Abstract
We show that reductive methylation of proteins can be used for highly sensitive NMR identification of conformational changes induced by metal- and small molecule binding, as well as protein–protein interactions. Reductive methylation of proteins introduces two 13C-methyl groups on each lysine in the protein of interest. This method works well even when the lysines are not actively involved in the interaction, due to changes in the microenvironments of lysine residues. Most lysine residues are located on the protein exterior, and the exposed 13C-methyl groups may exhibit rapid localized motions. These motions could be faster than the tumbling rate of the molecule as a whole. Thus, this technique has great potential in the study of large molecular weight systems which are currently beyond the scope of conventional NMR methods.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
While NMR studies of large proteins and protein–protein complexes have become more feasible due to the use of TROSY-based approaches on perdeuterated and methyl-protonated proteins (Kay and Gardner 1997; Pervushin et al. 1997; Tugarinov and Kay 2004), they are limited by the ability to produce proteins enriched with stable isotopes. Stable isotopes are most commonly introduced into recombinant proteins expressed in E. coli. It is becoming exceedingly evident that some proteins cannot easily be expressed in, and purified from, bacterial cells. Complex folding processes in a eukaryotic host are often required for proteins to achieve their proper conformations and post-translational modifications. Incorporation of stable isotopes is very difficult and often not feasible in these expression systems. This problem significantly hampers NMR-based characterization of a great number of large proteins and protein–protein complexes.
Here we show how incorporation of 13C methyl groups into the protein of interest via reductive methylation of lysine residues achieves sensitive NMR signal detection in a protein that is not otherwise isotopically enriched. In addition, we show that 13C methyl groups added to lysines could be detected with higher sensitivity than those on aliphatic amino acids in 13C enriched proteins. We postulate that the increased sensitivity in NMR signal detection may be related to superior relaxation properties of methyl groups on lysines and to degenerate chemical shifts in di-methylated sites. Thus, we propose that reductive methylation may facilitate NMR studies of large molecular weight proteins and multi-molecular protein complexes.
First introduced in 1968 (Means and Feeney 1968), reductive methylation was used by several NMR laboratories to study protein–protein and protein–nucleic acid interactions, primarily where lysine residues were directly involved in binding (Dick et al. 1988, 1989; Moore et al. 1998; Sparks et al. 1992). Numerous X-ray crystallography and NMR studies have shown that reductive methylation does not perturb protein structure (Gerken et al. 1982; Kurinov et al. 2000; Rayment 1997; Walter et al. 2006), and that methylated lysines still retain their positive charge (Rayment 1997). In addition, analysis of crystal packing in methylated lysozyme shows that reductive methylation is unlikely to induce artificial protein–protein interactions (Rypniewski et al. 1993). Zhang and Vogel used reductive methylation to measure pKa values of lysines in Calmodulin in the apo- and calcium-bound forms, as well as in complex with the myosin light chain kinase peptide (Zhang and Vogel 1993). Recently, a technique for NMR resonance assignment of methyl signals in reductively methylated proteins has been reported. It is based on mass-spectrometry and NMR measurement of rates of methylation (Macnaughtan et al. 2005). Despite these developments, the technique of reductive methylation still remains underutilized in the field of NMR.
In this paper we describe the use of reductive methylation of lysine residues with 13C enriched formaldehyde as a method for detection of protein–ligand interactions. The human calcium modulator protein Calmodulin (CaM) serves as our model system. We demonstrate that conformational changes in methylated CaM due to calcium binding, interactions with a peptide representing the hyper variable region (HVR) of human oncogene K-Ras4B, as well as small molecule (W7) binding can be detected. Evidence increasingly suggests that molecular interactions occur through the induced fit mechanism which usually involves a conformational change (Swain and Gierasch 2006; Valente et al. 2006). Therefore, the technique of reductive methylation could be used to study conformational rearrangements occurring in response to molecular interactions. In addition, this technique may be extremely useful to the pharmaceutical industry, since it provides a rapid and relatively inexpensive way to screen for potential drugs that alter protein conformations. Our studies suggest that the 13C methyl groups on lysines act as structural probes and can be used to identify protein–ligand interactions even when these residues are not directly involved in binding. This illustrates that reductive methylation could be a powerful technique in NMR studies.
Materials and methods
Human CaM was expressed and purified using modifications of previously described protocols (Gopalakrishna and Anderson 1982). Human CaM was cloned into the pET42a expression-vector (Novagen), using the NdeI and XhoI restriction sites. The protein was expressed under standard conditions in LB media using E. coli BL21-A1 cells (Invitrogen). The cells were harvested by centrifugation at 6,000 rpm at 4°C. The pellet was resuspended in 10 mM Tris/HCl pH 7.6, 5 mM CaCl2 and 10 mM β-mercaptoethanol and incubated at 65°C for 30 min. The suspension was centrifuged for 30 min at 18,000 rpm in an SS34 rotor at 4°C, and the protein was purified using a Phenyl-Sepharose® CL-4B column (Sigma). The column was washed three times with resuspension buffer and the protein was eluted using 10 mM Tris/HCl pH 7.6, 10 mM EDTA and 10 mM β-mercaptoethanol. The protein was dialyzed into 10 mM HEPES pH 7.6, 10 mM CaCl2 and 10 mM β-mercaptoethanol. Reductive methylation was performed as described (Means and Feeney 1968). Briefly, 20 μl of 1 M borane–ammonia complex (NH3–BH3) (Sigma) and 40 μl of 1 M 13C formaldehyde (Cambridge Isotope Laboratories, Inc) were added to 1 ml of protein and the reaction was incubated for 2 h with stirring at 4°C. The addition of borane–ammonia complex and 13C formaldehyde was repeated, and the mixture incubated for another 2 h. After the addition of another 10 μl of 1 M borane–ammonia complex, the reaction mixture was incubated at 4°C with stirring overnight. The reaction was stopped by adding glycine to 200 mM, and undesired reaction products as well as excess reagents were removed by dialysis against 10 mM Tris/HCl pH 7.6, 10 mM CaCl2 and 5 mM β-mercaptoethanol. The concentration of methylated CaM used in all of the 2-dimensional NMR experiments was 75 μM. To compare signal intensities of methylated CaM with uniformly 13C-labeled CaM, 13C CaM was prepared as described above except that the cells were grown in M9 media containing 13C glucose. All 1H–13C-edited HSQC spectra were collected at 25°C on a 600 MHz Bruker spectrometer equipped with a cryogenic probe at eight scans and 256 indirect points. The data were processed using NMRPipe software (Delaglio et al. 1995). The HVR of K-Ras4B has the amino acid sequence ‘KEKMSKDGKKKKKKSKTKCVIM’. W-7, the CaM antagonist, was purchased from Biomol International, LP.
Results and discussion
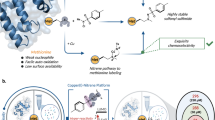
The usefulness of reductive methylation of lysines in NMR for detection of protein–ligand interactions is currently unappreciated. Here we demonstrate the potential of this easy-to-implement and largely non-perturbing technique. Reductive methylation of CaM was performed by adding 13C enriched formaldehyde to the protein solution in the presence of a reducing agent. Formaldehyde reacts with the ε-amino group of the lysine residue to form a Schiff base intermediate (Rayment 1997). Under reducing conditions the intermediate is rapidly converted to a secondary amine as illustrated in the reaction scheme shown in Fig. 1a. The reaction proceeds one more round in the presence of sufficient formaldehyde to yield a tertiary amine as shown in Fig. 1b. It should be noted that reductive methylation does not change the intrinsic charge on the lysine residue (Rayment 1997). The completeness of CaM methylation was confirmed by MALDI-TOF mass-spectrometry which indicated the presence of eight di-methylated lysines and a di-methylated N-terminus (Supplementary data, Fig. 1S).
Schematic of the reductive methylation reaction showing addition of one methyl group to an ε-amino group of a lysine residue or the N-terminal amino group through the formation of a Schiff base intermediate (a). In the presence of sufficient formaldehyde the reaction proceeds rapidly to give the di-methylated product (b)
Lysine side chains that are commonly found on the protein exterior experience unobstructed rotations. The frequency of these rotations may exceed the molecular tumbling rate. We therefore hypothesized that the methyl groups on lysine residues may possess favorable relaxation properties due to the low order parameter of the lysine side chains. To test this hypothesis we compared signal intensity of 13C methyl groups in 50 μM 13C-labeled CaM with that of 13C methyl groups introduced by reductive methylation on lysines in 50 μM unlabeled CaM. Figure 2 shows that 13C methyl groups on lysine residues can be detected by NMR with approximately ten times higher sensitivity than methyl groups in aliphatic amino acids. This characteristic makes methyl groups on lysines unique structural probes. We postulate that the order parameter of the methyl axis rotation is lower for the methyls at the ends of lysine side-chains leading to favorable relaxation times and subsequently higher intensities in NMR spectra compared to aliphatic methyls which may be buried in the hydrophobic core of the protein molecule. This implies that reductive methylation may be used to study high molecular weight protein systems by NMR. It should also be noted that the chemical shifts of the two 13C methyl groups on each lysine are often degenerate which also contributes to the higher sensitivity of the signals.
To test if the methyl groups on lysines are sensitive to a conformational change in the protein of interest, 1H–13C HSQC spectra of CaM in the presence and absence of Ca2+ were compared (Fig. 3). In the spectrum of apo-CaM only four out of eight expected signals belonging to methyl groups on lysines were observed due to signal overlap. The signal overlap may result from the fact that, due to their solvent exposure and motion averaging, the methyl groups on lysines may possess very similar chemical shift values. This constitutes a potential limitation of the use of reductive methylation in NMR. Also, in some cases where the lysines are critical for the function of the protein reductive methylation may affect protein function (Ganzhorn et al. 1996). Addition of Ca2+ causes significant shifts in three peaks and the appearance of an additional peak (Fig. 3) which illustrates the conformational differences between the Ca2+-loaded and apo-loaded forms of CaM (Komeiji et al. 2002; Weljie et al. 2003). The conformational change induced by Ca2+ binding to CaM is responsible for exposing hydrophobic pockets in the N- and C-terminal globular domains and renders CaM competent for binding to its targets (Weljie et al. 2003; Ye et al. 2006). The observation shown in Fig. 3 indicates that reductive methylation can be used for NMR detection of a conformational change resulting from metal binding.
Next, we studied binding of the K-Ras4B HVR to Ca2+-CaM. The HVR peptide was added to Ca2+-CaM in the molar ratio of 2:1. Using a fluorescent HVR fusion construct, it has been shown that HVR mediates binding of human K-Ras4B to CaM in vivo (Fivaz and Meyer 2005). The HVR is required for CaM mediated removal of K-Ras4B from the plasma membrane in response to PDGF signaling (Liao et al. 2006). Ca2+-loaded CaM is known to bind to its various targets using hydrophobic patches on its N- and C-terminal domains (Fallon et al. 2005; Meador et al. 1992; Weljie et al. 2003; Ye et al. 2006). Several peptides have been shown to induce a conformational change in Ca2+-loaded CaM upon binding. Examples are the CaM binding region of the smooth muscle myosin light chain kinase, the CaM binding region of calcineurin, and the hydrophobic IQ domain of the cardiac Ca(v)1.2 calcium channel (Fallon et al. 2005; Meador et al. 1992; Ye et al. 2006). Binding of the K-Ras4B HVR may have a similar effect. The overlay of the CaM 1H–13C HSQC spectra obtained in the presence and absence of HVR shows differences in three chemical shifts, which suggests that the HVR of K-Ras4B binds to CaM (Fig. 4). Thus, we propose that the 13C methyl groups on lysines can be used to detect conformational changes caused by protein–protein interactions. Several CaM structures have shown that all lysine residues are surface exposed, and therefore are not likely to be involved in direct interactions with the peptide of interest (Fallon et al. 2005; Meador et al. 1992; Ye et al. 2006). Although the observation of HVR binding to CaM is not novel, the use of reductive methylation for detection of this interaction confirms the utility of this technique for discovery of protein–protein interactions that are accompanied by a conformational change.
Furthermore, the technique of reductive methylation would be particularly significant if it could be used in the pharmaceutical industry for development of drug binding assays. To test our hypothesis that small molecule binding can be detected by NMR of reductively methylated lysines, a known Ca2+-CaM antagonist, W7 was used (Tanaka et al. 1982). W7 has been demonstrated to induce a conformational change in CaM. The solution structure of the Ca2+-CaM-W7 complex shows that none of the lysines are directly involved in W7 binding to the hydrophobic pocket on each of the two domains of CaM (Osawa et al. 1998).
W7 was dissolved in 50% ethanol and added to CaM in the molar ratio of 2:1. Ethanol was used as a co-solvent because W7 has limited solubility in aqueous solution. The same solution without drug was added to CaM as a negative control. The final amount of ethanol in both samples was 0.19%. 1H–13C HSQC spectra were acquired at 600 MHz on methylated CaM and CaM-W7 complex. The overlay of the two spectra shows chemical shift changes in four peaks (Fig. 5a). This illustrates that reductive methylation may be successfully used to discover small molecule binding with high sensitivity. Current NMR-based drug screening techniques rely on the use of relatively expensive and time consuming isotopic enrichment to detect small molecule binding. Alternatively, small molecule binding can be detected by observing ligand signals in diffusion based, NOE pumping or transferred NOE experiments (Zartler and Shapiro 2006). However, these methods only allow identification of low affinity binding. Our results indicate that small molecule binding causes a change in the microenvironment of lysine residues, even when they are not directly involved in the interaction with the protein. This suggests that the use of reductive methylation may provide a viable alternative to the available methods because it offers high sensitivity NMR signal detection at low protein concentrations and would allow identification of low and high affinity binding molecules. In our experiments, signals from methylated CaM could be detected at concentrations as low as 10 μM after just eight scans (data not shown). This would be particularly useful in screening small molecules for their ability to interact with a protein that is difficult to label with 13C and 15N and/or to produce in sufficient quantity to conduct conventional NMR experiments.
(a) Overlay of 1H–13C HSQC spectra of Ca2+-CaM with 0.19% ethanol (blue) as a control and Ca2+-CaM bound to W7 dissolved in 50% ethanol (red). W7 in 50% ethanol was added to Ca2+-CaM in the molar ratio of 2:1 (final ethanol concentration was 0.19%). (b) Overlay of 1H–13C HSQC spectra of Ca2+-CaM with 0.19% ethanol in the absence (blue) and presence of l-tryptophan (red) in the molar ratio 2:1 as a negative control. l-tryptophan is structurally similar to W7 but does not act as a CaM antagonist
To address the possibility that the observed chemical shift changes in lysine methyl groups may be due to crowding of hydrophobic molecules near the protein surface we also tested the effects of l-Tryptophan at the same molar ratio to Ca2+-CaM as in the case of W7. l-Tryptophan does not bind specifically to CaM and does not act as a CaM antagonist (Ouyang and Vogel 1998). Addition of l-Tryptophan did not produce any significant chemical shift changes (Fig. 5b) which confirms that the technique does not yield false positive results, and can be used reliably to detect specific molecular interactions that induce conformational changes.
Conclusion
In conclusion, we have demonstrated that reductive methylation offers a rapid, inexpensive, and sensitive way to detect conformational changes in the protein of interest due to metal binding, as well as interactions with peptides and small molecules. This is significant, since molecular interactions are often accompanied by a conformational change. Due to a reduction in NMR spectrum complexity, it may also be used as an initial approach to identify proteins that exhibit multiple conformations in slow exchange.
Abbreviations
- CaM:
-
Calmodulin
- HVR:
-
Hyper variable region of K-Ras4B
- HSQC:
-
Heteronuclear single quantum coherence
References
Delaglio F, Grzesiek S, Vuister GW, Zhu G, Pfeifer J, Bax A (1995) NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J Biomol NMR 6:277–293
Dick LR, Sherry AD, Newkirk MM, Gray DM (1988) Reductive methylation and 13C NMR studies of the lysyl residues of fd gene 5 protein. Lysines 24, 46, and 69 may be involved in nucleic acid binding. J Biol Chem 263:18864–18872
Dick LR, Geraldes CF, Sherry AD, Gray CW, Gray DM (1989) 13C NMR of methylated lysines of fd gene 5 protein: evidence for a conformational change involving lysine 24 upon binding of a negatively charged lanthanide chelate. Biochemistry 28:7896–7904
Fallon JL, Halling DB, Hamilton SL, Quiocho FA (2005) Structure of calmodulin bound to the hydrophobic IQ domain of the cardiac Ca(v)1.2 calcium channel. Structure 13:1881–1886
Fivaz M, Meyer T (2005) Reversible intracellular translocation of KRas but not HRas in hippocampal neurons regulated by Ca2+/calmodulin. J Cell Biol 170:429–441
Ganzhorn AJ, Lepage P, Pelton PD, Strasser F, Vincendon P, Rondeau JM (1996) The contribution of lysine-36 to catalysis by human myo-inositol monophosphatase. Biochemistry 35:10957–10966
Gerken TA, Jentoft JE, Jentoft N, Dearborn DG (1982) Intramolecular interactions of amino groups in 13C reductively methylated hen egg-white lysozyme. J Biol Chem 257:2894–2900
Gopalakrishna R, Anderson WB (1982) Ca2+-induced hydrophobic site on calmodulin: application for purification of calmodulin by phenyl-Sepharose affinity chromatography. Biochem Biophys Res Commun 104:830–836
Kay LE, Gardner KH (1997) Solution NMR spectroscopy beyond 25 kDa. Curr Opin Struct Biol 7:722–731
Komeiji Y, Ueno Y, Uebayasi M (2002) Molecular dynamics simulations revealed Ca(2+)-dependent conformational change of Calmodulin. FEBS Lett 521:133–139
Kurinov IV, Mao C, Irvin JD, Uckun FM (2000) X-ray crystallographic analysis of pokeweed antiviral protein-II after reductive methylation of lysine residues. Biochem Biophys Res Commun 275:549–552
Liao J, Planchon SM, Wolfman JC, Wolfman A (2006) Growth factor-dependent AKT activation and cell migration requires the function of c-K(B)-Ras versus other cellular ras isoforms. J Biol Chem 281:29730–29738
Macnaughtan MA, Kane AM, Prestegard JH (2005) Mass spectrometry assisted assignment of NMR resonances in reductively 13C-methylated proteins. J Am Chem Soc 127:17626–17627
Meador WE, Means AR, Quiocho FA (1992) Target enzyme recognition by calmodulin: 2.4 Å structure of a calmodulin–peptide complex. Science 257:1251–1255
Means GE, Feeney RE (1968) Reductive alkylation of amino groups in proteins. Biochemistry 7:2192–2201
Moore GR, Cox MC, Crowe D, Osborne MJ, Rosell FI, Bujons J, Barker PD, Mauk MR, Mauk AG (1998) N epsilon, N epsilon-dimethyl-lysine cytochrome c as an NMR probe for lysine involvement in protein–protein complex formation. Biochem J 332(Pt 2):439–449
Osawa M, Swindells MB, Tanikawa J, Tanaka T, Mase T, Furuya T, Ikura M (1998) Solution structure of calmodulin-W-7 complex: the basis of diversity in molecular recognition. J Mol Biol 276:165–176
Ouyang H, Vogel HJ (1998) Melatonin and serotonin interactions with calmodulin: NMR, spectroscopic and biochemical studies. Biochim Biophys Acta 1383:37–47
Pervushin K, Riek R, Wider G, Wuthrich K (1997) Attenuated T2 relaxation by mutual cancellation of dipole–dipole coupling and chemical shift anisotropy indicates an avenue to NMR structures of very large biological macromolecules in solution. Proc Natl Acad Sci USA 94:12366–12371
Rayment I (1997) Reductive alkylation of lysine residues to alter crystallization properties of proteins. Methods Enzymol 276:171–179
Rypniewski WR, Holden HM, Rayment I (1993) Structural consequences of reductive methylation of lysine residues in hen egg white lysozyme: an X-ray analysis at 1.8-A resolution. Biochemistry 32:9851–9858
Sparks DL, Phillips MC, Lund-Katz S (1992) The conformation of apolipoprotein A-I in discoidal and spherical recombinant high density lipoprotein particles. 13C NMR studies of lysine ionization behavior. J Biol Chem 267:25830–25838
Swain JF, Gierasch LM (2006) The changing landscape of protein allostery. Curr Opin Struct Biol 16:102–108
Tanaka T, Ohmura T, Hidaka H (1982) Hydrophobic interaction of the Ca2+-calmodulin complex with calmodulin antagonists. Naphthalenesulfonamide derivatives. Mol Pharmacol 22:403–407
Tugarinov V, Kay LE (2004) An isotope labeling strategy for methyl TROSY spectroscopy. J Biomol NMR 28:165–172
Valente AP, Miyamoto CA, Almeida FC (2006) Implications of protein conformational diversity for binding and development of new biological active compounds. Curr Med Chem 13:3697–3703
Walter TS, Meier C, Assenberg R, Au KF, Ren J, Verma A, Nettleship JE, Owens RJ, Stuart DI, Grimes JM (2006) Lysine methylation as a routine rescue strategy for protein crystallization. Structure 14:1617–1622
Weljie AM, Yamniuk AP, Yoshino H, Izumi Y, Vogel HJ (2003) Protein conformational changes studied by diffusion NMR spectroscopy: application to helix-loop-helix calcium binding proteins. Protein Sci 12:228–236
Ye Q, Li X, Wong A, Wei Q, Jia Z (2006) Structure of calmodulin bound to a calcineurin peptide: a new way of making an old binding mode. Biochemistry 45:738–745
Zartler ER, Shapiro MJ (2006) Protein NMR-based screening in drug discovery. Curr Pharm Des 12:3963–3972
Zhang M, Vogel HJ (1993) Determination of the side chain pKa values of the lysine residues in calmodulin. J Biol Chem 268:22420–22428
Acknowledgement
The research was supported by the American Cancer Society, Illinois Division Grant ACS 08-14 to Vadim Gaponenko. NMR spectra were recorded using the NMR facility at the University of Illinois at Chicago funded by the NIH Grant P41 GM68944 to Peter GW Gettins.
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Abraham, S.J., Hoheisel, S. & Gaponenko, V. Detection of protein–ligand interactions by NMR using reductive methylation of lysine residues. J Biomol NMR 42, 143–148 (2008). https://doi.org/10.1007/s10858-008-9274-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10858-008-9274-y