
Sano Y, Katsumata M, Okuno K (1986) Genetic studies of speciation in cultivated rice. 5. Inter-and intraspecific differentiation in the waxy gene expression of rice. Euphytica 35:1–9
Sasaki A, Ashikari M, Ueguchi-Tanaka M, Itoh H, Nishimura A, Swapan D, Ishiyama K, Saito T, Kobayashi M, Khush GS et al (2002) Green revolution: a mutant gibberellin-synthesis gene in rice. Nature 416:701–702
Standards Press of China (1999a) National Standard of People’s Republic of China. High quality paddy, GB2905-82. Standards Press of China, Beijing
Standards Press of China (1999b) National Standard of People’s Republic of China. High quality paddy, GB/T17891-1999. Standards Press of China, Beijing
Tan YF, Li JX, Yu SB, Xing YZ, Xu CG, Zhang Q (1999) The three important traits for cooking and eating quality of rice grains are controlled by a single locus in an elite rice hybrid, Shanyou 63. Theor Appl Genet 99:642–648
Tian ZX, Qian Q, Liu QQ, Yan MX, Liu XF, Yan CJ, Liu GF, Gao ZY, Tang SZ, Zeng DL, Wang YH, Yu JM, Gu MH, Li JY (2009) Allelic diversity in rice starch biosynthesis pathway leads to a diverse array of rice eating and cooking qualities. Proc Natl Acad Sci USA 106:21760–21765
Umemoto T, Aoki N (2005) Single-nucleotide polymorphisms in rice starch synthase IIa that alter starch gelatinization and starch association of the enzyme. Funct Plant Biol 32:763–768
Umemoto T, Yano M, Satoh H, Shomura A, Nakamura Y (2002) Mapping of a gene responsible for the difference in amylopectin structure between japonica-type and indica-type rice varieties. Theor Appl Genet 104:1–8
Wang SC, Zeng ZB (2004) Windows QTL Cartographer 2.0. Department of Statistics, North Carolina State University, Raleigh, NC, 2001–2004. http://statgen.ncsu.edu/qtlcart/WQTLCart.htm
Wang ZY, Wu ZL, Xing YY, Zheng FG, Guo XL, Zhang WG, Hong MM (1990) Nucleotide sequence of rice waxy gene. Nucleic Acids Res 18(19):5898
Wang LQ, Liu WJ, Xu Y, He YQ, Luo LJ, Xing YZ, Xu CG, Zhang Q (2007) Genetic basis of 17 traits and viscosity parameters characterizing the eating and cooking quality of rice grain. Theor Appl Genet 115:463–476
Wang LQ, Zhong M, Li XH, Yuan DJ, Xu YB, Liu H, He YQ, Luo LJ, Zhang Q (2008) The QTL controlling amino acid content in grain of rice (Oryza sativa) were co-localized with the regions involved in amino acid metabolism pathway. Mol Breed 21:127–137
Wu HK, Liu SJ, Jiang L, Zhang WW, Wang YH, Ren YL, Han XH, Liu F (2009) Relationship between protein composition and total protein content and starch RVA profile properties in rice. Chin J Rice Sci 23:421–426
Xu FF, Zhang G, Tong C, Sun X, Corke H, Sun M, Bao J (2013) Association mapping of starch physicochemical properties with starch biosynthesizing genes in waxy rice (Oryza sativa L.). J Agric Food Chem 61:10110–10117
Yan CJ, Tian ZX, Fang YW, Yang YC, Li J, Zeng SY, Gu SL, Xu CW, Tang SZ, Gu MH (2011) Genetic analysis of starch paste viscosity parameters in glutinous rice (Oryza sativa L.). Theor Appl Genet 122:63–76
Yan B, Wang Y, Gao GJ, Zhang QL, Liu X, He YQ (2012) QTL Mapping and epistasis analysis of the protein content in brown rice. Mol Plant Breed 10:594–599 (in Chinese with English abstract)
Yang J, Hu C, Hu H, Yu R, Xia Z, Ye X, Zhu J (2008) QTLNetwork: mapping and visualizing genetic architecture of complex traits in experimental populations. Bioinformatics 24:721–723
Zhong M, Wang L, Yuan D, Luo L, Xu C, He YQ (2011) Identification of QTL affecting protein and amino acid contents in rice. Rice Sci 18:187–195
Acknowledgments
We thank Professor Robert A. McIntosh (Plant Breeding Institute, University of Sydney) for editing this manuscript. This work was supported in part by the National Program on R&D of Transgenic Plants (2014ZX0800936B), the National 863 Project (2012AA101102), the National Natural Science Foundation of China (31371701), the Foundation of Ministry of Agriculture (CARS-01-03, 2012G2) and the Bill & Melinda Gates Foundation.
Author information
Authors and Affiliations
Corresponding author
Additional information
Bao Yan and Nassirou Tondi Yacouba have contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yan, B., Tondi Yacouba, N., Chen, J. et al. Analysis of minor quantitative trait loci for eating and cooking quality traits in rice using a recombinant inbred line population derived from two indica cultivars with similar amylose content. Mol Breeding 34, 2151–2163 (2014). https://doi.org/10.1007/s11032-014-0170-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11032-014-0170-8