Abstract
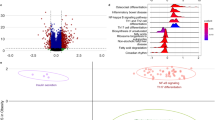
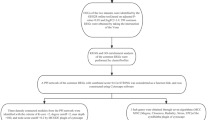
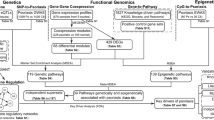
The pathogeneses of psoriasis and metabolic syndrome are closely related; however, the underlying biological mechanisms are yet to be clarified. A psoriasis training set was downloaded from the Gene Expression Omnibus database and analyzed to identify the differentially expressed genes (|logFC|> 1 and adjust P < 0.05). Differentially expressed genes for metabolic syndrome were obtained from the GeneCards, Online Mendelian Inheritance in Man, and DisGeNET databases, and crosstalk genes were obtained for multiple enrichment analysis after identifying the disease intersection. Characteristic crosstalk genes were screened using the least absolute shrinkage and selection operator regression model and random forest tree model, and the genes with area under the receiver operating characteristic curve > 0.7 were selected for validation by the two validation sets. Differential analyses of immune cell infiltration were performed on psoriasis lesion and control samples using the CIBERSORT and ImmuCellAI methods, and correlation analyses were performed between the screened signature crosstalk genes and immune cell infiltration. Significant crosstalk genes were analyzed based on the psoriasis area and severity index and on the responses to biological agents. We found five signature genes (NLRX1, KYNU, ABCC1, BTC, and SERPINB4) were screened based on two machine learning algorithms, and NLRX1 was validated. The infiltration of multiple immune cells in psoriatic lesions and non-lesions was associated with NLRX1 expression. NLRX1 was found to be associated with psoriasis severity and response rate after the use of biologics. NLRX1 could be a significant crosstalk gene for psoriasis and metabolic syndrome.








Similar content being viewed by others
Availability of Data and Materials
The data of this study are openly available in Gene Expression Omnibus (GEO, https://www.ncbi.nlm.nih.gov/geo/) database.
References
Boehncke, W.H., and M.P. Schön. 2015. Psoriasis. Lancet 386: 983–994. https://doi.org/10.1016/S0140-6736(14)61909-7.
Gisondi, P., A.C. Fostini, I. Fossà, G. Girolomoni, and G. Targher. 2018. Psoriasis and the metabolic syndrome. Clinics in Dermatology 36: 21–28. https://doi.org/10.1016/j.clindermatol.2017.09.005.
Kassi, E., P. Pervanidou, G. Kaltsas, and G. Chrousos. 2011. Metabolic syndrome: Definitions and controversies. BMC Medicine 9: 48. https://doi.org/10.1186/1741-7015-9-48.
Wu, J.J., A. Kavanaugh, M.G. Lebwohl, R. Gniadecki, and J.F. Merola. 2022. Psoriasis and metabolic syndrome: Implications for the management and treatment of psoriasis. Journal of the European Academy of Dermatology and Venereology 36: 797–806. https://doi.org/10.1111/jdv.18044.
Corbetta, S., R. Angioni, A. Cattaneo, P. Beck-Peccoz, and A. Spada. 2006. Effects of retinoid therapy on insulin sensitivity, lipid profile and circulating adipocytokines. European Journal of Endocrinology 154: 83–86. https://doi.org/10.1530/eje.1.02057.
Shimobayashi, M., V. Albert, B. Woelnerhanssen, I.C. Frei, D. Weissenberger, A.C. Meyer-Gerspach, N. Clement, S. Moes, M. Colombi, J.A. Meier, M.M. Swierczynska, P. Jenö, C. Beglinger, R. Peterli, and M.N. Hall. 2018. Insulin resistance causes inflammation in adipose tissue. The Journal of Clinical Investigation 128: 1538–1550. https://doi.org/10.1172/JCI96139.
Newman, A.M., C.L. Liu, M.R. Green, A.J. Gentles, W. Feng, X. Yue, C.D. Hoang, M. Diehn, and A.A. Alizadeh. 2015. Robust enumeration of cell subsets from tissue expression profiles. Nature Methods 12: 453–457. https://doi.org/10.1038/nmeth.3337.
Miao, Y.R., Q. Zhang, Q. Lei, M. Luo, G.Y. Xie, H. Wang, and A.Y. Guo. 2020. ImmuCellAI: A unique method for comprehensive T-cell subsets abundance prediction and its application in cancer immunotherapy. Advanced Science (Weinh) 7: 1902880. https://doi.org/10.1002/advs.201902880.
Banerjee, S., A. Biehl, M. Gadina, S. Hasni, and D.M. Schwartz. 2017. JAK-STAT signaling as a target for inflammatory and autoimmune diseases: Current and future prospects. Drugs 77: 521–546. https://doi.org/10.1007/s40265-017-0701-9.
Peng, H., D. Xian, J. Liu, S. Pan, R. Tang, and J. Zhong. 2020. Regulating the polarization of macrophages: A promising approach to vascular dermatosis. Journal of Immunology Research 2020: 8148272. https://doi.org/10.1155/2020/8148272.
Soleimani, R., M. Mohammadi, S.A. Saghebi, A. Taghipour, A.K. Vakilzadeh, and J.T. Afshari. 2020. Comparison of Th1/Th2 and Treg/Th17 ratios between wet and dry cupping therapies in Persian medicine. Avicenna Journal of Phytomedicine 10: 24–34.
Funes, S.C., M. Rios, J. Escobar-Vera, and A.M. Kalergis. 2018. Implications of macrophage polarization in autoimmunity. Immunology 154: 186–195. https://doi.org/10.1111/imm.12910.
Sun, C., L. Sun, H. Ma, Y. Jianxia Peng, K.D. Zhen, G. Liu, W. Ding, and Y. Zhao. 2012. The phenotype and functional alterations of macrophages in mice with hyperglycemia for long term. Journal of Cellular Physiology 227: 1670–1679. https://doi.org/10.1002/jcp.22891.
Kaneko, F., N. Itoh, H. Yoshida, M. Suzuki, and I. Ono. 1991. The cell-components and cytokines in the subcorneal microabscess of psoriasis. Fukushima Journal of Medical Science 37: 103–112.
Dunphy, S.E., C.M. Sweeney, G. Kelly, A.M. Tobin, B. Kirby, and C.M. Gardiner. 2017. Natural killer cells from psoriasis vulgaris patients have reduced levels of cytotoxicity associated degranulation and cytokine production. Clinical Immunology 177: 43–49. https://doi.org/10.1016/j.clim.2015.10.004.
Arican, O., M. Aral, S. Sasmaz, and P. Ciragil. 2005. Serum levels of TNF-alpha, IFN-gamma, IL-6, IL-8, IL-12, IL-17, and IL-18 in patients with active psoriasis and correlation with disease severity. Mediators of Inflammation 2005: 273–279. https://doi.org/10.1155/MI.2005.273.
Roh, N.K., S.H. Han, H.J. Youn, Y.R. Kim, Y.W. Lee, Y.B. Choe, and K.J. Ahn. 2015. Tissue and serum inflammatory cytokine levels in Korean psoriasis patients: A comparison between plaque and guttate psoriasis. Annals of Dermatology 27: 738–743. https://doi.org/10.5021/ad.2015.27.6.738.
Michalak-Stoma, A., J. Bartosińska, M. Kowal, M. Juszkiewicz-Borowiec, A. Gerkowicz, and G. Chodorowska. 2013. Serum levels of selected Th17 and Th22 cytokines in psoriatic patients. Disease Markers 35: 625–631. https://doi.org/10.1155/2013/856056.
Takahashi, H., H. Tsuji, Y. Hashimoto, A. Ishida-Yamamoto, and H. Iizuka. 2010. Serum cytokines and growth factor levels in Japanese patients with psoriasis. Clinical and Experimental Dermatology 35: 645–649. https://doi.org/10.1111/j.1365-2230.2009.03704.x.
Lowes, M.A., T. Kikuchi, J. Fuentes-Duculan, I. Cardinale, L.C. Zaba, A.S. Haider, E.P. Bowman, and J.G. Krueger. 2008. Psoriasis vulgaris lesions contain discrete populations of Th1 and Th17 T cells. The Journal of Investigative Dermatology 128: 1207–1211. https://doi.org/10.1038/sj.jid.5701213.
Yilmaz, S.B., N. Cicek, M. Coskun, O. Yegin, and E. Alpsoy. 2012. Serum and tissue levels of IL-17 in different clinical subtypes of psoriasis. Archives of Dermatological Research 304: 465–469. https://doi.org/10.1007/s00403-012-1229-1.
Sato, Y., E. Ogawa, and R. Okuyama. 2020. Role of innate immune cells in psoriasis. International Journal of Molecular Sciences 21: 6604. https://doi.org/10.3390/ijms21186604.
Rosário, M., and I. Azevedo. 2010. Chronic inflammation in obesity and the metabolic syndrome. Mediators of Inflammation 2010: 289645. https://doi.org/10.1155/2010/289645.
Grandl, G., and C. Wolfrum. 2018. Hemostasis, endothelial stress, inflammation, and the metabolic syndrome. Seminars in Immunopathology 40: 215–224. https://doi.org/10.1007/s00281-017-0666-5.
Demir, E., N.O. Harmankaya, İK. Utku, G. Açıksarı, T. Uygun, H. Özkan, and B. Demir. 2019. The relationship between epicardial adipose tissue thickness and serum interleukin-17a level in patients with isolated metabolic syndrome. Biomolecules 9: 97. https://doi.org/10.3390/biom9030097.
Snäkä, T., and N. Fasel. 2020. Behind the scenes: nod-like receptor X1 controls inflammation and metabolism. Frontiers in Cellular and Infection Microbiology 10: 609812. https://doi.org/10.3389/fcimb.2020.609812.
Chen, L., S.Q. Cao, Z.M. Lin, S.J. He, and J.P. Zuo. 2021. NOD-like receptors in autoimmune diseases. Acta Pharmacologica Sinica 42: 1742–1756. https://doi.org/10.1038/s41401-020-00603-2.
Li, S., P. Kang, W. Zhang, Z. Jian, Q. Zhang, X. Yi, S. Guo, W. Guo, Q. Shi, B. Li, P. Yuanmin He, L.L. Song, K. Li, G. Wang, T. Gao, and C. Li. 2020. Activated NLR family pyrin domain containing 3 (NLRP3) inflammasome in keratinocytes promotes cutaneous T-cell response in patients with vitiligo. The Journal of Allergy and Clinical Immunology 145: 632–645. https://doi.org/10.1016/j.jaci.2019.10.036.
Tervaniemi, M.H., S. Katayama, T. Skoog, H.A. Siitonen, J. Vuola, K. Nuutila, R. Sormunen, A. Johnsson, S. Linnarsson, S. Suomela, E. Kankuri, J. Kere, and O. Elomaa. 2016. NOD-like receptor signaling and inflammasome-related pathways are highlighted in psoriatic epidermis. Scientific Reports 6: 22745. https://doi.org/10.1038/srep22745.
Tang, L., Y. Cheng, C. Zhu, L. Chao Yang, Y.Z. Liu, L. Wen, X. Zhang, F. Zhou, and S. Yang. 2018. Integrative methylome and transcriptome analysis to dissect key biological pathways for psoriasis in Chinese Han population. Journal of Dermatological Science 91: 285–291. https://doi.org/10.1016/j.jdermsci.2018.06.001.
Costa, F.R., M.C. Françozo, G.G. de Oliveira, A. Ignacio, A. Castoldi, D.S. Zamboni, S.G. Ramos, N.O. Câmara, M.R. de Zoete, N.W. Palm, R.A. Flavell, J.S. Silva, and D. Carlos. 2016. Gut microbiota translocation to the pancreatic lymph nodes triggers NOD2 activation and contributes to T1D onset. The Journal of Experimental Medicine 213: 1223–1239. https://doi.org/10.1084/jem.20150744.
Gao, Y., J. Li, S. Chu, Z. Zhang, N. Chen, L. Li, and L. Zhang. 2020. Ginsenoside Rg1 protects mice against streptozotocin-induced type 1 diabetic by modulating the NLRP3 and Keap1/Nrf2/HO-1 pathways. European Journal of Pharmacology 866: 172801. https://doi.org/10.1016/j.ejphar.2019.172801.
Fekete, T., D. Bencze, E. Bíró, S. Benkő, and K. Pázmándi. 2021. Focusing on the cell type specific regulatory actions of NLRX1. International Journal of Molecular Sciences 22: 1316. https://doi.org/10.3390/ijms22031316.
Singh, K., L. Sripada, A. Lipatova, M. Roy, P. Prajapati, D. Gohel, K. Bhatelia, P.M. Chumakov, and R. Singh. 2018. NLRX1 resides in mitochondrial RNA granules and regulates mitochondrial RNA processing and bioenergetic adaptation. Biochimica et Biophysica Acta. Molecular Cell Research 1865: 1260–1276. https://doi.org/10.1016/j.bbamcr.2018.06.008.
Yang, Q., X. Jiean, Q. Ma, Z. Liu, V. Sudhahar, Y. Cao, L. Wang, X. Zeng, Y. Zhou, M. Zhang, X. Yiming, Y. Wang, N.L. Weintraub, C. Zhang, T. Fukai, W. Chaodong, L. Huang, Z. Han, T. Wang, D.J. Fulton, M. Hong, and Y. Huo. 2018. PRKAA1/AMPKalpha1-driven glycolysis in endothelial cells exposed to disturbed flow protects against atherosclerosis. Nature Communications 9: 4667. https://doi.org/10.1038/s41467-018-07132-x.
Kornberg, M.D., P. Bhargava, P.M. Kim, V. Putluri, A.M. Snowman, N. Putluri, P.A. Calabresi, and S.H. Snyder. 2018. Dimethyl fumarate targets GAPDH and aerobic glycolysis to modulate immunity. Science 360: 449–453. https://doi.org/10.1126/science.aan4665.
Soares, F., I. Tattoli, M.A. Rahman, S.J. Robertson, A. Belcheva, D. Liu, C. Streutker, S. Winer, D.A. Winer, A. Martin, D.J. Philpott, D. Arnoult, and S.E. Girardin. 2014. The mitochondrial protein NLRX1 controls the balance between extrinsic and intrinsic apoptosis. The Journal of Biological Chemistry 289: 19317–19330. https://doi.org/10.1074/jbc.M114.550111.
Tattoli, I., L.A. Carneiro, M. Jéhanno, J.G. Magalhaes, Y. Shu, D.J. Philpott, D. Arnoult, and S.E. Girardin. 2008. NLRX1 is a mitochondrial NOD-like receptor that amplifies NF-kappaB and JNK pathways by inducing reactive oxygen species production. EMBO Reports 293–300. https://doi.org/10.1038/sj.embor.7401161.
Novoszel, P., M. Holcmann, G. Stulnig, C. De Sa Fernandes, V. Zyulina, I. Borek, M. Linder, A. Bogusch, B. Drobits, T. Bauer, C. Tam-Amersdorfer, P.M. Brunner, G. Stary, L. Bakiri, E.F. Wagner, H. Strobl, and M. Sibilia. 2021. Psoriatic skin inflammation is promoted by c-Jun/AP-1-dependent CCL2 and IL-23 expression in dendritic cells. EMBO Molecular Medicine e12409. https://doi.org/10.15252/emmm.202012409.
Hammouda, M., A. Ford, Y. Liu, and J. Zhang. 2020. The JNK signaling pathway in inflammatory skin disorders and cancer. Cells 857. https://doi.org/10.3390/cells9040857.
Chu, X., W. Songwei, and R. Raju. 2019. NLRX1 regulation following acute mitochondrial injury. Frontiers in Immunology 10: 2431. https://doi.org/10.3389/fimmu.2019.02431.
Acknowledgements
We thank all authors for their contributions.
Funding
The authors declare that no funds, grants, or other support were received during the preparation of this manuscript.
Author information
Authors and Affiliations
Contributions
Yang Zhou wrote the manuscript and generated figures. Lu Han, Ziting Wang, Ning Guan, Runan Fang, Yue Wan, and Zeyu Yang contributed to editing the manuscript. Jianhong Li and Qing Ni supervised the study and approved the submission of this research article. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhou, Y., Han, L., Wang, Z. et al. Bioinformatic Analysis of the Potential Common Pathogenic Mechanisms for Psoriasis and Metabolic Syndrome. Inflammation 46, 1381–1395 (2023). https://doi.org/10.1007/s10753-023-01815-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10753-023-01815-4