Abstract
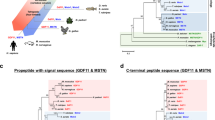
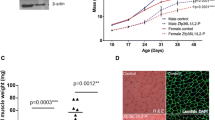
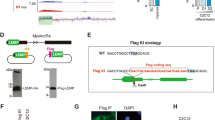
The growth and differentiation factor-11 (GDF-11) gene is thought to code for a single protein that plays a crucial role in regulating the development of multiple tissues. In this study, we aimed to investigate if the GDF-11 gene has another transcript and, if so, to characterise this transcript and determine its tissue-specific and developmental expression. We have identified a novel transcript of GDF-11 in mouse muscle, which contains the 3′ region of intron 1, exon 2, exon 3 and 3′UTR, and has two transcription initiation sites and a single termination site. We named the novel transcript GDF-11ΔEx1 because it does not contain exon 1 of canonical GDF-11. The GDF-11ΔEx1 transcript was expressed in the skeletal muscles, heart, brain and kidney, but was undetectable in the liver and gut. The concentration of the GDF-11ΔEx1 transcript was increased in gastrocnemius muscles from three to 6 weeks of age, a period of accelerated muscle growth, steadily declined thereafter and was higher in male than female mice (P < 0.001 for age and sex). GDF-11ΔEx1 cDNA was predicted to code for a putative N-terminal-truncated propeptide and the canonical ligand for GDF-11. However, propeptide-specific antibodies could not identify proteins of the expected size in skeletal muscle. Interestingly, in silico analysis of the GDF-11ΔEx1 RNA predicted a secondary structure with the potential to coordinate multiple protein interactions as a molecular scaffold. Therefore, we postulate that GDF-11ΔEx1 may act as a long non-coding RNA to regulate the transcription of canonical GDF-11 and/or other genes in skeletal muscle and other tissues.







Similar content being viewed by others
References
McPherron AC, Lawler AM, Lee SJ (1999) Regulation of anterior/posterior patterning of the axial skeleton by growth/differentiation factor 11. Nat Genet 22:260–264. doi:10.1038/10320
Nakashima M, Toyono T, Akamine A, Joyner A (1999) Expression of growth/differentiation factor 11, a new member of the BMP/TGFbeta superfamily during mouse embryogenesis. Mech Dev 80:185–189
Gamer LW, Wolfman NM, Celeste AJ, Hattersley G, Hewick R, Rosen V (1999) A novel BMP expressed in developing mouse limb, spinal cord, and tail bud is a potent mesoderm inducer in Xenopus embryos. Dev Biol 208:222–232. doi:10.1006/dbio.1998.9191
Gamer LW, Cox KA, Small C, Rosen V (2001) Gdf11 is a negative regulator of chondrogenesis and myogenesis in the developing chick limb. Dev Biol 229:407–420. doi:10.1006/dbio.2000.9981
Xing F, Tan X, Zhang PJ, Ma J, Zhang Y, Xu P, Xu Y (2007) Characterization of amphioxus GDF8/11 gene, an archetype of vertebrate MSTN and GDF11. Dev Genes Evol 217:549–554. doi:10.1007/s00427-007-0162-3
Jeanplong F, Falconer SJ, Thomas M, Matthews KG, Oldham JM, Watson T, McMahon CD (2012) Growth and differentiation factor-11 is developmentally regulated in skeletal muscle and inhibits myoblast differentiation. Open J Mol Integr Physiol 2:127–138. doi:10.4236/ojmip.2012.24018
McPherron AC, Lawler AM, Lee SJ (1997) Regulation of skeletal muscle mass in mice by a new TGF-beta superfamily member. Nature 387:83–90. doi:10.1038/387083a0
Lee SJ, McPherron AC (2001) Regulation of myostatin activity and muscle growth. Proc Natl Acad Sci USA 98:9306–9311. doi:10.1073/pnas.151270098
Rebbapragada A, Benchabane H, Wrana JL, Celeste AJ, Attisano L (2003) Myostatin signals through a transforming growth factor beta-like signaling pathway to block adipogenesis. Mol Cell Biol 23:7230–7242
Oh SP, Yeo CY, Lee Y, Schrewe H, Whitman M, Li E (2002) Activin type IIA and IIB receptors mediate Gdf11 signaling in axial vertebral patterning. Genes Dev 16:2749–2754. doi:10.1101/gad.1021802
Lee SJ, Reed LA, Davies MV, Girgenrath S, Goad ME, Tomkinson KN, Wright JF, Barker C, Ehrmantraut G, Holmstrom J, Trowell B, Gertz B, Jiang MS, Sebald SM, Matzuk M, Li E, Liang LF, Quattlebaum E, Stotish RL, Wolfman NM (2005) Regulation of muscle growth by multiple ligands signaling through activin type II receptors. Proc Natl Acad Sci USA 102:18117–18122. doi:10.1073/pnas.0505996102
Covi JA, Kim HW, Mykles DL (2008) Expression of alternatively spliced transcripts for a myostatin-like protein in the blackback land crab, Gecarcinus lateralis. Comp Biochem Physiol A 150:423–430. doi:10.1016/j.cbpa.2008.04.608
Castelhano-Barbosa EC, Gabriel JE, Alvares LE, Monteiro-Vitorello CB, Coutinho LL (2005) Temporal and spatial expression of the myostatin gene during chicken embryo development. Growth Dev Aging 69:3–12
Huang KL, Wang JW, Han CC, Liu HH, Li L, Dai F, Pan Z, Xu F, He H, Xu H (2011) Developmental expression and alternative splicing of the duck myostatin gene. Comp Biochem Physiol Part D Genomics Proteomics 6:238–243. doi:10.1016/j.cbd.2011.04.002
Garikipati DK, Gahr SA, Roalson EH, Rodgers BD (2007) Characterization of rainbow trout myostatin-2 genes (rtMSTN-2a and -2b): genomic organization, differential expression, and pseudogenization. Endocrinology 148:2106–2115. doi:10.1210/en.2006-1299
Jeanplong F, Falconer SJ, Oldham JM, Thomas M, Gray TS, Hennebry A, Matthews KG, Kemp FC, Patel K, Berry C, Nicholas G, McMahon CD (2013) Discovery of a mammalian splice variant of myostatin that stimulates myogenesis. PLoS ONE 8:e81713. doi:10.1371/journal.pone.0081713
Lundby C, Nordsborg N, Kusuhara K, Kristensen KM, Neufer PD, Pilegaard H (2005) Gene expression in human skeletal muscle: alternative normalization method and effect of repeated biopsies. Eur J Appl Physiol 95:351–360. doi:10.1007/s00421-005-0022-7
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685
Tukey JW (1953) Some selected quick and easy methods of statistical analysis. Trans N Y Acad Sci 16:88–97
Oldham JM, Osepchook CC, Jeanplong F, Falconer SJ, Matthews KG, Conaglen JV, Gerrard DF, Smith HK, Wilkins RJ, Bass JJ, McMahon CD (2009) The decrease in mature myostatin protein in male skeletal muscle is developmentally regulated by growth hormone. J Physiol 587:669–677. doi:10.1113/jphysiol.2008.161521
Zuker M (2003) Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res 31:3406–3415
Lee TI, Young RA (2000) Transcription of eukaryotic protein-coding genes. Annu Rev Genet 34:77–137. doi:10.1146/annurev.genet.34.1.77
Funkenstein B, Olekh E (2010) Growth/differentiation factor-11: an evolutionary conserved growth factor in vertebrates. Dev Genes Evol 220:129–137. doi:10.1007/s00427-010-0334-4
Esquela AF, Lee SJ (2003) Regulation of metanephric kidney development by growth/differentiation factor 11. Dev Biol 257:356–370
McPherron AC, Huynh TV, Lee SJ (2009) Redundancy of myostatin and growth/differentiation factor 11 function. BMC Dev Biol 9:24. doi:10.1186/1471-213X-9-24
Dinger ME, Pang KC, Mercer TR, Mattick JS (2008) Differentiating protein-coding and noncoding RNA: challenges and ambiguities. PLoS Comput Biol 4:e1000176. doi:10.1371/journal.pcbi.1000176
Kapranov P, St Laurent G, Raz T, Ozsolak F, Reynolds CP, Sorensen PH, Reaman G, Milos P, Arceci RJ, Thompson JF, Triche TJ (2010) The majority of total nuclear-encoded non-ribosomal RNA in a human cell is ‘dark matter’ un-annotated RNA. BMC Biol 8:149. doi:10.1186/1741-7007-8-149
St Laurent G, Savva YA, Kapranov P (2012) Dark matter RNA: an intelligent scaffold for the dynamic regulation of the nuclear information landscape. Front Genet 3:57. doi:10.3389/fgene.2012.00057
Wang KC, Chang HY (2011) Molecular mechanisms of long noncoding RNAs. Mol Cell 43:904–914. doi:10.1016/j.molcel.2011.08.018
St Laurent G, Shtokalo D, Tackett MR, Yang Z, Eremina T, Wahlestedt C, Urcuqui-Inchima S, Seilheimer B, McCaffrey TA, Kapranov P (2012) Intronic RNAs constitute the major fraction of the non-coding RNA in mammalian cells. BMC Genom 13:504. doi:10.1186/1471-2164-13-504
Acknowledgements
We thank Dr Neil Cox for his help with statistical analysis. We also thank Linda Tseng for technical assistance, and Drs Christine Couldrey and Adrian Molenaar for critically reading the manuscript. This study was supported by a research grant (C10X0703) from the Foundation of Research Science and Technology, New Zealand.
Conflict of interest
Authors declare no competing financial interests.
Data deposition
All novel DNA sequences were deposited to GenBank under the following accession numbers: KC561772, KC561773 and KC561774.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Jeanplong, F., Falconer, S.J., Oldham, J.M. et al. Identification and expression of a novel transcript of the growth and differentiation factor-11 gene. Mol Cell Biochem 390, 9–18 (2014). https://doi.org/10.1007/s11010-013-1949-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11010-013-1949-3