Abstract
Studies on the molecular mechanisms of dehydration tolerance have been largely limited to plants and invertebrates. Currently, research in whole body dehydration of complex animals is limited to cognitive and behavioral effects in humans, leaving the molecular mechanisms of vertebrate dehydration relatively unexplored. The present review summarizes studies to date on the African clawed frog (Xenopus laevis) and examines whole-body dehydration on physiological, cellular and molecular levels. This aquatic frog is exposed to seasonal droughts in its native habitat and can endure a loss of over 30% of its total body water. When coping with dehydration, osmoregulatory processes prioritize water retention in skeletal tissues and vital organs over plasma volume. Although systemic blood circulation is maintained in the vital organs and even elevated in the brain during dehydration, it is done so at the expense of reduced circulation to the skeletal muscles. Increased hemoglobin affinity for oxygen helps to counteract impaired blood circulation and metabolic enzymes show altered kinetic and regulatory parameters that support the use of anaerobic glycolysis. Recent studies with X. laevis also show that pro-survival pathways such as antioxidant defenses and heat shock proteins are activated in an organ-specific manner during dehydration. These pathways are tightly coordinated at the post-transcriptional level by non-coding RNAs, and at the post-translational level by reversible protein phosphorylation. Paired with ongoing research on the X. laevis genome, the African clawed frog is poised to be an ideal animal model with which to investigate the molecular adaptations for dehydration tolerance much more deeply.


Similar content being viewed by others
References
Abe AS (1995) Estivation in South American amphibians and reptiles. Brazilian J Med Biol Res 28:1241–1247
Loveridge PJ (1976) Strategies of water conservation in Southern African frogs. Zool Africana 11:319–333
Seymour RS, Lee AK (1974) Physiological adaptations of anuran amphibians to aridity: Australian prospects. Aust Zool 18:53–65
Pinder AW, Storey KB, Ultsch GR (1992) Estivation and hibernation. In: Feder ME, Burggren W (eds) Environmental biology of the Amphibia. University of Chicago Press, Chicago, pp 250–274
Storey KB (2002) Life in the slow lane: molecular mechanisms of estivation. Comp Biochem Physiol – A Mol Integr Physiol 133:733–754. https://doi.org/10.1016/S1095-6433(02)00206-4
Balinsky JB, Cragg MM, Baldwin E (1961) The adaptation of amphibian waste nitrogen excretion to dehydration. Comp Biochem Physiol 3:236–244
Groom DJE, Kuchel L, Richards JG (2013) Metabolic responses of the South American ornate horned frog (Ceratophrys ornata) to estivation. Comp Biochem Physiol Part B Biochem Mol Biol 164:2–9. https://doi.org/10.1016/j.cbpb.2012.08.001
Tinsley RC, Kobel HR (1996) The biology of Xenopus. Clarendon Press, Oxford
Hillman SS (1978) The roles of oxygen delivery and electrolyte levels in the dehydrational death of Xenopus laevis. J Comp Physiol B 128:169–175
Balinsky JB, Choritz EL, Coe CG, van der Schans GS (1967) Amino acid metabolism and urea synthesis in naturally aestivating Xenopus laevis. Comp Biochem Physiol 22:59–68. https://doi.org/10.1016/0010-406x(67)90166-1
Jokumsen A, Weber RE (1980) Haemoglobin-oxygen binding properties in the blood of Xenopus laevis, with special reference to the influences of aestivation and of temperature and salinity acclimation. J Exp Biol 86:19–37
Jørgensen CB (1997) Urea and amphibian water economy. Comp Biochem Physiol A Physiol 117:161–170
Malik AI, Storey KB (2009) Activation of extracellular signal-regulated kinases during dehydration in the African clawed frog, Xenopus laevis. J Exp Biol 212:2595–2603
Hillman SS (1978) Some effects of dehydration on internal distributions of water and solutes in Xenopus laevis. Comp Biochem Physiol Part A Physiol 61:303–307
Hillman SS, Sommerfeldt RW (1981) Microsphere studies of amphibian systemic blood flow redistribution during dehydration, hypovolemia, and salt load. J Exp Zool 218:305–308
Wu C-W, Tessier SN, Storey KB (2017) Regulation of the insulin-Akt signaling pathway and glycolysis during dehydration stress in the African clawed frog Xenopus laevis. Biochem Cell Biol 95:663–671. https://doi.org/10.1139/bcb-2017-0117
Childers CL, Storey KB (2016) Post-translational regulation of hexokinase function and protein stability in the aestivating frog Xenopus laevis. Protein J 35:61–71. https://doi.org/10.1007/s10930-016-9647-0
Dawson NJ, Biggar Y, Malik AI, Storey KB (2018) Increased transcript levels and kinetic function of pyruvate kinase during severe dehydration in aestivating African clawed frogs, Xenopus laevis. Comp Biochem Physiol B Biochem Mol Biol 0–1. https://doi.org/10.1016/j.cbpb.2018.01.003
Katzenback BA, Dawson NJ, Storey KB (2014) Purification and characterization of a urea sensitive lactate dehydrogenase from the liver of the African clawed frog, Xenopus laevis. J Comp Physiol B 184:601–611. https://doi.org/10.1007/s00360-014-0824-1
Childers CL, Storey KB (2019) Purification and characterization of a urea sensitive lactate dehydrogenase from skeletal muscle of the African clawed frog, Xenopus laevis. J Comp Physiol B Biochem Syst Environ Physiol 189:271–281. https://doi.org/10.1007/s00360-018-1200-3
Cowan KJ, Storey KB (2003) Mitogen-activated protein kinases: new signaling pathways functioning in cellular responses to environmental stress. J Exp Biol 206:1107–1115. https://doi.org/10.1242/jeb.00220
Mitchell TJ, John S (2005) Signal transducer and activator of transcription (STAT) signalling and T-cell lymphomas. Immunology 114:301–312
Chung J, Uchida E, Grammer TC, Blenis J (1997) STAT3 serine phosphorylation by ERK-dependent and -independent pathways negatively modulates its tyrosine phosphorylation. Mol Cell Biol 17:6508–6516. https://doi.org/10.1128/MCB.17.11.6508
Krishna M, Narang H (2008) The complexity of mitogen-activated protein kinases (MAPKs) made simple. Cell Mol Life Sci 65:3525–3544
Bhattacharya S, Ray RM, Johnson LR (2005) STAT3-mediated transcription of Bcl-2, Mcl-1 and c-IAP2 prevents apoptosis in polyamine-depleted cells. Biochem J 392:335–344. https://doi.org/10.1042/BJ20050465
Burnell AM, Tunnacliffe ALAN (2011) Gene induction and desiccation stress in nematodes. In: Molecular and physiological basis of nematode survival. CABI, Wallingford, pp 126–156
Huang Z, Banton MC, Tunnacliffe A (2010) Modeling anhydrobiosis: activation of the mitogen-activated protein kinase ERK by dehydration in both human cells and nematodes. J Exp Zool Part A Ecol Genet Physiol 313A:660–670. https://doi.org/10.1002/jez.637
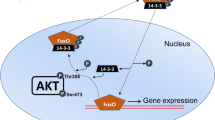
Malik AI, Storey KB (2011) Transcriptional regulation of antioxidant enzymes by FoxO1 under dehydration stress. Gene 485:114–119. https://doi.org/10.1016/j.gene.2011.06.014
Zhang Y, Luu BE, Storey KB (2018) FoxO4 activity is regulated by phosphorylation and the cellular environment during dehydration in the African clawed frog, Xenopus laevis. Biochim Biophys Acta Gen Subj 1862:1721–1728. https://doi.org/10.1016/j.bbagen.2018.05.002
Giraud-Billoud M, Rivera-Ingraham GA, Moreira DC et al (2019) Twenty years of the ‘Preparation for Oxidative Stress’ (POS) theory: Ecophysiological advantages and molecular strategies. Comp Biochem Physiol -Part A Mol Integr Physiol 234:36–49. https://doi.org/10.1016/j.cbpa.2019.04.004
Moreira DC, Oliveira MF, Liz-Guimarães L et al (2017) Current trends and research challenges regarding “preparation for oxidative stress”. Front Physiol 8:702. https://doi.org/10.3389/fphys.2017.00702
Zhu H, Itoh K, Yamamoto M et al (2005) Role of Nrf2 signaling in regulation of antioxidants and phase 2 enzymes in cardiac fibroblasts: protection against reactive oxygen and nitrogen species-induced cell injury. FEBS Lett 579:3029–3036. https://doi.org/10.1016/j.febslet.2005.04.058
Jung K-A, Choi B-H, Nam C-W et al (2013) Identification of aldo-keto reductases as NRF2-target marker genes in human cells. Toxicol Lett 218:39–49. https://doi.org/10.1016/j.toxlet.2012.12.026
Malik AI, Storey KB (2009) Activation of antioxidant defense during dehydration stress in the African clawed frog. Gene 442:99–107. https://doi.org/10.1016/j.gene.2009.04.007
Feder ME, Hofmann GE (1999) Heat-shock proteins, molecular chaperones, and the stress response: evolutionary and ecological physiology. Annu Rev Physiol 61:243–282. https://doi.org/10.1146/annurev.physiol.61.1.243
Tuttle AM, Gauley J, Chan N, Heikkila JJ (2007) Analysis of the expression and function of the small heat shock protein gene, hsp27, in Xenopus laevis embryos. Comp Biochem Physiol Part A Mol Integr Physiol 147:112–121. https://doi.org/10.1016/j.cbpa.2006.12.003
Ali A, Salter-Cid L, Flajnik MF, Heikkila JJ (1996) Isolation and characterization of a cDNA encoding a Xenopus 70-kDa heat shock cognate protein, Hsc70.I. Comp Biochem Physiol – B Biochem Mol Biol 113:681–687. https://doi.org/10.1016/0305-0491(95)02081-0
Heikkila JJ (2010) Heat shock protein gene expression and function in amphibian model systems. Comp Biochem Physiol Part A Mol Integr Physiol 156:19–33. https://doi.org/10.1016/j.cbpa.2010.01.024
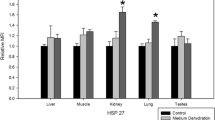
Luu BE, Wijenayake S, Malik AI, Storey KB (2017) The regulation of heat shock proteins in response to dehydration in Xenopus laevis. Cell Stress Chaperones 23:1–9
Garrido C, Bruey J-M, Ducasse C et al (2000) Hsp27 negatively regulates cell death by interacting with cytochrome c. Nat Cell Biol 2:645–652. https://doi.org/10.1038/35023595
Concannon CG, Orrenius S, Samali A (2001) Hsp27 inhibits cytochrome c-mediated caspase activation by sequestering both pro-caspase-3 and cytochrome c. Gene Expr 9:195–201
Goldfarb SB, Kashlan OB, Watkins JN et al (2006) Differential effects of Hsc70 and Hsp70 on the intracellular trafficking and functional expression of epithelial sodium channels. Proc Natl Acad Sci U S A 103:5817–5822. https://doi.org/10.1073/pnas.0507903103
Lu HAJ, Sun TX, Matsuzaki T et al (2007) Heat shock protein 70 interacts with aquaporin-2 and regulates its trafficking. J Biol Chem 282:28721–28732. https://doi.org/10.1074/jbc.M611101200
Kampinga HH, Craig EA (2010) The HSP70 chaperone machinery: J proteins as drivers of functional specificity. Nat Rev Mol Cell Biol 11:579–592. https://doi.org/10.1038/nrm2941
Greene MK, Maskos K, Landry SJ (1998) Role of the J-domain in the cooperation of Hsp40 with Hsp70. Proc Natl Acad Sci U S A 95:6108–6113
Suh WC, Burkholder WF, Lu CZ et al (1998) Interaction of the Hsp70 molecular chaperone, DnaK, with its cochaperone DnaJ. Proc Natl Acad Sci U S A 95:15223–15228
Cajo GC, Horne BE, Kelley WL et al (2006) The role of the DIF motif of the DnaJ (Hsp40) co-chaperone in the regulation of the DnaK (Hsp70) chaperone cycle. J Biol Chem 281:12436–12444. https://doi.org/10.1074/jbc.M511192200
Zou J, Guo Y, Guettouche T et al (1998) Repression of heat shock transcription factor HSF1 activation by HSP90 (HSP90 complex) that forms a stress-sensitive complex with HSF1. Cell 94:471–480. https://doi.org/10.1016/S0092-8674(00)81588-3
Bartel DP (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116:281–297
Storey KB, Wu C-W (2013) Stress response and adaptation: a new molecular toolkit for the 21st century. Comp Biochem Physiol A Mol Integr Physiol 165:417–428. https://doi.org/10.1016/j.cbpa.2013.01.019
Storey KB (2015) Regulation of hypometabolism: insights into epigenetic controls. J Exp Biol 218:150–159
Wu CW, Biggar KK, Storey KB (2013) Dehydration mediated microRNA response in the African clawed frog Xenopus laevis. Gene 529:269–275
Luu BE, Storey KB (2015) Dehydration triggers differential microRNA expression in Xenopus laevis brain. Gene 573:64–69
Gao Y, Su J, Guo W et al (2015) Inhibition of miR-15a promotes BDNF expression and rescues dendritic maturation deficits in MeCP2-deficient neurons. Stem Cells 33:1618–1629. https://doi.org/10.1002/stem.1950
Varendi K, Kumar A, Härma MA, Andressoo JO (2014) MIR-1, miR-10b, miR-155, and miR-191 are novel regulators of BDNF. Cell Mol Life Sci 71:4443–4456
Yin KJ, Deng Z, Hamblin M et al (2010) Peroxisome proliferator-activated receptor regulation of miR-15a in ischemia-induced cerebral vascular endothelial injury. J Neurosci 30:6398–6408. https://doi.org/10.1523/JNEUROSCI.0780-10.2010
Xu L-J, Ouyang Y-B, Xiong X et al (2015) Post-stroke treatment with miR-181 antagomir reduces injury and improves long-term behavioral recovery in mice after focal cerebral ischemia. Exp Neurol 264:1–7. https://doi.org/10.1016/j.expneurol.2014.11.007
Moon J, Xu L, Giffard RG (2013) Inhibition of microRNA-181 reduces forebrain ischemia-induced neuronal loss. J Cereb Blood Flow Metab 33:1976–1982. https://doi.org/10.1038/jcbfm.2013.157
Hawkins LJ, Storey KB (2020) MicroRNA expression in the heart of Xenopus laevis facilitates metabolic adaptation to dehydration. Genomics. https://doi.org/10.1016/j.ygeno.2020.04.003
Ha M, Kim VN (2014) Regulation of microRNA biogenesis. Nat Rev Mol Cell Biol 15:509–524
Hadj-Moussa H, Storey KB (2018) Micromanaging freeze tolerance: the biogenesis and regulation of neuroprotective microRNAs in frozen brains. Cell Mol Life Sci 75:1–13
Biggar KK, Storey KB (2014) New approaches to comparative and animal stress biology research in the post-genomic era: a contextual overview. Comput Struct Biotechnol J
Biggar KK, Biggar Y, Storey KB (2015) Identification of a novel dehydration responsive gene, drp10, from the African clawed frog, Xenopus laevis. J Exp Zool A Ecol Genet Physiol 323:375–381. https://doi.org/10.1002/jez.1930
Hawkins LJ, Luu BE, Storey KB (2018) Selection of reference genes for accurate RT-qPCR analysis of dehydration tolerance in Xenopus laevis. Gene Rep 13:192–198. https://doi.org/10.1016/j.genrep.2018.10.006
Session AM, Uno Y, Kwon T et al (2016) Genome evolution in the allotetraploid frog Xenopus laevis. Nature 538:336–343. https://doi.org/10.1038/nature19840
Acknowledgments
The authors thank JM Storey for editorial review of the manuscript. Research in the Storey lab was supported by a Discovery grant (#6793) from the Natural Sciences and Engineering Research Council of Canada (NSERC) to KBS, an NSERC postgraduate scholarship to BEL, and an Ontario Graduate Scholarship to LJH. Thanks to JM Weber and SPJ Brooks for their discussions that inspired this review. BioRender was used in the illustration of the figures.
Funding
Research in the Storey lab was supported by a Discovery Grant (#6793) from the Natural Sciences and Engineering Research Council of Canada (NSERC) to KBS, an NSERC postgraduate scholarship to BEL, and an Ontario Graduate Scholarship to LJH.
Author information
Authors and Affiliations
Contributions
BEL and LJH wrote the manuscript. All authors contributed to editing the manuscript and approved of the final version.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Luu, B.E., Hawkins, L.J. & Storey, K.B. Insights from a vertebrate model organism on the molecular mechanisms of whole-body dehydration tolerance. Mol Cell Biochem 476, 2381–2392 (2021). https://doi.org/10.1007/s11010-021-04072-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11010-021-04072-x