Abstract
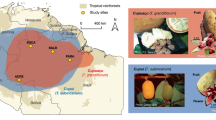
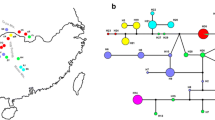
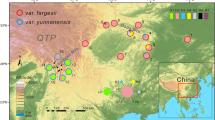
Jacaratia mexicana A. DC. (Caricaceae) is a tropical tree distributed throughout Mexico and Mesoamerica. Some evidence in Mexico indicates the presence of an incipient domestication process in this species. Phylogeographical analyses can potentially determine contemporary patterns of gene flow, isolation between population lineages, as well as historical processes such as population bottlenecks or expansions on their geographical areas. In this study we reconstruct the phylogeographical patterns in populations of J. mexicana A. DC., in order to find differences between genetic variation among wild and cultivated populations utilizing chloroplast DNA and nuclear DNA sequences. We generate a Bayesian phylogenetic tree, to estimate the divergence time between clades using calibrated mutation rates. We also infer the demographic history of these populations using neutrality tests among wild and cultivated accessions. We identified higher levels of haplotype and nucleotide diversity for the cpDNA and ITS types in wild populations than in domesticated populations. These results indicate a reduction of genetic diversity derived from human selection on domestication traits. Neutrality test suggests population expansion detected by the significant negative values of Fu’s Fs in the cultivated populations of this specie. These process results in an excess of rare polymorphism with the fixation of certain advantageous mutation throughout time, this implication are in accordance with the role of the strong selection in the fruit traits of J. mexicana. The dated phylogeny constructed with BEAST program indicated a dispersion pattern for the J. mexicana ancestors across the South Pacific and South Eastern populations during the late Pliocene. Posterior dispersion and divergence in the clades from Central Mexico and North Pacific are in agreement with the episodes of mountain-building in different regions of Mexico.



Similar content being viewed by others
References
Aradhya MK, Manshardt RM, Zee F, Morden CW (1999) A phylogenetic analysis of the genus Carica L. (Caricaceae) based on restriction fragment length variation in a cpDNA intergenic spacer region. Genet Resour Crop Evol 46:579–586
Avise JC (2000) Phylogeography. The history and formation of species. Harvard University Press, Cambridge
Baldwin BG (1992) Phylogenetic utility of the internal transcribed spacers of nuclear ribosomal DNA in plants: an example from the Compositae. Mol Phylogenet Evol 1:3–16
Bandelt HJ, Forster P, Sykes BC, Richards MB (1995) Mitochondrial portraits of human populations. Genetics 141:743–753
Benz BF (2001) Archaelogical evidence of teosinte domestication from Guila naquitz, Oaxaca. PNAS 98:2104–2106
Briones MR (2002) Timbiriche y cuaguayote: plantas milenarias en extinción. Hypatia 4:1–3
Brown JW, Rest JS, García-Moreno J, Sorenson MD, Mindell DP (2008) Strong mitochondrial DNA support for a Cretaceous origin of modern avian lineages. BMC Biol 6:1170–1186
Caicedo AL, Schaal BA (2004) Population structure and phylogeography of Solanum pimpinellifolium inferred from a nuclear gene. Mol Ecol 13:1871–1882
Chiang TY, Schaal BA, Peng T (1998) Universal primers for amplification and sequencing a noncoding spacer between the atpB and rbcL genes of chlorolast DNA. Bot Bull Acad Sin 39:245–250
Crandall ED, Frey MA, Grosberg RK, Barber PH (2008) Contrasting demographic history and phylogeographical patterns in two Indo-Pacific gastropods. Mol Ecol 17:611–626
Dane F, Lang P (2004) Comparative analysis of cpDNA variability in wild and cultivated Citrullus species: implications for evolution of watermelon. Am J Bot 91:1922–1929
Doebley J (1989) Isozymic evidence and the evolution of crop plants. In: Soltis D, Soltis P (eds) Isozymes in plant biology. Dioscorides Press, Portland, Oregon, pp 165–191
Drummond AJ, Ho SYW, Phillips MJ, Rambaut A (2006) Relaxed phylogenetics and dating with confidence. PLOS Biol 4:699–710
Ellstrand NC, Prentice HC, Hancock JF (1999) Gene flow and introgression from domesticated plants into their wild relatives. Ann Rev Ecol Syst 30:539–563
Excoffier L, Laval G, Schneider S (2005) Arlequin ver. 3.0: an integrated software package for population genetics data analysis. EBO 1:47–50
Fernández IA, Aguilar JF, Panero JL, Nieto GF (2001) A phylogenetic analysis of Doronicum (Asteraceae: Senecioneae) based on morphological, nuclear ribosomal (ITS) and chloroplast (TrnL-F) evidence. Mol Phylogenet Evol 20:41–64
Fu YX (1996) New statistical tests of neutrality for DNA samples from a population. Genetics 143:557–570
Gepts P, Papa R (2002) Evolution during domestication. Encyclopedia of life sciences. Macmillan Publisher Ltd, New York, pp 1–6
Graham A (1999) Studies in Neotropical paleobotany. XIII. An Oligo-Miocene palynoflora from Simojovel (Chiapas). Am J Bot 86:17–31
Guizar N, Sánchez V (1991) Guia para el reconocimiento de los Principales árboles del Alto Balsas. Universidad Autonoma de Chapingo, México, pp 1–207
Guo YL, Ge S (2005) Molecular phylogeny of Oryzeae (Poaceae) based on DNA sequences from chloroplast, mitochondrial, and nuclear genomes. Am J Bot 92:1548–1558
Hamrick JL, Godt MJW (1997) Allozyme diversity in cultivated crops. Crop Sci 37:26–31
Harlan JR (1975) Our vanishing genetic resources. Science 188:618–621
Harpending HC, Sherry ST, Rogers AR, Stoneking M (1993) The genetic structure of ancient human populations. Curr Anthropol 34:483–496
Heiser CB (1979) Origins of some cultivated new world plants. Ann Rev Ecol Evol Syst 10:309–326
Hillman GC, Davies MS (1990) Measured domestication rates in wild wheats and barley under primitive cultivation, and their archaeological implications. J World Prehist 4:157–222
Innan H, Stephan W (2000) The coalescent in an exponentially growing metapopulation and its application to Arabidopsis thaliana. Genetics 155:2015–2019
Jakob SS, Martínez-Meyer E, Blattner FR (2009) Phylogeographic analyses and paleodistribution modeling indicate Pleistocene in situ survival of Hordeum species (Poaceae) in Southern Patagonia without genetic or spatial restriction. MBE 26:907–923
Koenig R, Gepts P (1989) Allozime diversity in wild Phaseolus vulgaris: further evidence for two major centers of genetic diversity. Theor Appl Genet 78:809–817
López A, Luján L (2001) El pasado indígena. Fondo de Cultura Económica. El Colegio de México, México
Lozano-García MS, Ortega-Guerrero B (1997) Late Quaternary environmental changes of the central part of the Basin of Mexico; correlation between Texcoco and Chalco Basins. Rev Palaeobot Palynol 99:77–93
Lozano-Garcia MS, Xelhuantzi-Lopez MS (1997) Some problems in the late Quaternary pollen records of central Mexico and Zacapu. Quat Int 43:117–123
Metcalfe SE, O’Hara SL, Caballero M, Davies SJ (2000) Records of late Pleistocene Holocene climatic change in Mexico: a review. Quat Sci Rev 19:699–721
Miller A, Schaal B (2006) Domestication and the distribution of genetic variation in wild and cultivated populations of the Mesoamerican fruit tree Spondias purpurea L. (Anacardiaceae). Mol Ecol 15:1467–1480
Miranda F, Hernández-X E (1963) Los tipos de vegetación de México y su clasificación. Vol Soc Bot Mex 28:29–178
Moore BR, Donoghue MJ (2007) Correlates of diversification in the plant clade Dipsacales: geographic movement and evolutionary innovations. Am Nat 170:28–55
Moreno NP (1980) Flora de Veracruz. INIREB 10:1–17
Niembro R (1986) Árboles y Arbustos utiles de México: naturales e introducidas. Limusa, Mexico, p 206
Piperno DR, Moreno JE, Iriarte J, Holst I, Lachniet M, Jones JG, Ranere AJ, Castanzo R (2007) Late pleistocene and holocene environmental history of the iguala valley, central balsas watershed of Mexico. PNAS 104:11874–11881
Posada D, Crandall KA (1998) Modeltest: testing the model of DNA substitution. Bioinformatics 14:817–818
Rambaut A (2006) FigTree v1.1. Institute of Evolutionary Biology University of Edinburgh, Edinburgh
Ramos-Onsins SE, Rozas J (2002) Statistical properties of neutrality test against population growth. Mol Biol Evol 19:2092–2100
Raven P, Axelrod D (1974) Angiosperm biogeography and past continental movements. Ann Miss Bot Gard 61:539–673
Richardson JE, Pennington RT, Pennington TD, Hollingsworth PM (2001) Rapid diversification of a species-rich genus of neotropical rain forest trees. Science 293:2242–2245
Rogers AR, Harpending H (1992) Population growth makes waves in the distribution of pairwise genetic differences. Mol Biol Evol 9:552–569
Rossotti A, Ferrari L, López-Martínez M, Rosas-Elguera J (2002) Geology of the boundary between the Sierra Madre Occidental and the Trans-Mexican Volcanic Belt in the Guadalajara region, western Mexico. Rev Mex Cienc Geol 19:1–15
Rozas J, Rozas R (1999) Dnasp version 3.0: an integrated program for molecular population genetics and molecular evolution analysis. Bioinformatics 15:174–175
Rutschmann F (2006) Molecular dating of phylogenetic trees: a brief review of current methods that estimate divergence times. Divers Distrib 12:35–48
Rzedowski J, Equihua M (1987) Atlas Cultural de México: Flora. Planeta/INAH. Grupo Editorial Planeta, Mexico, DF, p 223
Sanders WT, Price B (1968) Mesoamerica: The Evolution of a Civilization. Random House, New York
Schaal BA, Hayworth DA, Olsen KM, Rauscher JT, Smith WA (1998) Phylogeographic studies in plants: problems and prospects. Mol Ecol 7:465–474
Smith BD (1997) The initial domestication of Cucurbita pepo in the Americas 10, 000 years ago. Science 276:932–934
Standley CP (1924) Trees and shrubs of México. Washington Government Printing Office, USA
Stock M, Sicilia A, Belfiore NM, Buckley D, Lo Bruto S, Lo Valvo M, Arculeo M (2008) Post-Messinian evolutionary relationships across the Sicilian channel: mitochondrial and nuclear markers link a new green toad from Sicily to African relatives. BMC Evol Biol 8:56
Tavaré S (1986) Some probabilistic and statistical problems in the analysis of DNA sequences. In: Miura RM (ed) Some mathematical questions in biology – DNA sequence analysis. American Mathematical Society, Providence, RI, pp 57–86
Valenzuela D, Ceballos G (2000) Habitat selection, home range, and activity of the white-nosed coati (Nasua narica) in a Mexican tropical dry forest. J Mammal 81:810–819
Van Devender TR (1990) Late quaternary vegetation and climate of the Chihuahuan Desert, United States and Mexico. In: Betancourt JL, Van Devender TR, Martin PS (eds) Packrat middens. The last 40,000 years of biotic change. University of Arizona Press, Tucson, pp 104–133
Vavilov NI (1926) The centres of origin of cultivated plants. Bull Appl Bot Genet Plant Breed 16:1–248
Walsh BM, Hoot SB (2001) Phylogenetic of Capsicum based on AtpB-rbcL and waxy genes. Int J Plant Sci 162:1409–1418
White TJ, Bruns T, Lee S, Taylor JW (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: Innis MA, Gelfand DH, Sninsky JJ, White TJ (eds) PCR protocols: a guide to methods and applications. Academic Press, Inc, New York, pp 315–322
Wolfe KH, Li WH, Sharp PM (1987) Rates of nucleotide substitution vary greatly among plant mitochondria, chloroplast, and nuclear DNAs. Proc Natl Acad Sci USA 84:9054–9058
Wright SI, Gaut BS (2005) Molecular population genetics and the search for adaptive evolution in plants. Mol Biol Evol 22:506–519
Acknowledgments
We are grateful to AF Castorena, FX Gonzalez-Cozatl, E Arellano, E Quijano, O Pacheco, R Ramírez and B Maldonado for technical assistance and suggested improvements to the manuscript. Also thanks to JC Juarez and J Almonte for assistance in the field collection.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Arias, D., Peñaloza-Ramírez, J., Dorado, O. et al. Phylogeographic patterns and possible incipient domestication of Jacaratia mexicana A. DC. (Caricaceae) in Mexico. Genet Resour Crop Evol 57, 1227–1238 (2010). https://doi.org/10.1007/s10722-010-9569-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-010-9569-1