Abstract
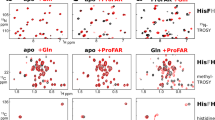
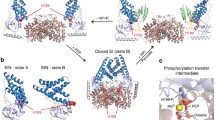
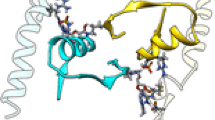
IGPS is a 51 kDa heterodimeric enzyme comprised of two proteins, HisH and HisF, that catalyze the hydrolysis of glutamine to produce NH3 in the HisH active site and the cyclization of ammonia with N′-[(5′-phosphoribulosyl)formimino]-5-aminoimidazole-4-carboxamide-ribonucleotide (PRFAR) in HisF to produce imidazole glycerol phosphate (IGP) and 5-aminoimidazole-4-carboxamide ribotide (AICAR). Binding of PRFAR and IGP stimulates glutaminase activity in the HisH enzyme over 5,000 and 100-fold, respectively, despite the active sites being >25 Å apart. The details of this long-range protein communication process were investigated by solution NMR spectroscopy and CPMG relaxation dispersion experiments. Formation of the heterodimer enzyme results in a reduction in millisecond motions in HisF that extend throughout the protein. Binding of lGP results in an increase in protein-wide millisecond dynamics evidenced as severe NMR line broadening and elevated R ex values. Together, these data demonstrate a grouping of flexible residues that link the HisF active site with the protein interface to which HisH binds and provide a model for the path of communication between the IGPS active sites.










Similar content being viewed by others
Abbreviations
- AICAR:
-
5-Aminoimidazole-4-carboxamide ribotide
- CPMG:
-
Carr-Purcell-Meiboom-Gill
- HisF-IGPS:
-
Isotopically labeled HisF in complex with HisH
- IGP:
-
Imidazole glycerol phosphate
- IGPS:
-
Imidiazole glycerol phosphate synthase (a heterodimer of HisF and HisH)
- PRFAR:
-
N′-[(5′-Phosphoribulosyl)formimino]-5-aminoimidazole-4-carboxamide-ribonucleotide
References
Amaro RE, Myers RS, Davisson VJ, Luthey-Schulten ZA (2005) Structural elements in IGP synthase exclude water to optimize ammonia transfer. Biophys J 89:475–487
Amaro RE, Sethi A, Myers RS, Davisson VJ, Luthey-Schulten ZA (2007) A network of conserved interactions regulates the allosteric signal in a glutamine amidotransferase. Biochemistry 46:2156–2173
Beach H, Cole R, Gill M, Loria JP (2005) Conservation of μs–ms enzyme motions in the apo- and substrate-mimicked state. J Am Chem Soc 127:9167–9176
Bhattacharya A, Kurochkin AV, Yip GN, Zhang Y, Bertelsen EB, Zuiderweg ER (2009) Allostery in Hsp70 chaperones is transduced by subdomain rotations. J Mol Biol 388:475–490
Boyer JA, Lee AL (2008) Monitoring aromatic picosecond to nanosecond dynamics in proteins via 13C relaxation: expanding perturbation mapping of the rigidifying core mutation, V54A, in eglin c. Biochemistry 47:4876–4886
Bruschweiler S, Schanda P, Kloiber K, Brutscher B, Kontaxis G, Konrat R, Tollinger M (2009) Direct observation of the dynamic process underlying allosteric signal transmission. J Am Chem Soc 131:3063–3068
Chaudhuri BN, Lange SC, Myers RS, Chittur SV, Davisson VJ, Smith JL (2001) Crystal structure of imidazole glycerol phosphate synthase: a tunnel through a (beta/alpha)8 barrel joins two active sites. Structure 9:987–997
Chittur SV, Chen Y, Davisson VJ (2000) Expression and purification of imidazole glycerol phosphate synthase from Saccharomyces cerevisiae. Protein Expr Purif 18:366–377
Chittur SV, Klem TJ, Shafer CM, Davisson VJ (2001) Mechanism for acivicin inactivation of triad glutamine amidotransferases. Biochemistry 40:876–887
Delaglio F, Grzesiek S, Vuister G, Zhu G, Pfeifer J, Bax A (1995) NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J Biomol NMR 6:277–293
DeLano WL (2005) MacPyMOL: a PyMOL-based molecular graphics application for MacOS X. DeLano Scientific LLC, South San Francisco, CA, USA
Douangamath A, Walker M, Beismann-Driemeyer S, Vega-Fernandez MC, Sterner R, Wilmanns M (2002) Structural evidence for ammonia tunneling across the (beta alpha)(8) barrel of the imidazole glycerol phosphate synthase bienzyme complex. Structure 10:185–193
Eaton WA, Henry ER, Hofrichter J, Mozzarelli A (1999) Is cooperative oxygen binding by hemoglobin really understood? Nat Struct Biol 6:351–358
Grzesiek S, Stahl SJ, Wingfield PT, Bax A (1996) The CD4 determinant for downregulation by HIV-1 Nef directly binds to Nef. Mapping of the Nef binding surface by NMR. Biochemistry 35:10256–10261
Kimmel JL, Reinhart GD (2000) Reevaluation of the accepted allosteric mechanism of phosphofructokinase from Bacillus stearothermophilus. Proc Natl Acad Sci U S A 97:3844–3849
Klem TJ, Davisson VJ (1993) Imidazole glycerol phosphate synthase—the glutamine amidotransferase in histidine biosynthesis. Biochemistry 32:5177–5186
Klem TJ, Chen Y, Davisson VJ (2001) Subunit interactions and glutamine utilization by Escherichia coli imidazole glycerol phosphate synthase. J Bacteriol 183:989–996
Kneller DG, Kuntz ID (1993) Ucsf sparky—an Nmr display, annotation and assignment tool. J Cell Biochem 25:4–254
Koshland DE, Nemethy G, Filmer D (1966) Comparison of experimental binding data and theoretical models in proteins containing subunits. Biochemistry 5:365–385
Krahn JM, Kim JH, Burns MR, Parry RJ, Zalkin H, Smith JL (1997) Coupled formation of an amidotransferase interdomain ammonia channel and a phosphoribosyltransferase active site. Biochemistry 36:11061–11068
Kuenzler M, Balmelli T, Egli CM, Paravicini G, Braus GH (1993) Cloning, primary structure, and regulation of the HIS7 gene encoding a bifunctional glutamine amidotransferase: cyclase from Saccharomyces cerevisiae. J Bacteriol 175:5548–5558
Lang D, Thoma R, Henn-Sax M, Sterner R, Wilmanns M (2000) Structural evidence for evolution of the beta/alpha barrel scaffold by gene duplication and fusion. Science 289:1546–1550
Lipchock J, Loria JP (2008) 1H, 15N, and 13C resonance assignment of imidazole glycerol phosphate (IGP) synthase protein HisF from Thermotoga maritima. Biomol NMR Assign 2:219–221
Loria JP, Rance M, Palmer AG (1999a) A relaxation-compensated Carr-Purcell-Meiboom-Gill sequence for characterizing chemical exchange by NMR spectroscopy. J Am Chem Soc 121:2331–2332
Loria JP, Rance M, Palmer AG (1999b) A TROSY CPMG sequence for characterizing chemical exchange in large proteins. J Biomol NMR 15:151–155
Luz Z, Meiboom S (1963) Nuclear magnetic resonance study of the protolysis of trimethylammonium ion in aqueous solution—order of the reaction with respect to solvent. J Chem Phys 39:366–370
Massi F, Wang C, Palmer AG 3rd (2006) Solution NMR and computer simulation studies of active site loop motion in triosephosphate isomerase. Biochemistry 45:10787–10794
Massiere F, Badet-Denisot MA (1998) The mechanism of glutamine-dependent amidotransferases. Cell Mol Life Sci 54:205–222
Monod J, Wyman J, Changeux J-P (1965) On the nature of allosteric transitions: a plausible model. J Mol Biol 12:88–118
Mulder FA, Skrynnikov NR, Hon B, Dahlquist FW, Kay LE (2001) Measurement of slow (micros-ms) time scale dynamics in protein side chains by (15)N relaxation dispersion NMR spectroscopy: application to Asn and Gln residues in a cavity mutant of T4 lysozyme. J Am Chem Soc 123:967–975
Myers RS, Jensen JR, Deras IL, Smith JL, Davisson VJ (2003) Substrate-induced changes in the ammonia channel for imidazole glycerol phosphate synthase. Biochemistry 42:7013–7022
Omi R, Mizuguchi H, Goto M, Miyahara I, Hayashi H, Kagamiyama H, Hirotsu K (2002) Structure of imidazole glycerol phosphate synthase from Thermus thermophilus HB8: open–closed conformational change and ammonia tunneling. J Biochem 132:759–765
Pervushin K, Riek R, Wider G, Wuthrich K (1997) Attenuated T2 relaxation by mutual cancellation of dipole-dipole coupling and chemical shift anisotropy indicates an avenue to NMR structures of very large biological macromolecules in solution. Proc Natl Acad Sci U S A 94:12366–12371
Rudolph J, Stubbe J (1995) Investigation of the mechanism of phosphoribosylamine transfer from glutamine phosphoribosylpyrophosphate amidotransferase to glycinamide ribonucleotide synthetase. Biochemistry 34:2241–2250
Schachman HK (1988) Can a simple model account for the allosteric transition of aspartate transcarbamoylase? J Biol Chem 263:18583–18586
Sinha SC, Chaudhuri BN, Burgner JW, Yakovleva G, Davisson VJ, Smith JL (2004) Crystal structure of imidazole glycerol-phosphate dehydratase: duplication of an unusual fold. J Biol Chem 279:15491–15498
Stevens SY, Sanker S, Kent C, Zuiderweg ER (2001) Delineation of the allosteric mechanism of a cytidylyltransferase exhibiting negative cooperativity. Nat Struct Biol 8:947–952
Thoden JB, Holden HM, Wesenberg G, Raushel FM, Rayment I (1997) Structure of carbamoyl phosphate synthetase: a journey of 96 A from substrate to product. Biochemistry 36:6305–6316
Velyvis A, Yang YR, Schachman HK, Kay LE (2007) A solution NMR study showing that active site ligands and nucleotides directly perturb the allosteric equilibrium in aspartate transcarbamoylase. Proc Natl Acad Sci U S A 104:8815–8820
Volkman BF, Lipson D, Wemmer DE, Kern D (2001) Two-state allosteric behavior in a single-domain signaling protein. Science 291:2429–2433
Yu EW, Koshland DE Jr (2001) Propagating conformational changes over long (and short) distances in proteins. Proc Natl Acad Sci U S A 98:9517–9520
Zalkin H, Smith JL (1998) Enzymes utilizing glutamine as an amide donor. Adv Enzymol Relat Areas Mol Biol 72:87–144
Zhu G, Xia Y, Nicholson LK, Sze KH (2000) Protein dynamics measurements by TROSY-based NMR experiments. J Magn Reson 143:423–426
Acknowledgments
JPL acknowledges support from the NIH (R01-GM070823), JML was supported by an NIH biophysical training grant (5T32GM008283). We thank Joanna Dunn (Yale University) for help in constructing His-tagged HisH and Simon Lipchock for delaying his birth until after completion of this manuscript. The authors thank V. Jo Davisson for helpful suggestions.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Lipchock, J., Loria, J.P. Millisecond dynamics in the allosteric enzyme imidazole glycerol phosphate synthase (IGPS) from Thermotoga maritima . J Biomol NMR 45, 73–84 (2009). https://doi.org/10.1007/s10858-009-9337-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10858-009-9337-8